The term “integrated circuit” is no longer limited to computer chips, and the term “logic board” is no longer limited to computer chassis. These terms are being applied to biological systems by synthetic biologists. What’s more, the synthetic biologists aren’t being metaphorical. They are being literal. They are starting to assemble biological systems from plug-and-play components. And these systems, like computer systems, are meant to be programmable.
Currently, most synthetic biology components are molecular pathways—switches, oscillators, gated pathways, and so on—that are being used to reengineer, and thus repurpose, living cells. Some repurposed cells are enabling exciting applications in drug discovery and development. Sometimes the cells serve as especially flexible and powerful biopharmaceutical production platforms. Sometimes the cells themselves are the drugs. That is, the cells are “living drugs.”
These possibilities will be discussed at an upcoming conference, the Synthetic Biology–Based Therapeutics Summit. The conference, which will be held virtually December 7–9, has scheduled many impressive speakers, several of whom have contributed their insights to this article.
Most of the insights in this article pertain to tools and platforms that are already being used to develop therapies. However, these therapies are, in a sense, mere preliminaries. The real takeaway is that the new tools and platforms reflect synthetic biology’s commitment to programmability. Accordingly, they may be seen as steps to a new frontier—a place where the bold and ambitious needn’t feel obliged to consider only the easiest, most obvious target that a disease might present. For example, if a disease should present multiple targets, the sensible course might be to hit them all at once, via delivery vehicles that provide exquisite spatiotemporal control over therapeutic administration.
A larger set of CRISPR tools
Mammoth Biosciences searches for novel CRISPR systems by sifting through metagenomics data. According to Lucas Harrington, PhD, Mammoth’s co-founder and CSO, the company has genomics data about microbes from a vast number of sites (including volcanoes, hot springs, and Antarctic ice); from varied specimen types (including animal stools); and from living organisms (including humans). “Mining this massive amount of data,” he says, “feeds new applications.”
Mammoth takes both a top-down and bottom-up approach to find the best tools for each application to leverage the full CRISPR toolbox. The company’s “discovery first” approach addresses how the enzymes work, and the bottom-up perspective defines the specifications of enzymes that address unmet needs, including those in the healthcare space.
For example, Mammoth used a discovery-to-application process to develop the DETECTR system. Mammoth’s scientists discovered that after a certain Cas enzyme complexed with a guide RNA and found a DNA target, it became a nonspecific cutter that could also cleave a reporter molecule. Because reporter molecules can be added to serve as a fluorescent, colorimetric, or electrical signal, a cutting event may provide a real-time result that a target sequence has been found.
Mammoth recently indicated that it was preparing an Emergency Use Authorization (EUA) application for a high-throughput, CRISPR-based COVID-19 test called DETECTR BOOST. The company has other respiratory viral targets in the pipeline. The pandemic illustrated the need to differentiate infectious agents, such as influenza, respiratory syncytial virus, and SARS-CoV-2.
“The power of the CRISPR assay is its flexibility,” stresses Janice Chen, PhD, Mammoth’s co-founder and CTO. “You can program the guide RNA to go after different targets, allowing multiplexing to simultaneously detect different target sequences and get a differential diagnosis from a single sample.”
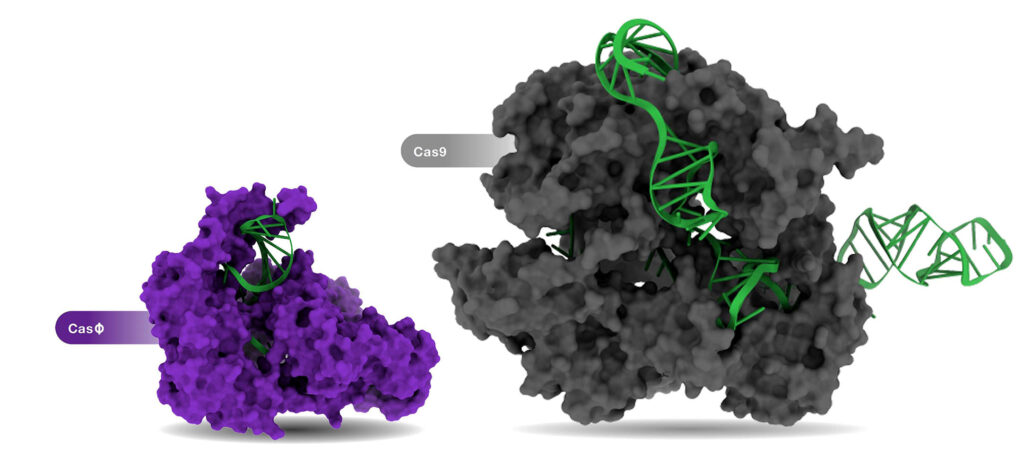
On the therapeutics side, one of the most intense areas of inquiry is the search for compact CRISPR proteins. Legacy CRISPR proteins are very large, and delivery to desired tissues can be difficult.
