Biotechnology in the 21st century is beginning to bear the fruit promised at the beginning of the Human Genome Project over two decades ago. The massive amounts of information derived from that and from subsequent species genome projects have buoyed extensive advances in molecular biology, including gene regulation and epigenetics.
Parallel advances in computing power have dovetailed elegantly with the huge increase in molecular biology information, greatly facilitating researchers’ ability to analyze abundant amounts of data. The synergistic combination of biological information and computing power is not only rapidly advancing our understanding of biological systems, but also ushering in a new era of precision medicine.
Increased genomic data and accelerated computing power helped cultivate the field of spatial biology. Quickly becoming a critical tool for rapidly advancing precision medicine, spatial biology is helping unravel the mysteries of the local cellular environment’s influence on the function and behavior of individual cells.1,2
To unlock the secrets of single-cell and subcellular biology, companies such as NanoString Technologies have developed spatial biology platforms that can characterize molecules down to single-cell and subcellular resolution. Such fine-tuned resolution is crucial for defining baseline and condition-dependent gene expression levels across multiple cell types in healthy and diseased tissues. Broader knowledge of a healthy tissue’s molecular biology allows for reliable identification of relevant differences when the tissue environment changes, particularly in the case of tumors.
Since specific tissue environments depend on the various cell types present, their proximity to each other, and the timing of signals, the ability to gain a more precise understanding of the spatial context of molecular events in intact tissues is critical. Spatial biology provides visualization and quantitation of that context.
Suite of solutions
NanoString’s suite of spatial biology solutions includes the CosMx™ Spatial Molecular Imager (SMI), the GeoMx® Digital Spatial Profiler (DSP), and an integrated analysis tool, the AtoMx™ Spatial Informatics Platform (SIP). Capable of quantifying RNA or protein expression from tissue regions down to subcellular resolution, these technologies increase the capacity of the conventional techniques—in situ hybridization (ISH) and immunofluorescence (IF)—by using automated workflows and enabling high-plex multiomics on FFPE or fresh-frozen tissue samples.
NanoString’s CosMx SMI and GeoMx DSP platforms also have integrated informatics, further shortening the time needed to glean information from precious tissue samples. NanoString’s suite of spatial biology platforms enables scientists to utilize all the information available from every sample.
The newest tool in NanoString’s spatial analysis continuum, the CosMx SMI, provides the greatest level of cellular resolution at the highest plex available. The CosMx SMI expands on the technology of the GeoMx DSP by employing fluorescent molecular barcodes in multiple cycles of nucleic acid hybridization on intact biological samples to measure RNA and protein levels, then overlays single-cell definition and subcellular resolution through use of high-resolution imaging and multianalyte cell segmentation (Figure 1).
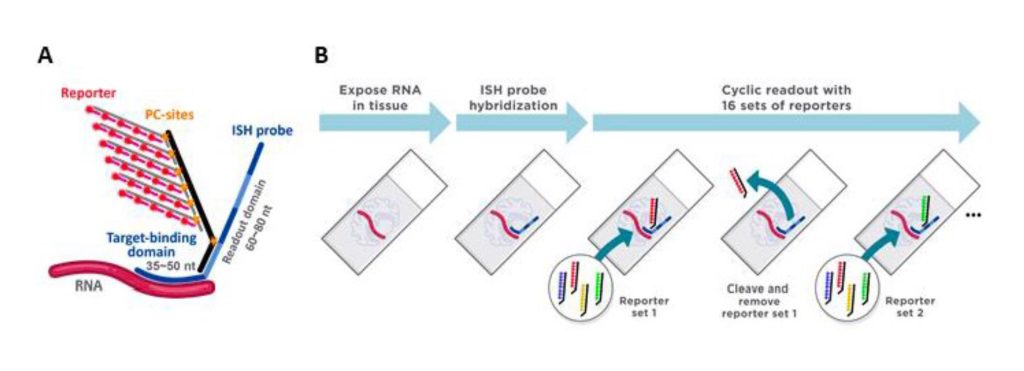
With the ability to analyze gene expression profiles of 1,000 transcripts of RNA and 64-plex proteins in a single experiment, the CosMx SMI is the highest-plex multiomic imager available, ensuring no important biology is missed, even for low-expressing genes.2 Already the most widely adopted spatial imager, the CosMX SMI can classify cell type, cell state, cellular function, and cell-to-cell interaction in a tissue by tracking gene expression changes and ligand-receptor interactions.
