Circulating tumor DNA (ctDNA) is frequently vastly outnumbered by the amount of non-cancer DNA in the blood, especially in cancers with a low tumor burden, or in certain types of cancer such as gliomas and renal cancers. Improving detection methods is of paramount importance to allow the identification of the early stages of cancer through the process of liquid biopsy. Two recent papers led by the Rosenfeld group at the University of Cambridge report improvements in the detection of cell-free tumor DNA (cftDNA), advancing its potential to revolutionize how cancer is diagnosed and treated. In one paper, the researchers focus on fragment length to improve the specific detection of cftDNA among other cell free DNA (cfDNA). With the other, the researchers target cftDNA in the cerebrospinal fluid (CSF) of glioma patients.
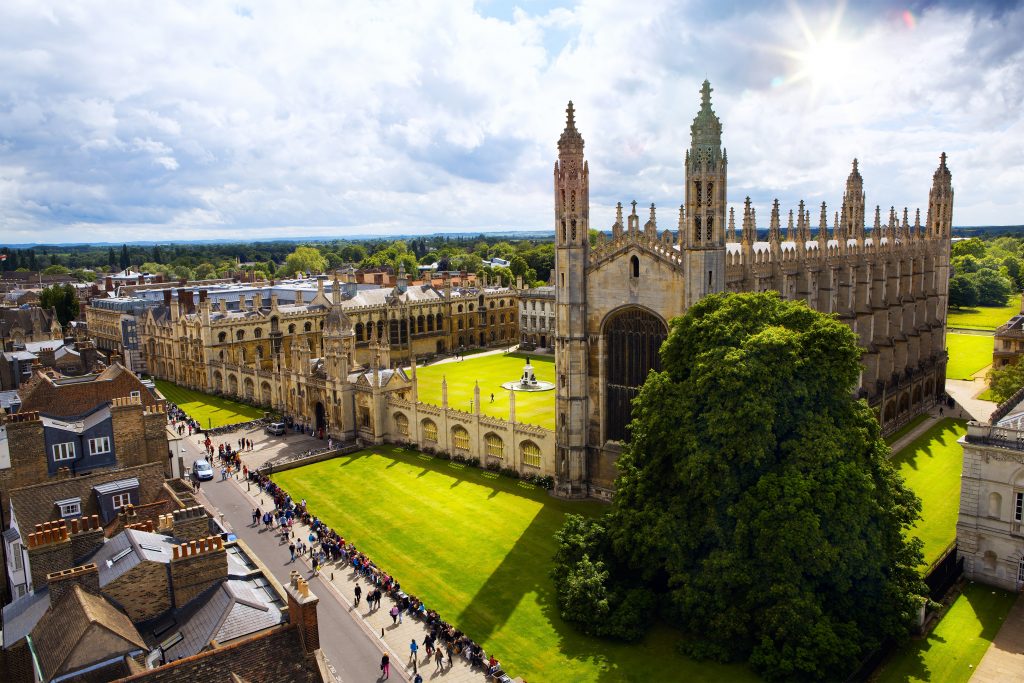

Fragment Size Matters
In healthy individuals, white blood cells are the primary source of cfDNA. There are expected size patterns of cfDNA based on multiple factors, such as the DNA’s availability for cleavage due to interaction with histones. Under certain conditions, such as patients with cancer, other cell sources may contribute to cfDNA in the form of ctDNA. The first paper from this team of researchers suggests that harnessing the distinct size distribution of ctDNA fragments could improve their detection. Published in Science Translational Medicine, the paper is titled “Enhanced detection of circulating tumor DNA by fragment size analysis”.
The authors hypothesized that “differences in fragment length of circulating DNA could be exploited to enhance sensitivity for detecting the presence of ctDNA.” In addition, that ctDNA could be utilized for noninvasive genomics analysis of cancer such as the identification of clinically actionable mutations and somatic copy number alterations (SCNAs).

Florent Mouliere, Ph.D., assistant professor in the department of pathology, Amsterdam UMC, Vrije Universiteit Amsterdam, co-first author on the study, and his colleagues, observed the expected cfDNA size pattern in plasma samples from 65 healthy controls. However, in 344 plasma samples from 200 patients with multiple, different forms of cancer, they noticed varying size distributions. Specifically, they found shorter fragments of 20 to 150 bp more frequently in patients with melanoma, breast, ovarian, lung, colorectal, and cholangiocarcinoma cancer types, but not in plasma samples from other cancers.