“Although the systems we are developing have less bulk and machinery than Cas9, we have been able to get higher accuracy due to hundreds of millions of years of natural selection,” Harrington asserts. “We can couple our compact proteins with advanced CRISPR-based techniques, such as base editing, and take advantage of the precision of the edits and the delivery advantages of a smaller protein like Cas14. It is exciting to think about all the different things you can do.”
A discovery engine for programmable cell therapies
“Our technology for integrated circuit–modified T cells allows the installation of more synthetic therapeutic function into T-cell therapies to treat cancers,” says Aaron Cooper, PhD, senior director of synthetic biology at ArsenalBio. “We consider multiple functions that address multiple problems to create a multidimensional cell therapeutic.”
ArsenalBio generates integrated circuit–modified T cells by making use of its PrimeR logic gates, its CARchitecture derived gene expression controls, and its CellFoundry-mediated nonviral manufacturing platform. The integrated circuit, which incorporates multifunctional DNA, can be used to program a T cell to recognize two antigens and to kill tumor cells only when both antigens are present. This approach can overcome targeting difficulties in solid tumors and reduce off-target effects. The integrated circuit can also tune aspects of T-cell biology to increase cellular proliferation and potency, as well as increase resistance to the suppressive tumor microenvironment.
A key challenge is delivering the DNA for the integrated circuit into cells. The integrated circuit is too large for viral delivery. “We have to use a CRISPR-based delivery approach,” Cooper explains. “It can accommodate more DNA and is faster to implement. We can compress iterations and quickly test, learn, and build.”
Another challenge is safety. To address this challenge, ArsenalBio ensures that the DNA is inserted into the genome at a site far from any genes. “Being able to choose a site means we are able to predict the activity of the inserted program more than we could with random integration with a virus or transposon,” Cooper explains. “We can test hypotheses across many T-cell samples and get a high degree of confidence of what we will see in vivo.”
AresenalBio has been using integrated circuits that contain about 8.3 kb of DNA. The large amount of real estate creates a vast design palette to explore.
“The evolution will be in identifying additional mechanisms to go after in a tumor,” Cooper declares. “A lot of ideas exist, and we need to start executing on more than one at a time for a single therapy. Genetic logic boards can allow you to go after multiple facets all at once so that you can tackle antigen recognition, extend the longevity of therapies, and engineer therapies to be more resistant to suppressors that are in the tumor microenvironment.”
A pair of next-generation drug discovery platforms
SyntheX has two drug discovery platforms: ToRPPIDO and ToRNeDO. They rely on functional intracellular drug selection as opposed to in vitro screening. ToRPPIDO allows discovery of compounds that can disrupt specific protein-protein interactions (PPIs) within cells, whereas ToRNeDO achieves the inverse. It discovers compounds that bring together two proteins, an E3 ubiquitin ligase of choice and a neosubstrate of interest, to achieve targeted protein degradation.
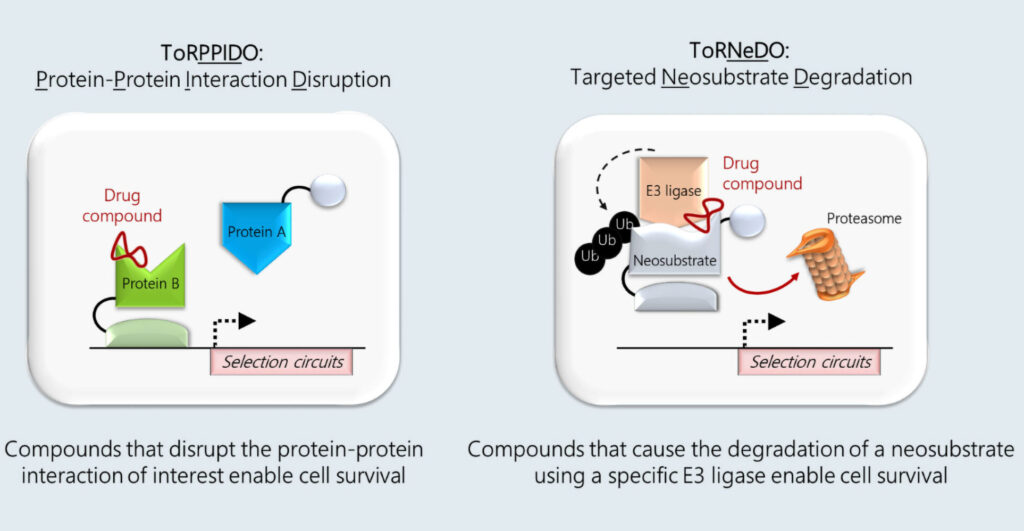
Both technologies rely on the concept of strong and robust negative selection. “In that sense, negative selection is analogous to how a NOR logic gate works,” says Maria Soloveychik, PhD, SyntheX’s co-founder and CEO. “Neither counter-selection nor double-negative reporter circuits can produce a level of negative selection stringency that is robust enough to screen billions of permutations with a very low false-discovery rate. We screen for potent compounds from high-diversity libraries (>1010) with very few off-target hits.
