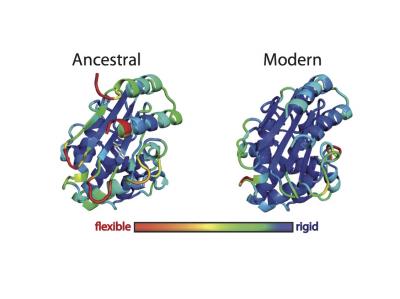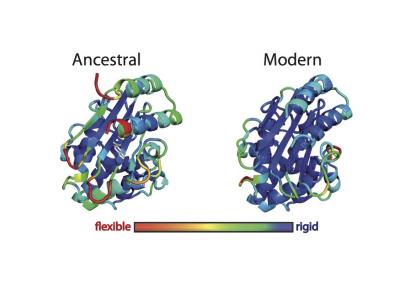
Ancient proteins have been “resurrected” to provide clues that protein engineers could use to generate enzymes that contain new active sites and perform novel functions. In this image, ancestral and modern forms of ß-lactamase are shown in three dimensions. The colors indicate the root mean square fluctuation (RMSF) of each amino acid, which reflects the overall flexibility of the system, as determined by computer simulations. [Jose Manuel Sanchez-Ruiz and Valeria A. Risso/Facultad de Ciencias University of Granada]
Using an experimental setup that could be called Precambrian Park, a team of scientists has brought back long-extinct enzymes that contain clues to the evolution of enzymatic functionalities. No, the team did not include Jurassic Park’s Dr. Ian Malcolm, just a group of biochemists interested in solving a fundamental problem in protein science: How is it that nature has been so good at creating enzymes with diverse functions, whereas rational approaches to protein engineering seem, well, unsatisfactory?
The team, which included members from the University of Granada (Spain), the University of Uppsala (Sweden), the “Instituto de Quimica Fisica Rocasolano” (Madrid, Spain), and the Georgia Institute of Technology (U.S.), did more than resurrect ancient proteins. It revisited an old proposal, forwarded years ago by Roy Jensen, M.D., a scientist currently at the University of Kansas Medical Center. Dr. Jensen suggested that modern enzymes, which tend to be highly specialized, evolved from primordial enzymes that could catalyze a diversity of reactions.
This idea is consistent with the experience of bioengineers. When they apply rational design approaches while trying to generate new active sites, bioengineers usually create proteins that show low levels of catalysis. They have come to believe that completely new active sites are unlikely to emerge in noncatalytic protein scaffolds.
In hopes of taking guidance from nature, the international team of scientists put Dr. Jensen’s idea to the test. The scientists extrapolated backward from known ancestral proteins to arrive at protein designs that likely existed in Precambrian times. Then the scientists subjected synthetic versions of these proteins to structural analysis. The scientists also demonstrated that these proteins could serve as scaffolds for engineering novel enzyme functions.
Detailed results from this work appeared July 18 in the journal Nature Communications, in an article entitled “De Novo Active Sites for Resurrected Precambrian Enzymes.” According to the article’s authors, these results suggest a mechanism for the emergence of primordial enzymes and highlight the potential of ancestral reconstruction as a tool for protein engineering.
“Here, we use resurrected Precambrian proteins as scaffolds for protein engineering and demonstrate that a new active site can be generated through a single hydrophobic-to-ionizable amino acid replacement that generates a partially buried group with perturbed physico-chemical properties,” the authors wrote. “We provide experimental and computational evidence that conformational flexibility can assist the emergence and subsequent evolution of new active sites by improving substrate and transition-state binding, through the sampling of many potentially productive conformations.”
The current study focused on using resurrected Precambrian β-lactamases as scaffolds for the engineering of completely new active sites. Precambrian β-lactamases are proteins approximately 3 billion years old. If we simplify, the scientists did for these proteins what the scientists in Jurassic Park did for dinosaurs: bring ancient forms back to life, so that they can be studied to better understand how complexity in species comes about.
How is it possible to resurrect ancestral protein? Proteins are made from various combinations of amino acid building blocks, with a nearly endless variety of complexity and function. Researchers have compiled large databases full of the proteins sequences. By comparing today's sequences to each other within an evolutionary framework, scientists can reasonably infer the sequence of an ancestral protein from which the modern versions descended using models of sequence evolution.
“The properties of these ancestral proteins (Precambrian β-lactamases) confer high structural stability and promiscuous enzymatic activity, meaning that they are capable of reacting with a variety of substances. These properties support the biotechnological potential of Precambrian protein resurrection because both high stability and enhanced promiscuity are desirable features in protein scaffolds for laboratory directed evolution and molecular design,” explained Valeria A. Risso, a researcher at University of Granada and first author of the paper.
“We have found that a minimalist design to introduce a de novo activity (catalysis of the Kemp elimination, a common benchmark in de novo enzyme design) fails when performed on modern β-lactamases but is highly successful when using the scaffolds of hyperstable/promiscuous Precambrian β-lactamases,” underlined Eric A. Gaucher, Ph.D., from the Institute for Bioengineering and Biosciences, Georgia Institute of Technology.
For their experiment, the team used three structural biology beamlines at the ESRF, the European Synchrotron in Grenoble (France): ID29, ID23-1, and the fully automated “hands-off” beamline MASSIF-1, as well as the Xaloc beamline at Alba, the Spanish synchrotron.
“Three-dimensional structural information derived from the data obtained at the ESRF was essential for the interpretation of the work, as it led to a high-resolution structure of the new active site and provided conclusive evidence of the role of protein reorganization in the emergence of the new function,” noted José A. Gavira, Ph.D., co-corresponding author, from the University of Granada.
The current study confirms the potential of ancestral reconstruction as a tool for protein engineering. “We provide experimental and computational evidence that a laboratory-resurrected ancestral enzyme will make much better scaffolds for new function engineering due to its high stability and dynamics features,” explained José M. Sanchez-Ruiz, Ph.D.
The innovative combination of bioinformatics, computational biology, structural biology, and biophysics allowed researchers to delve deep into evolutionary time and change the course of an enzyme's evolutionary potential. “Learning more about primordial life, and how it can be remanipulated, will open up a lot of new avenues for science, and shed light on the puzzle of how complex biological systems evolve at the most fundamental molecular level,” noted Lynn Kamerlin, Ph.D., co-corresponding author, from the department of cell and molecular biology, Uppsala University.