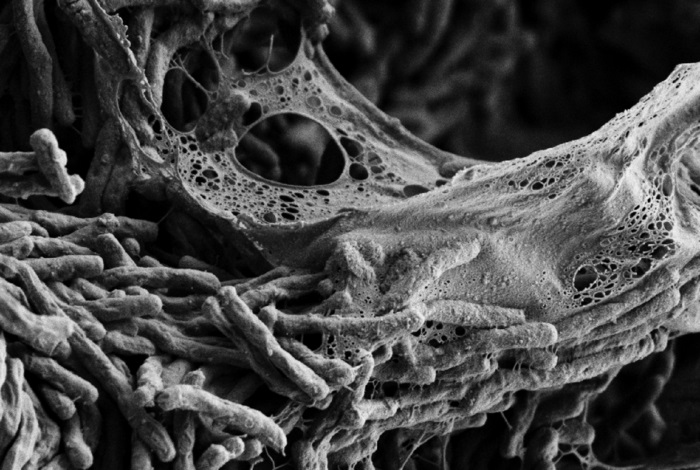
Tuberculosis (TB) is the second leading cause of death due to infectious disease globally. As resistance grows, there is an urgent need for the development of new strategies to treat and diagnose infection. Now, researchers from the University of Wisconsin-Madison and collaborators have identified how the bacterium that causes TB can evolve rapidly in response to new environments. These findings could inform the development of antibiotics targeted at biofilm growth.
Their findings are published in the journal eLife in a paper titled, “Rapid adaptation of a complex trait during experimental evolution of Mycobacterium tuberculosis.”
“TB, caused by Mycobacterium tuberculosis (M. tb), is a leading cause of death due to infectious disease,” the researchers wrote. “TB is not traditionally associated with biofilms, but M. tb biofilms are linked with drug and immune tolerance and there is increasing recognition of their contribution to the recalcitrance of TB infections. Here, we used M. tb experimental evolution to investigate this complex phenotype and identify candidate loci controlling biofilm formation.”
Researchers evolved populations of M. tb in the lab and found that it could form thick biofilms due to mutations in genetic regions.
“TB remains a challenging infection to treat due to the bacterium’s ability to persist in the face of antibiotic and immune pressure, and to acquire novel drug resistances,” explained Madison Youngblom, a graduate student in the lab of senior author Caitlin Pepperell, MD, associate professor, University of Wisconsin-Madison, and co-first author of the study alongside Tracy Smith, science communicator at New York Genome Center. “To better treat and control TB, we need to understand the sources of the bacterium’s robustness and identify its vulnerabilities. We wanted to learn more about how it is able to form biofilms by discovering the genes and genetic regions involved in biofilm growth, as well as how the bacterium evolves in response to changes in its environment.”
Their findings revealed that each strain was able to adapt rapidly to environmental pressure, with the growth of a thicker and therefore more robust biofilm.
The genetic regions that mutated during the experiment, causing this biofilm growth, were mostly regulators such as regX3, phoP, embR, and Rv2488c. “These regulators control the activity of multiple genes, meaning a single mutation can cause many changes to occur in one go,” Youngblom explained. “This is an efficient process that we observed when we looked at the different characteristics of the bacteria, such as their cell size and growth rate.”
“Bacteria are prone to growing as biofilms in many contexts, including the infection of humans and other hosts, and during colonization of natural and built environments,” said Pepperell. “In a medical context, the insights gained from our work could be used to explore potential new antibiotics that are better able to attack bacteria that grow this way. We imagine such biofilm-directed therapies for TB would likely be add-ons to conventional therapy to help shorten and simplify current treatment strategies.”