In a new study published in the journal Cell (“Engineered virus-like particles for efficient in vivo delivery of therapeutic proteins”), a team of researchers led by David Liu, PhD, professor at Harvard University and the Broad Institute, engineered DNA-free virus-like particles (eVLPs) that can efficiently deliver base editors or Cas9 ribonucleoproteins to multiple organs with minimal off-target effects.
While base editing provides a powerful protocol for treating currently untreatable genetic disorders, efficient delivery of base editors to relevant cell types, and leaving no telltale footprints in the form of persistent expression of editing enzymes that could lead to off-target effects, remain a challenge.
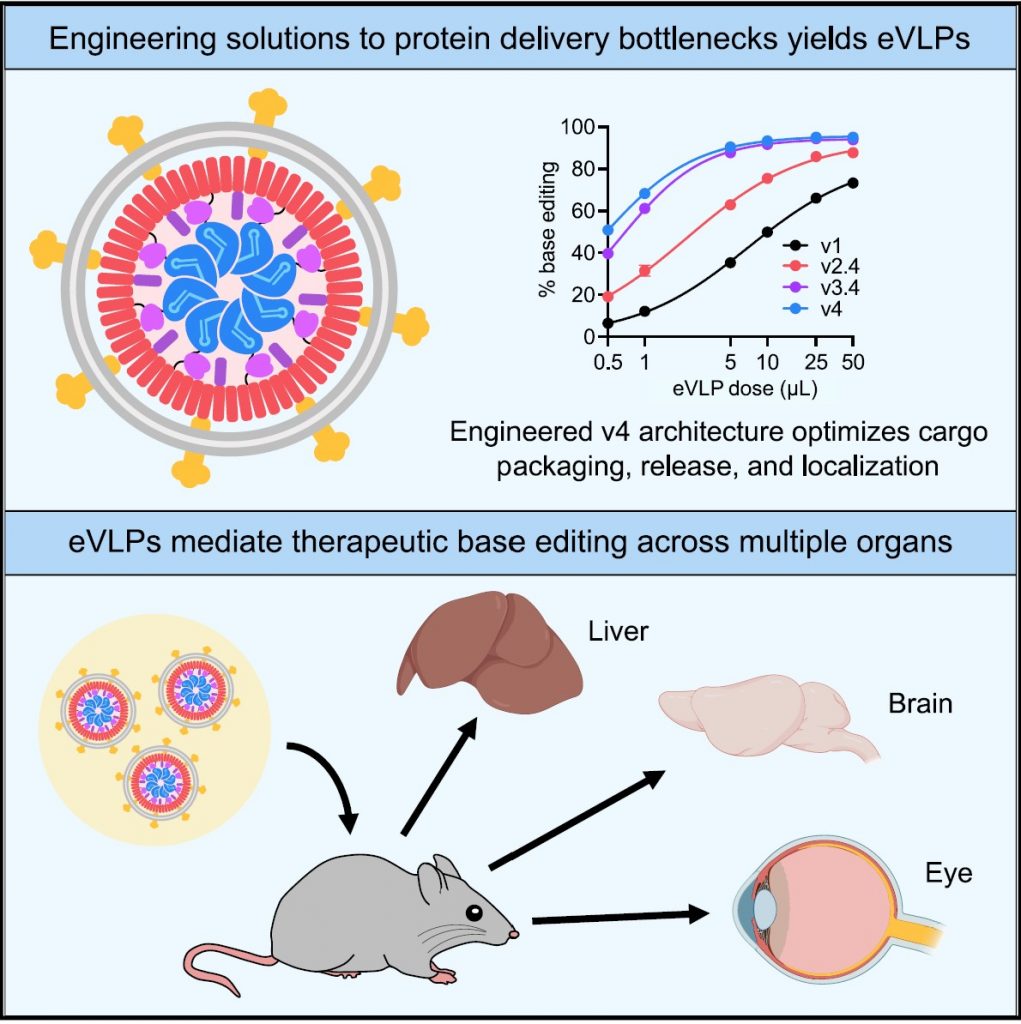
The fourth-generation eVLPs reported in the current study overcome earlier design limitations including the packaging, release, and targeting of contents. The authors demonstrated the successful delivery of base-editing machinery in primary mouse and human cell types and in three diseases modeled in mice. Through single injections, they attained therapeutic levels of organ-specific base editing in the brain, eye, and liver that significantly enhanced editing efficiency over earlier attempts.
“For gene editing as an approach to treat genetic diseases in vivo, delivery of the gene editor to the right tissue remains a challenge. Though lipid nanoparticles have emerged as a robust approach for delivery to the liver, accessing organs beyond the liver has been tough,” said Sekar Kathiresan, MD, co-founder and CEO of Verve Therapeutics.