Precise cell segmentation by the CosMx SMI is a critical attribute of an in situ analyzer and is achieved by protein imaging and a proprietary deep-learning algorithm.2 The CosMx SMI can pinpoint gene expression in tissues and cells, as demonstrated in non-small cell lung and breast cancers, and it can localize molecular interactions involved in a tissue’s cellular biology.2
Understanding the molecular biology at work in cellular neighborhoods such as the tumor microenvironment will be further accelerated as NanoString continues to refine the CosMx SMI, increasing the assay capacity to ultra-high-plex RNA (approximating whole transcriptome analysis), and further fine-tuning sensitivity and increasing throughput.
Ability to extract voluminous amounts of data
Spatial biology platforms such as the CosMx SMI and the GeoMx DSP can extract voluminous amounts of data from the tissue of choice. To date, massive amounts of data have been a data analysis challenge, forcing scientists either to develop custom data analysis applications on their own, or employ someone who can. NanoString has developed a user-friendly cloud-based informatics solution, known as the AtoMx SIP, for efficiently managing data generated by high-capacity spatial biology platforms. In combination with the CosMx SMI, the AtoMx SIP represents the first fully integrated spatial biology solution designed to go from initial study design to finished peer-reviewed publication.
Accessible from a desktop computer, the AtoMx SIP was developed by NanoString to be user-friendly and have step-by-step support, freeing researchers to concentrate on finding answers in their data instead of spending time developing analytical tools. In addition, the AtoMx SIP is flexible, is able to modify the parameters of a specific experiment for robust data analysis, and has open-source integration, so the research community can collaborate on improving the application with custom code.
Based in the cloud, the AtoMx SIP also facilitates research collaborations with easy data sharing within and across institutions. Initially launching with the CosMx SMI, the AtoMx SIP will also be compatible with the GeoMx DSP, creating a fully integrated suite of spatial biology informatics for analyzing data generated by both platforms.
Anchoring NanoString’s spatial biology platforms is the GeoMx DSP, firmly established as a workhorse for whole transcriptome analysis. Already used widely for large-scale science, the GeoMx DSP is tried and trusted, with more than 375 systems already installed and over 175 publications in the literature. With tunable tissue region of interest selection driven by morphology, the GeoMX DSP is more flexible than fixed arrays and provides a better understanding of the biology in the tissue of interest. In a recent publication,3 the GeoMx DSP was used to identify CD44 in tumor cells as a biomarker for sensitivity to PD-1 axis blockade in non-small cell lung cancer.
Using a spatially resolved proteomic profiling strategy to localize CD44 expression to the tumor compartment but not the immune compartment, an association was established in anti-PD-1-treated patients between CD44 expression and clinical benefit with extended progression-free survival.3 The ability of the GeoMx DSP to analyze RNA and protein on the same slide made it possible to establish this correlation between biomarker and treatment outcome by simultaneously assaying cellular compartments and gene expression levels. Automated to scale, the GeoMx DSP will continue to deliver consistent, reliable results of standardized proteogenomics analysis for precious samples in multisite cancer studies.1
Research and development at NanoString continues to push the envelope by developing innovations in the spatial continuum, including: 1) ever-greater plex capabilities in transcriptome and proteomic capacity; 2) continued refinement of sensitivity to achieve single transcript and protein expression detection; 3) increases in workflow efficiency achieved by decreasing turnaround time without sacrificing reliability; 4) continual increases in the capacity and speed of data analysis informatics capabilities; and 5) integration of artificial intelligence and machine learning capabilities into a suite of spatial biology products—the CosMx SMI, GeoMx DSP, and AtoMx platforms.
To learn more and see our products up close, come visit NanoString at the upcoming Advances in Genome Biology and Technology (AGBT) event. Or visit our website for more information about the CosMx SMI, GeoMx DSP, and AtoMx SIP platforms, as well as our full line of products.
Geoffrey Hummelke, PhD, is a technical writer for NanoString Technologies.
References
1. Bergholtz H, Carter JM, Cesano A, et al. Best Practices for Spatial Profiling for Breast Cancer Research with the GeoMx® Digital Spatial Profiler. Cancers (Basel) 2021; 13(17): 4456. DOI: 10.3390/cancers13174456. DOI: 10.3390/cancers13174456.
2. He S, Bhatt R, Brown C, et al. High-plex imaging of RNA and proteins at subcellular resolution in fixed tissue by spatial molecular imaging. Nat. Biotechnol. Published October 6, 2022. Accessed December 8, 2022. DOI: 10.1038/s41587-022-01483-z.
3. Moutafi MK, Molero M, Martinez Morilla S, et al. Spatially resolved proteomic profiling identifies tumor cell CD44 as a biomarker associated with sensitivity to PD-1 axis blockade in advanced non-small-cell lung cancer. J. Immunother. Cancer 2022; 10(8): e004757. DOI: 10.1136/jitc-2022-004757.

