May 15, 2010 (Vol. 30, No. 10)
Double-Quenched Probes Designed to Be Stable at High Temperatures and Minimize Fluorescence
Within the field of molecular biology, quantitative polymerase chain reaction (qPCR) processes enable the simultaneous amplification and quantification of target DNA sequences. As an important experimental procedure, qPCR can be applied to a broad range of different scientific study areas, including drug discovery and medical research. Carried out in three basic phases—denaturing, annealing, and elongation—one PCR experiment typically consists of 20 to 40 cycles in order to obtain significant exponential growth of the target sequence.
As the basis of real-time monitoring, dual-labeled probes (also known as fluorescence-quenched or self-quenching probes) are important tools. Oligonucleotide probes that are complementary to a section of the target DNA can be labeled with a 5´ fluorescent dye and a 3´ quencher.
When in close proximity, the quencher will effectively block the fluorescence signal emission from the fluorophore. However, the 5´ to 3´ exonuclease activity of the DNA polymerase enzyme during the elongation phase of each PCR cycle will digest the 5´ end of the probe, separating it from the quencher, resulting in an increase in fluorescence. The level of fluorescence emitted will, therefore, be directly proportional to the amount of PCR product produced.
Common methods of fluorescent quenching use fluorescent resonance transfer (FRET), which consists of energy transfer via dipolar coupling between the fluorophore and quencher. However, this requires an overlap between the absorption and emission spectra, which complicates the already complex process of probe design, since quenchers are only functional within a limited wavelength range.
Probe Design
The efficiency of quenching via methods such as FRET is extremely sensitive to the distance between the fluorophore and the quencher (RFQ). By increasing the effectiveness of the quenching, background fluorescence is reduced and sensitivity is improved.
Duplex stability and melting temperature (Tm) are also important factors when considering probe design, since each denaturing phase requires that high temperatures are reached (95–96°C) to ensure sufficient separation of the DNA duplex. In subsequent annealing steps, the temperature must be lowered below the Tm to enable target nucleic acids and primers to re-bind. At this point, temperatures must remain high enough to avoid nonspecific hybridization.
The quencher and fluorophore are typically separated by between 20 and 30 bases, which limits quencher efficiency. In order to increase the effectiveness of the probe and decrease the RFQ, shorter oligonucleotides can be synthesized. However, these sequences will only hybridize strongly to complementary DNA at lower temperatures, which are unfavorable for qPCR assays.
Even with the use of Tm-enhancing modifications, such as the inclusion of LNA bases or minor groove binders, probe length is still usually between 14 and 18 bases. Any method that permits the placement of the fluorophore and quencher within closer proximity of one another will improve quencher efficiency and subsequently enhance the probe performance.
For this reason, Integrated DNA Technologies (IDT) has developed a double-quenched probe that is stable at higher temperatures and minimizes the occurrence of background fluorescence, enabling researchers to gain highly precise and sensitive data.
Double-Quenched Probes
A number of factors must be considered when designing a self-quenching probe, including ease of synthesis, fluorophore and quencher compatibility, duplex stability, and probe specificity. The newly developed double-quenched probe incorporates a ZEN™ quencher.
The quencher is placed internally at a fixed distance from the 5´-fluorophore, regardless of probe length, in addition to the standard 3´ quencher (Figure 1). The quencher is inserted into the oligo using a nontraditional, performance-enhancing attachment method that requires a special linker. This, in combination with the unique chemical structure of the ZEN quencher, stabilizes duplex formation (increasing Tm) instead of disrupting duplex formation (decreasing Tm) as normally occurs when nonbase modifying groups interrupt a DNA sequence. Overall, this leads to increased functional performance of the probe.
During the annealing phase of each PCR cycle, the primers and double-quenched probe both bind complementary sections of the DNA. During the elongation phase, polymerization of the new DNA strand is initiated from the primers. Once the polymerase reaches the bound probe, its 5´ to 3´ exonuclease activity degrades the probe, thereby physically separating the quencher from the fluorophore (Figure 1). As a result, fluorescence can be measured and will increase in real-time with the exponential increase in PCR product.