“In addition to the built-in selection circuits, we wanted to expand the chemical space we could sample to achieve the desired functions. For PPI disruption, it is unlikely to find a high-affinity disruptor from a canonical small-molecule library. Therefore, we added a second engineering layer. The same cells that are parsing the selection readouts are also generating highly diversified libraries of genetically encoded molecules (~10 billion) to select from.”
Since PPI interfaces tend to be large, flat surfaces, libraries are focused on peptides and mini-protein scaffolds with diversified variable regions to find hits that can orthosterically block the interaction or to discover new surfaces or pockets that can allosterically interfere with the PPI. This approach lets evolutionary selection reveal the best compounds for producing the desired functional readout in the presence of the engineered selection circuits.
The cellular selection platforms bypass many in vitro bottlenecks. Structural information about the targets of interest is not required, and there is no need to limit constructs to those that can be produced recombinantly and purified in high concentrations. SyntheX identifies molecules that target the proteins of interest while they are expressed in an intracellular milieu and under physiological conditions.
This is particularly useful for ToRNeDO, where the combination of engineered and encoded systems enables the use of a multitude of the more than 600 E3 ligases predicted to be present in the human genome. By screening small-molecule libraries, ToRNeDO can identify functional molecular glues for degraders of a neosubstrate of interest with a desired E3 ligase.
A bacterial platform for vaccine and drug delivery
“Our engineered Clostridium Assisted Drug Development (CADD) platform provides versatility and broad clinical applicability ranging from vaccines to immunotherapies,” says Edward Green, PhD, founder and CEO, CHAIN Biotechnology.
The spore-forming anaerobic host species are well suited for oral delivery in a pill or capsule. The spores survive the highly acidic and proteolytic conditions found in the stomach and upper gastrointestinal tract, and when the spores reach the lower gastrointestinal tract, they germinate to produce viable cells.
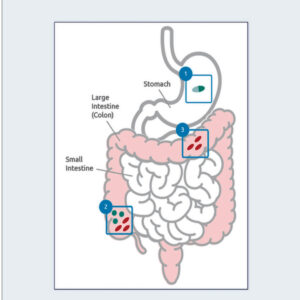
1. Acid-resistant spores are formulated into tablets or capsules for oral dosing.
2. Rapid germination and cell growth commences in the colon.
3. Cell products impact the host (gut mucosa, immune system) and gut microbiota.
CHAIN Biotech produces spores that have been engineered to release vaccines and drugs. So, when these spores reach the lower gastrointestinal tract, they germinate to prophylactic or therapeutic effect. For example, drugs may be released that impact the gut microbiome and mucosa and activate the immune system.
The host bacterial strain engineered by CHAIN Biotech has a long history of being used safely. The strain naturally produces butyrate in the colon, a beneficial short-chain fatty acid supporting growth of a healthy gut microbiome.
The bacterial cells do not permanently colonize. After spore ingestion stops, the bacteria and secreted therapeutics wash out of the colon. Managing the amount and frequency of spore ingestion is primarily used to control dosage. The spore-based biotherapeutics are easy to manufacture and highly stable, and they do not require cold chain logistics for global distribution.
In most cases, precise engineering relates to a single gene and a gene product providing a describable mechanism of action. “For vaccine applications,” Green details, “we design specific antigenic peptides targeting immune cells in the gut mucosa with the potential to stimulate both mucosal and systemic immunity, which most vaccines delivered by injection do not induce.”
CHAIN Biotech has been focusing on oral vaccines to treat or immunize against infectious disease. The company has preclinical projects on human papilloma virus (HPV), human rotavirus (HRV), and SARS-CoV-2.
The HPV vaccine is designed to work therapeutically by clearing chronic HPV infection and preventing cervical cancer. In vivo proof-of-principle data for this vaccine have been generated in collaboration with the University of Oxford. The HRV prophylactic vaccine mainly targets malnourished children in low- and middle-income countries to prevent and treat childhood diarrheal disease. The oral SARS-CoV-2 candidate is designed to offer a long-term, global, and sustainable vaccine solution, especially for emerging variants.
