![A more definite consensus sequence has been determined for the human Initiator, a core promoter element. The human Initiator appears to be located precisely at the start site of more than half of all human genes. [UC San Diego]](https://genengnews.com/wp-content/uploads/2018/08/Jan26_2017_UCSD_DNASequence7211927491-1.jpg)
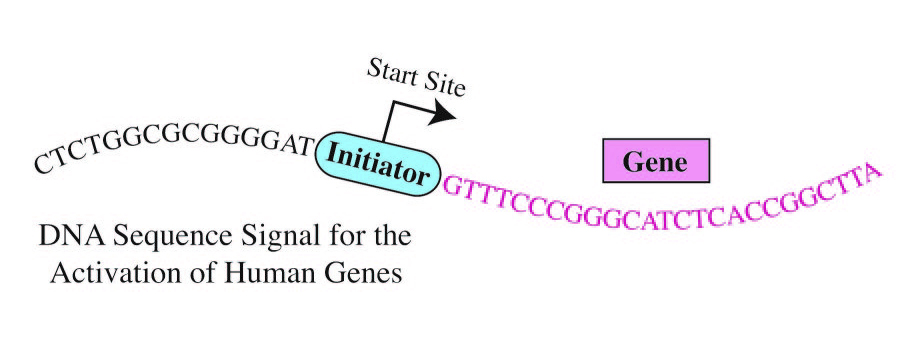
A more definite consensus sequence has been determined for the human Initiator, a core promoter element. The human Initiator appears to be located precisely at the start site of more than half of all human genes. [UC San Diego]
Transcription promoter elements, like history, may not repeat themselves, but they often rhyme. Consider the Initiator (Inr) element, a core promoter that initiates transcription and regulates the activity of more than half of all human genes. Its consensus sequence may be expressed in various ways. For example, in nucleic acid notation, this sequence has been expressed as YYANWYY. Now, as a result of a study conducted by scientists at UC San Diego, it may also be expressed, more precisely, as BBCA+1BW.
This development appeared online January 20 in the journal Genes & Development, in an article entitled, “The Human Initiator Is a Distinct and Abundant Element That Is Precisely Positioned in Focused Core Promoters.” This article, which will appear in print February 10, presents results that could help scientists better understand how human genes are turned on and off.
“Here we show that the human Inr has the consensus of BBCA+1BW at focused promoters in which transcription initiates at a single site or a narrow cluster of sites,” wrote the article’s authors. “The analysis of 7678 focused transcription start sites revealed 40% with a perfect match to the Inr and 16% with a single mismatch outside of the CA+1 core.”
The authors added that TATA-like sequences are underrepresented in Inr promoters. Both the TATA box and the Inr facilitate the binding of transcription factors.
“There are many sequence signals that control gene activity in human cells, and the Inr is the most commonly occurring sequence at the start sites of genes,” said James T. Kadonaga, a professor of molecular biology who led the UC San Diego scientists. “The solution of the human Inr code will enable us to explore new frontiers in gene regulation. In the future, it will be possible to use the code to identify other regulatory signals and, in this way, gain a more complete understanding of how human genes are turned on and off.”
“It is essential for the cell to control the activity of each of its tens of thousands of genes, because the improper control of gene activity can lead to adverse outcomes such as cell death or the formation of a cancer cell.”
That's where the human Inr comes in.
First observed by Pierre Chambon and his colleagues in Strasbourg, France in 1980, the human Inr and its role in gene activation were articulated in 1989 by two MIT biologists, Stephen Smale and David Baltimore at MIT, who revealed in the 1990s the approximate sequence code of the Inr.
Since then, however, other scientists had proposed a number of different sequences for the human Inr, but none of them were found to be consistently associated with the start sites of human genes. As a result, the true Inr sequence code remained a mystery until now.
The new consensus sequence is written in nucleic acid notation, for which the most familiar characters are A, C, G, and T. These characters correspond to the DNA nucleotides adenine, cytosine, guanine, and thymine, respectively. Nucleic acid notation, however, also includes characters that can be used to spell out degenerate sequences, that is, sequences that retain their identity and function despite containing base locations that may be occupied by alternative bases. For example, in nucleic acid notation, Y stands for cytosine or thymine (C/T); N, for any of the four nucleobases (A/C/G/T); W, for adenine or thymine (A/T); and B, for cytosine, guanine, or thymine (C/G/T). The “+1” subscript is used to indicate the specific nucleobase that may serve as a transcription start site.
Kadonaga and his team employed emerging genomic techniques and devised novel computational strategies to unlock the DNA sequence code for the human Inr. They also discovered that this sequence is located precisely at the start site of more than half of all human genes, underlining the importance of the human Inr in the human genome.