Due to the smaller RFQ achieved with double-quenched probes, background fluorescence is significantly reduced. As such, the initial fluorescence signal measured is much lower, which makes any change in fluorescence easy to detect and functionally increases assay sensitivity.
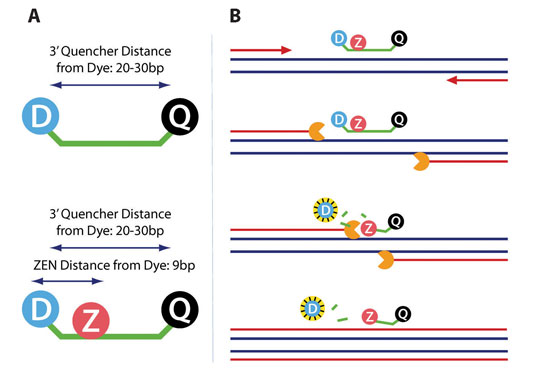
Figure 1. (A) The incorporation of an internal ZEN quencher decreases the distance between quencher and dye to only 9 base pairs. (B) The 5’ to 3’ exonuclease activity of the polymerase degrades the probe, physically separating the quenchers from the fluorophore, which results in measurable fluorescence.
In addition, this design enables longer probes to be used with no loss of sensitivity or quality of quenching because the internal quencher is always the same distance from the fluorophore, regardless of probe length. This is especially useful for AT-rich sequences, where long probes may be needed. In order to test the effectiveness of this double-quencher design, several qPCR reactions were run on a real-time PCR instrument, using a variety of different probes.
Since it is such an important issue to ensure that signal-to-noise ratio remains high, background fluorescence needs to be maintained at a minimal level. Significantly lower background fluorescence is observed when using the new double-quenching probe in comparison to various other dual-labeled probes (Figure 2).
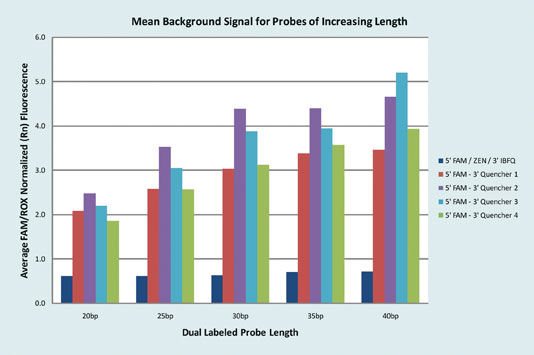
Figure 2. Probes with five different quenchers and five different base pair lengths (20, 25, 30, 35, and 40) were tested for a total of 25 different probe types. Six replicates of each probe type were mixed with 10 ng/µL cDNA and a gene-expression master mix and run under standard cycling conditions. Mean background (Rn) measurements were calculated to determine the average background fluorescent signal.
Precision and accuracy are vital in any experimental set-up. Incorporating double-quenched probes produces extremely accurate (Figure 3) and precise data in comparison to a variety of traditional probes.
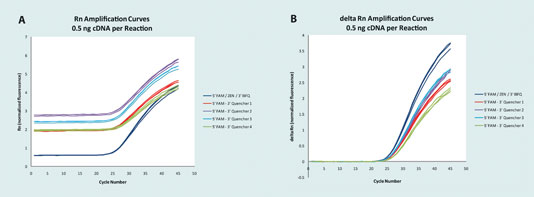
Figure 3 (A and B). 5’ FAM probes with five different quenchers: Each probe was synthesized to target the ACTB locus and all were run in triplicate with the same primer products and 0.5 ng of cDNA.
Conclusion
As a sensitive technique with many uses, it is vital that data obtained from qPCR is as accurate and reliable as possible. IDT’s new double-quenched probe, with the internal ZEN quencher, enables researchers to obtain highly sensitive results due to the significant reduction in background fluorescence.
Furthermore, providing a solution in which the quencher remains close to the fluorophore, regardless of probe length, enables the use of longer probes with a higher Tm, which are able to remain stable at higher temperatures. Researchers can, therefore, be confident in the data obtained from their qPCR.
Mark Behlke, M.D., Ph.D. ([email protected]), is CSO, Scott Rose, Ph.D. ([email protected]), is director, molecular biology, and Jaime Sabel is a technical writer at Integrated DNA Technologies.