CHAIN Biotech has also developed a therapeutic candidate for treating ulcerative colitis based on secretion of an anti-inflammatory metabolite called β-hydroxybutyrate. “We have proof-of-concept efficacy data in a disease model,” Green points out. “We will partner to clinically advance this candidate.”
A novel RNA reporter system
Circularis Biotechnology has developed a system to discover, enhance, and validate promoters. At the core of the system are highly structured and self-circularizing RNA reporters. The system does not require cellular cofactors to function, and it can work in any cell type.
Next-generation sequencing is used to follow the expression of barcoded RNA circles in massively parallel report assays to examine hundreds to millions of promoters or promoter variants simultaneously. To take advantage of a wide range of genomic sequences, a combination of human, nonhuman primate, and mouse sequences are used to identify promoters.
“We populate the pool of promoters for testing bioinformatically using a combination of databases such as GENCODE to determine the sequences around the transcription start site,” says Paul Feldstein, PhD, founder and CSO, Circularis Biotechnology. “And we use GTEx to find genes that show tissue- or cell-type-specific expression.”
To identify enhancer regions, a library is built that contains genomic DNA fragments that have been cloned upstream of a core promoter of interest with the barcoded reporter downstream. This library is run through cells, the RNA barcodes are read out, and the genomic fragment are associated to increase library members with cell-type specific expression.
“Assay development is challenging because of the dependence on model organisms, such as mice and rats,” Feldstein notes. “These promoter sequences do not necessarily show the same tissue and cell type specificity when put into human or nonhuman primate cells.”
This necessitates using human cell lines as often as possible and cell lines representing a broad range of tissue types to identify promoters with the greatest specificity. Induced pluripotent stem cells or organoid models are valuable alternatives. In an optimal situation, a parallel development track uses a mouse promoter for ease of development and a human or nonhuman primate promoter for clinical trials.
RNA sequencing and quantitative PCR are used to determine if a promoter construct is having off-target effects. If appropriate for the model organism, GFP can better examine a greater range of tissue types.
“We continue to look for improved terminators to better isolate gene expression within the insert from that outside in the genome,” Feldstein says. The Circularis technology can be applied to a variety of gene expression components including the effects on expression of UTRs, terminators, microRNA, and inducible systems.
A microfluidics-based protein engineering platform
A hybrid protein engineering strategy that combines rational design with evolution-based methods can be dramatically enhanced by applying the right technology. “We leverage Rosetta [a flexible computational protein design software] and tools from Bio-Prodict to identify beneficial mutations that enrich our variant libraries,” says Grant Murphy, PhD, director of protein engineering, Merck & Co.
Because Murphy’s group has a range of expression systems—such as Escherichia coli, Bacillus subtilis, Saccharomyces cerevisiae, CHO, and cell-free expression systems—it can engineer proteins that have different levels of complexity and post-translational modifications. Cell-free expression systems are highly scalable (µL to L) and can produce complex proteins for reasonable cost.
The high-throughput systems that build and express variant libraries are highly automated and key to sampling a large sequence space for challenging hosts such as CHO. Almost every variant produced with a high-throughput system is sequenced.
For protein function, a high-throughput, data-rich approach probes protein function in multiple dimensions, such as stability, solubility, and activity. “We leverage a wide variety of instruments to provide a holistic picture of the protein and how engineering is improving the protein along several dimensions,” Murphy notes.
Droplet-based microfluidics has been applied in protein engineering using cell-based or cell-free systems; however, sorting droplets has required the use of fluorescent surrogate substrates. “Using mass-activated droplet sorting (MADS) technology, we have developed an ultra-high-throughput, microfluidics-based platform for protein engineering that directly measures a desired function instead of having to use fluorescent surrogate substrates,” Murphy asserts. “This platform greatly expands the range of engineering projects that we can tackle.”
In addition to enhanced design, expression, and screening capabilities, new machine learning methods leverage the large internal datasets generated with high-throughput technology.
Advances in protein engineering and design have allowed us to make tremendous progress in optimizing protein structures. We can design specifically for stability and solubility, and we can replace immunogenic hot spots. However, our understanding of how structure and dynamics impact function (catalysis and binding) is still very limited.
If good structural information is available, we can use computational design to improve binding. In the absence of good structures, evolution-based methods are relied on with both positive and negative screens.
“The recent progress of protein design, structure prediction, and structural biology methods for correlated protein motions should enable us to better understand the subtle roles of structure and dynamics on function,” Murphy concludes.

