![This image shows the general transcription factor IID (blue) as it contacts DNA and recruits the polymerase (grey) for gene transcription. The start of the gene is shown with a flash of light. [Eva Nogales/Berkeley Lab]](https://genengnews.com/wp-content/uploads/2018/08/Mar24_2016_BerkeleyLab_TFIID8619637196-1.jpg)
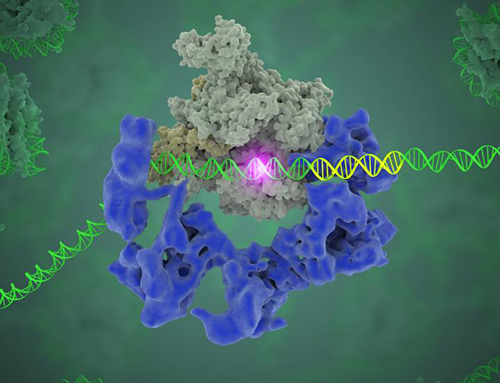
This image shows the general transcription factor IID (blue) as it contacts DNA and recruits the polymerase (grey) for gene transcription. The start of the gene is shown with a flash of light. [Eva Nogales/Berkeley Lab]
The expression of a gene may be likened to a seaborne invasion. Both are logistically complex operations that are hard to follow. Both involve arriving at the right landing site, establishing a beachhead, and only then staging a mechanized advance.
Beyond these generalities are transcriptional details that have been obscured by a biochemical fog of war. Yet a team of researchers based at Lawrence Berkeley National Laboratory has managed to achieve a high degree of situational awareness through the use of cryo-electron microscopy (cryo-EM). Researchers led by Eva Nogales have shown with unprecedented detail the role of the general transcription factor IID (TFIID). In particular, the researchers have observed how TFIID not only finds the right DNA to copy, but also musters the molecular machinery known as the transcription preinitiation complex (PIC). The PIC machinery ultimately positions RNA polymerase II so that it can start transcription proper.
These findings are important because they pave the way for scientists to understand and treat a host of malignancies. “Understanding this regulatory process in the cell is the only way to manipulate it or fix it when it goes bad,” said Dr. Nogales. “Gene expression is at the heart of many essential biological processes, from embryonic development to cancer. One day we'll be able to manipulate these fundamental mechanisms, either to correct for expression of genes that should or should not be present or to take care of malignant states where the process has gone out of control.”
The findings appeared March 23 the journal Nature, in an article titled, “Structure of Promoter-Bound TFIID and Insight into Human Pre-Initiation Complex Assembly.” The article presents the structure of human TFIID in complex with TFIIA and core promoter DNA, determined by single-particle cryo-electron microscopy at subnanometer resolution.
“All core promoter elements are contacted by subunits of TFIID, with TAF1 and TAF2 mediating major interactions with the downstream promoter,” the article’s authors indicated. “TFIIA bridges the TBP–TATA complex with lobe B of TFIID. We also present the cryo-electron microscopy reconstruction of a fully assembled human TAF-less PIC.”
Results such as these suggest how RNA polymerase II, the enzyme that carries out transcription, knows where, along DNA’s immense length, transcription should start. “DNA is a huge, huge molecule. Out of this soup, you have to find where this gene starts, so the polymerase knows where to start copying,” Dr. Nogales explained. “This transcription factor, TFIID, is the protein complex that does exactly that, by recognizing and binding to DNA core promoter regions.”
What Dr. Nogales and her team have been able to do is to visualize, with unprecedented detail, TFIID bound to DNA as it recognizes the start, or promoter, region of a gene. They have also found how it serves as a sort of beachhead for all the molecular machinery that needs to assemble at this position.
“TFIID has to do not only the binding of the DNA, recruitment, and serving as landing pad, it has to somehow do all that differently for different genes at any given point in the life of the organism,” Dr. Nogales added.
“We have generated the first ever structural model of the full human TFIID-based PIC,” declared Robert Louder, a biophysics graduate student in Dr. Nogales’ lab and the article’s first author. “Our model yields novel insights into human PIC assembly, including the role of TFIID in recruiting other components of the PIC to the promoter DNA and how the long observed conformational flexibility of TFIID plays a role in the regulation of transcription initiation.”
Proteins have traditionally been studied using X-ray crystallography, but that technique has not been possible for this kind of research. “TFIID has not been accessible to protein crystallography because there's not enough material to crystallize it, it has very flexible elements, and it is of a huge size,” Dr. Nogales noted. “All of those things we can overcome through cryo-EM.”
Cryo-EM, in which samples are imaged at cryogenic temperatures without need for dyes or fixatives, has been used since the 1980s in structural biology. With extensive computational analysis of the images, researchers are able to obtain three-dimensional structures. However, cryo-EM has undergone a revolution in the last few years with the advent of new detectors—developed, in fact, at Berkeley Lab—that improve resolution and reduce the amount of data needed by up to a hundred-fold.
While this study has revealed important new insights into gene expression, Dr. Nogales states that much work remains to be done. Next she plans to investigate how TFIID is able to recognize different sequences for different gene types and also how it is regulated by cofactors and activators.
“We are just at the beginning,” she remarked. “This complex, TFIID, is very, very critical. Now we have broken barriers in the sense that we can start generating atomic models and get into details of how DNA is being bound.”