“Our findings suggest eVLPs as an attractive alternative to other delivery strategies for the in vivo or ex vivo delivery of base editors, nucleases, and other proteins of therapeutic interest,” the authors noted.
“Here, Liu and colleagues develop a promising approach to deliver ribonucleoprotein—the base editor protein and the guide RNA,” said Kathiresan, who was not involved in this study. “Particular advantages of this approach seem to be the ability to manipulate the tissue to which the VLP will go to, as well as the short half-life of the base editor when delivered as a ribonucleoprotein.”
“It is exciting to follow the story and evolution of this alternative, nonviral eVLP system that will complement the current repertoire of delivery modalities for gene-editing applications,” said Jennifer Harbottle, PhD, senior scientist at Horizon, a Perkin Elmer company, who also wasn’t part of the study. “The transient nature of the delivery system, alongside its tissue tropism, allow high editing at the intended target site within the target organ and minimize the occurrence of potentially deleterious off target effects—that could occur with viral delivery systems, and even more so when delivering standard CRISPR-Cas rather than ABE.”
“Base editors provide a safer genome editing approach but continued expression of the cDNA encoding the base editor increases the chance of off-target effects in the host cell,” said Jyoti Jaiswal, PhD, senior investigator and associate director of academics for the Center for Genetic Medicine Research at Children’s National, Washington DC, who was not involved in the current study. “Banskota et al., present a means to effectively overcome this limitation by delivering the base editor as RNPs. The short half-life of RNPs cause the editing machinery to disappear from the cell not too long after delivery into the cells.”
The authors demonstrated the application of the improved fourth generation (v4) eVLP base editor delivery platform by correcting a mutation in the gene Rpe65 in a layer of cells behind the retina to restore vision in a mouse model of genetic blindness and knocking down the gene PCSK9 in mice to lower levels of low-density lipoprotein (LDL) cholesterol in the blood, reducing the risk of atherosclerotic cardiovascular disease. The team also installed a silent mutation in the Dmnt1 gene in newborn mice midbrains and cerebral cortices, showing eVLPs can get base editing molecules into hard-to-reach nooks in the central nervous system.
Liu said, “The in vivo delivery of proteins into target cells remains a major challenge that limits the impact of gene editing, gene therapy, and other macromolecular therapeutics. VLPs in principle offer some of the best aspects of viral delivery (high transduction efficiency, specific tissue tropism, inherent endosomal escape) and of nonviral delivery (short-lived exposure that minimizes off-target editing, no possibility of oncogenic DNA integration). But VLPs have shown low protein delivery efficiency in vivo thus far.”
Base editing has been used to correct pathogenic point mutations in mice and nonhuman primates, but earlier studies have mostly used adeno-associated viruses (AAVs) to deliver DNA that expresses base editors in target tissues. DNA delivery causes prolonged expression of base editing machinery in transduced cells engendering off-target footprints, possibilities of integrating viral DNA into the host genome, and an increased likelihood of cancer.
These drawbacks can be overcome by delivering proteins or ribonucleoproteins (RNPs) instead, which decreases off-target changes without compromising on-target edits. Liu and other labs have locally injected lipid nanoparticles (LNPs) in the ear and eye, carrying base editing RNPs in earlier studies but a generalizable protocol for delivering base editing RNPs to multiple organs and tissues in live animals has not been formulated, until now.
“To unlock the potential of VLPs to mediate potent in vivo delivery of therapeutically relevant proteins including base editors and Cas9 nuclease, we systematically identified and engineered solutions to three major bottlenecks—cargo release, cargo packaging and localization, and VLP component stoichiometry—that reduce the potency of VLP-mediated protein delivery,” said Liu. “Combining these three solutions resulted in fourth-generation eVLPs that greatly enhance protein delivery potency into mammalian cells both in vitro and in vivo. Delivery of base editors into a wide variety of immortalized and primary human and mouse cells both in tissue culture and in the liver, brain, and retina in live mice was much more potent than first-generation VLPs based on previously reported architectures, typically resulting in 5- to 40-fold higher editing levels.”
Furthermore, using eVLPs the authors reported negligible in vitro and in vivo off-target editing—an improvement over AAV or plasmid delivery. In addition, while LNP-mediated delivery of base editor mRNA is also transient, RNP delivery shortens exposure to editing agents even more, lowering the likelihood of off-target editing further.
“The eVLPs also carry virtually no risk of DNA integration given that they are DNA-free delivery systems,” said Liu.
A bonus to eVLP-mediated delivery is the possibility of engineering specific docking locations. “We could alter the cell-type preferences (tropism) of eVLPs by using different glycoproteins,” said Liu.
