Dr. Sankar Ghosh explains how factors outside of protein-coding genes play a role in celiac disease. [Columbia University Medical Center].
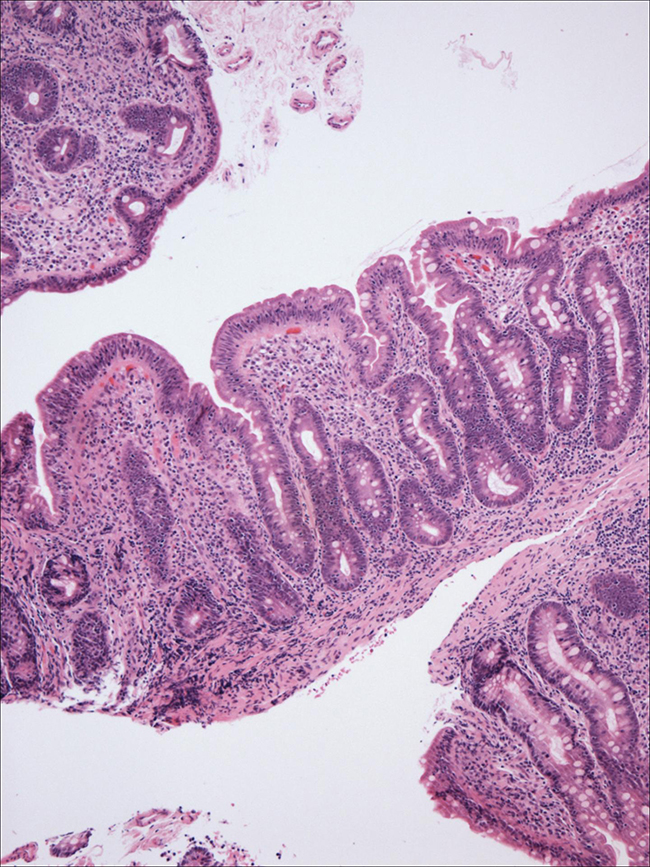
![Stained photomicrographs of small intestinal biopsies from newly diagnosed celiac disease patients showing marked architectural distortion that comprise total villous atrophy, crypt hyperplasia and increased intraepithelial lymphocytes. [Dr. Govind Bhagat, Dept of Pathology and Cell Biology, Columbia University Medical Center]](https://genengnews.com/wp-content/uploads/2018/08/111786_web1098132471-1.jpg)
Stained photomicrographs of small intestinal biopsies from newly diagnosed celiac disease patients showing marked architectural distortion that comprise total villous atrophy, crypt hyperplasia and increased intraepithelial lymphocytes. [Dr. Govind Bhagat, Dept of Pathology and Cell Biology, Columbia University Medical Center]
In the last several years, consumers have become increasingly aware of gluten—a protein found in cereal grains, wheat, rye, and barley—and its association with intestinal inflammation, specifically due to celiac disease (CD), an autoimmune digestive disorder in which genetically predisposed individuals develop an immune response to gluten. While gluten-free products are currently all the rage, researchers are still unsure as to the triggers for CD.
Now, researchers at Columbia University have identified a common variant in a noncoding RNA that may contribute to the intestinal inflammation that occurs in patients with CD. The investigators believe their findings point to a possible new risk factor for developing CD in people with CD risk genes.
Interestingly, while an estimated 40% of the population has the primary gene variant associated with CD, only 1% of people with these genes go on to develop intestinal inflammation and damage after ingesting gluten.
“We don't know why only a fraction of individuals with genetic risk factors for celiac disease actually become gluten intolerant,” explained co-author Peter Green, M.D., professor of medicine at Columbia University Medical Center (CUMC) and director of the Celiac Disease Center at Columbia University. “It is only through the dedicated work of translational scientists that we can begin to uncover the mechanisms that unleash the symptoms of celiac disease.”
The findings from this study were published recently in Science in an article entitled “A Long Noncoding RNA Associated with Susceptibility to Celiac Disease.”
The Columbia scientists focused on the ability of noncoding RNA, the portion of our genome that does not encode for the production of proteins, to regulate a variety of biological processes. Long noncoding RNAs (lncRNAs), which contain more than 200 nucleotides, are thought to play a role in autoimmune diseases and cancers by interacting with other RNA, DNA, and proteins. Yet, it was unclear whether changes in lncRNA genes could lead to the development of more complex diseases.
The researchers were able to demonstrate that a lncRNA variant was able to dampen the expression of celiac-associated genes by binding to a common family of proteins.
“Here we report on the identification and characterization of a lncRNA, lnc13, that harbors a celiac disease–associated haplotype block and represses expression of certain inflammatory genes under homeostatic conditions,” the authors wrote. “Lnc13 regulates gene expression by binding to hnRNPD, a member of a family of ubiquitously expressed heterogeneous nuclear ribonucleoproteins (hnRNPs). Upon stimulation, lnc13 levels are reduced, thereby allowing increased expression of the repressed genes.”
The celiac-associated variant of lnc13 binds poorly to these proteins, leading to increased expression of inflammatory genes. Moreover, the researchers discovered that patients with CD had unusually low levels of lnc13 in their intestines, suggesting that downregulation of this gene may contribute to the inflammation seen in CD.
“These findings add an important detail to our understanding of how celiac disease develops,” noted senior study author Sankar Ghosh, Ph.D., chairman of the department of microbiology and immunology at CUMC. “Given that the majority of the population consumes these grains, understanding the factors that put certain individuals at greater risk for the development of celiac disease will have a broad impact. In future studies, we hope to investigate factors that lead to suppression of lnc13, which may cause celiac disease in people who were previously able to tolerate gluten.”