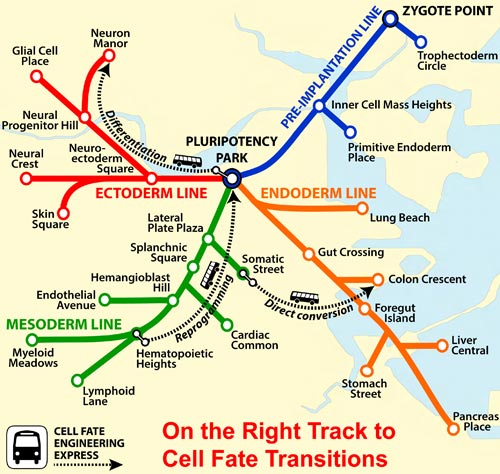
The differentiation of engineered stem cells may be imagined as a subway journey, where the genetic equivalents of missing a transfer or getting off at the wrong stop can take your stem cells far off course. In other words, your stem cells, when they finally detrain, may lack the characteristics you had in mind for them. But what if you had a map?
Such a map has, in fact, been created by scientists from Boston Children’s Hospital, the Wyss Institute, and Boston University. The map, called CellNet, is a computer algorithm, a network biology tool, that compares the gene regulatory networks (GRNs) of engineered cells with those of real-life counterpart cells in the body. With CellNet, researchers may assess:
- the quality of induced pluripotent stem cells (iPS cells) made by reprogramming blood cells or skin cells;
- the quality of specialized cells—such as liver, heart, muscle, brain or blood cells—made from either iPS cells or embryonic stem cells;
- the quality of specialized cells made from other specialized cells (such as liver cells made directly from skin cells);
- what specific improvements need to be made to the engineering process.
CellNet’s development and application have been described in back-to-back papers that appeared August 14 in the journal Cell. One paper, entitled, “CellNet: Network Biology Applied to Stem Cell Engineering,” indicated how CellNet may be used to quantify how closely engineered cell populations resemble their target cell types.
“Analyzing expression data from 56 published reports, we found that cells derived via directed differentiation more closely resemble their in vivo counterparts than products of direct conversion, as reflected by the establishment of target cell-type gene regulatory networks (GRNs),” wrote the authors. “Furthermore, we discovered that directly converted cells fail to adequately silence expression programs of the starting population and that the establishment of unintended GRNs is common to virtually every cellular engineering paradigm.”
The second paper, entitled “Dissecting Engineered Cell Types and Enhancing Cell Fate Conversion via CellNet,” illustrated how CellNet can be employed to improve direct conversion and to uncover unappreciated properties of engineered cells: “Using CellNet, we improved B-cell-to-macrophage conversion, transcriptionally and functionally, by knocking down predicted B-cell regulators.”
“Analyzing conversion of fibroblasts to induced hepatocytes (iHeps), CellNet revealed an unexpected intestinal program regulated by the master regulator Cdx2,” the authors continued. “We observed long-term functional engraftment of mouse colon by iHeps, thereby establishing their broader potential as endoderm progenitors and demonstrating direct conversion of fibroblasts into intestinal epithelium.”
Together, the papers may clarify uncertainty around which methods are best for stem cell engineering, advancing the use of cells derived from patient tissues to model disease, test potential drugs, and deliver treatments.
“To date, there has been no systematic means of assessing the fidelity of cellular engineering—to determine how closely cells made in a petri dish approximate natural tissues in the body,” said George Q. Daley, M.D., Ph.D., director of the stem cell transplantation program at Boston Children’s and senior investigator on both studies. “CellNet was developed to assess the quality of engineered cells and to identify ways to improve their performance.”