The authors have explored whether v4 base-editing eVLPs can be programmed to transduce specific cell types by modulating their envelop glycoproteins. For instance, they show base editing eVLPS with the FuG-B2 envelope glycoprotein targets the particles to efficiently transduce and edit a neuroblastoma cell line but not mouse fibroblasts. Swapping in glycoproteins opens the possibility of targeting eVLPS to other cell-specific targets.
“These results establish eVLPs as promising vehicles for therapeutic macromolecule delivery that combine advantages of both viral and nonviral delivery,” said Liu.
Toward therapeutic solutions, Liu’s team explores the ability of eVLPs to mediate base editing in adult animals. They target Pcsk9, a gene involved in cholesterol homeostasis whose function is naturally lost in a section of the human population, imparting an advantageous lower risk for hypercholesterolemia with no apparent adverse side effects.
The authors showed a single injection of eVLPs in mice results in therapeutic levels of base editing and reduces serum levels of Pcsk9 by 78% following 63% liver editing. They also showed v4 eVLPs are 26-fold more efficient in editing Pcsk9 in the mouse liver compared to a first-generation design attempting the same. This the authors noted, underscores “the importance of engineering VLP architectures for in vivo editing.”
The team also restored visual function in an adult mouse model of a retinal disorder (Leber congenital amaurosis, LCA) through a single subretinal injection of v4 base editing-eVLPs, achieving editing efficiencies and levels of rescue that are higher than those previously achieved.
“The accessibility of the eyes and their immune-privileged status may more readily enable the translation of new delivery modalities into preclinical and clinical studies,” the authors wrote. “Our data provide evidence of the therapeutic potential of BE-eVLPs as a means to correct pathogenic point mutations that cause ocular disorders.”
Harbottle said, “The study also demonstrates the strategic use of specific base editor variants to achieve the desired therapeutic outcome; in this case, swapping out ABE8e-NG for ABE7.10-NG which has a narrower editing window, thereby minimizing bystander editing and allowing successful rescue of the disease phenotype. It is never one-size-fits-all and this study highlights it beautifully.”
Stowaway alert
One limitation of using eVLPs is the inadvertent inclusion of stowaway proteins that are highly expressed in producer cell lines. During production, eVLPs bud out from the cytoplasm of producer cells following co-transfection with the base editor, viral envelop, and guide RNAs. Abundant proteins and RNAs in producer cells may stow away in eVLPs during the budding phase, resulting in undesirable immunogenicity. Studies to ascertain all contents of eVLPs will help evaluate and mitigate immunogenicity.
Kathiresan said, “A principal challenge of VLPs in the therapeutic context is manufacturing, including issues around yield and purity as eVLP producer cells can co-package other unwanted RNAs and proteins.”
“It is likely that similar cellular contents may be co-packaged within eVLPs. Although base editing-eVLP treatment was nontoxic both in vitro and in vivo, additional studies to fully characterize the protein and RNA contents of eVLPs and their effects on transduced cells will help improve our understanding of this delivery system,” the authors noted.
In the current study, the authors demonstrated efficient base editing in the liver and eyes. Improving editing efficiencies in other tissues could further expand the therapeutic potential of eVLPs. In addition, although the researchers demonstrate the transience of the eVLP-transported base editors, further pharmacokinetic studies will help determine dosage and quantify clearance of eVLPs and their cargo.
In closing
An upgrade of clinical approaches from treating superficial symptoms to eliminating the root cause of diseases is now possible through gene-editing tools that can change our DNA code. Base editing, a recent innovation in gene-editing technology, offers the ability to change our DNA code at single-letter resolution without hazardous slashes across both strands of the DNA helix.
Yet, safe, specific, and efficient delivery of the molecular machinery of base editing poses an obstacle between bench studies and its therapeutic application in eliminating once untreatable genetic diseases. Despite the kinks that still need to be ironed out in the new nonviral delivery platform, the current study takes a meticulous and systematic approach in overcoming these hurdles with alacrity.
“The elegant use of the combination of RNPs and VLPs offers a delivery system for transient, tailored, and targeted base editing. This delivery system also overcomes challenges such as antiviral immune response and genomic integration of viral DNA, posed by base editor delivery by viral vectors,” said Jaiswal.
“The broad therapeutic significance of this DNA-free and transient base editing is demonstrated by its use to treat three different organ-specific disease models, which further the impact of this approach. Future work to enhance the delivery of this short-acting base editing therapy in larger models could pave the path for its translation to the clinic,” Jaiswal added.
Harbottle said, “This body of work showcases a novel and effective delivery system for base editing which has a lot of potential in the therapeutic space where safety is paramount. I look forward to seeing further characterization, development, and application of this very promising system!”
“Overall, Liu and colleagues have developed an exciting new advance for the therapeutic delivery of gene editors,” said Kathiresan.