Mouliere told Clinical OMICs that the observation of a relative enrichment for ctDNA following size-selection of short fragments was expected. He adds that “our results confirmed this enrichment for ctDNA, and specific alterations and mutations with both in vitro and in silico methods.”
However, Moulier noted that “the results of the second half of the paper were more surprising.” He explained that, “by analyzing and leveraging with a machine learning model the genome-wide fragmentation of cfDNA with a simple and cheap sequencing method, we can drastically enhance the sensitivity for detecting minute amounts of tumor-derived molecules in the blood circulation.” They accomplished this without pre-existing knowledge about the mutations or copy number alterations in the patients. Mouliere added they applied this method to the most difficult cases, gliomas and renal cancer, which are often considered beyond the reach of liquid biopsy. In doing so, they were able to detect the majority of the cases (more 65%) with a plasma sample which Mouliere said was “unprecedented.”
Mouliere added that he would like to try this new concept that they developed to detect tumor-derived DNA by analyzing its genome-wide fragmentation, in other challenging situations such as early detection. “unfortunately, we had no access to early stage samples to demonstrate the potential for these cases, but works are underway to validate this new approach with these samples,” he said.
In a commentary on the work, also published in Science Translational Medicine, Ellen Heitzer, Ph.D., and Michael Speicher, M.D., Institute of Human Genetics, Diagnostic and Research Center for Molecular BioMedicine at the Medical University of Graz, Austria, wrote that the study “substantially extends the armamentarium of cfDNA-based diagnostic tests.” They noted in particular “the possibility of early identification of individuals with cancer offers the potential for new screening strategies and may change the therapeutic outcomes for these patients.”
Heitzer and Speicher detailed the next potentially significant steps for this field. For example, the necessity of confirmation of these results in large, multicenter clinical studies. Also, exploring the differences seen in the methods used for size selection (in silico vs. in vitro) and exploring the potential deleterious effects of size selection.
Cerebrospinal Fluid
In a second publication by Mouliere and colleagues entitled “Detection of cell-free DNA fragmentation and copy number alterations in cerebrospinal fluid from glioma patients,” published in EMBO Molecular Medicine, some members of the same team explored ways to improve the current liquid biopsy approaches used to detect glioma, a relatively common type of brain tumor that originates in the glial cells that surround and support neurons in the brain.
Because the detection rate of cftDNA in the plasma of glioma patients is extremely low, detection in cerebrospinal fluid (CSF) has been an enticing alternative approach. However, detection of cftDNA in CSF comes with many challenges and depends on the tumor’s location and heterogeneity.
Kevin Brindle, Ph.D., professor, department of biochemistry, University of Cambridge, and co-senior author, told Clinical OMICs that “sequencing ctDNA has the potential to guide treatment, but this has been difficult in the case of glioma because of the low levels of ctDNA in the plasma.” He added that “we have described methods that improve the sensitivity of ctDNA detection in the CSF, which bring us closer to the goal of using this approach to guide treatment of this deadly disease.”
The researchers used untargeted shallow whole-genome sequencing (sWGS) of cfDNA from the CSF of 13 glioma patients to identify SCNAs in five of the 13 patients. They also analyzed cfDNA size profiles to show an enrichment in fragments shorter than 145 bp in the patients which correlated with the presence of tumor-derived SCNAs in the CSF.
“The potential for the use of cftDNA in patients with brain tumors is huge.” He noted that he sees patients on a weekly basis whose experience could be improved through the analysis of cftDNA – from diagnosis to treatment planning and monitoring,” said Richard Mair, M.D., Division of Neurosurgery, University of Cambridge, co-first author on the study and a clinical neurosurgeon.
Mair added that, “the main issue has been the difficulty in detecting cftDNA in this group of patients. We have shown that the use of a simple and cheap technique, sWGS, in the cerebrospinal fluid of glioma patients is feasible and, together with fragment size analysis, may enable the selection of patients for more detailed genomic investigation.”
The authors noted that the combination of SCNAs and fragmentation analysis by sWGS of the cfDNA would provide a rapid, low-cost screening method that could provide tumor genomic information. In turn, this could target patients with high levels of cftDNA for larger-scale whole exome or whole-genome sequencing. And, extending the reach of the power of liquid biopsy to more patients with varying types of cancer could yield new diagnostics to advance this game-changing technology.
This article was originally published in the January/February 2019 issue of Clinical OMICs. For more content like this and details on how to get a free subscription, go to www.clinicalomics.com.