July 1, 2012 (Vol. 32, No. 13)
Quantitative Glycan Analysis for Biomarker Discovery and Characterization of Biotherapeutics
Glycomics is defined as the study of both free sugars and of the complex sugar chains (glycans) that reside in glycoconjugates such as glycoproteins and glycolipids. Given the effects of post-translational glycosylation on protein function, a major focus of glycomics is to characterize the glycosylation patterns on glycoproteins. This glycan analysis has direct relevance to healthcare, both as a source of novel biomarkers and as a means to characterize glycoprotein therapeutics.
The field of investigation for glycan biomarkers is wide open. Only a few glycan markers, the cancer markers CA19-9 and AFP-L3, for example, have so far entered into routine clinical use. Over 50% of proteins in the human body, including most secreted proteins, are glycosylated. The glycans in these conjugates play a role in a wide range of biologic processes including cell adhesion and signaling, and changes in glycoforms have been observed across a wide range of diseases including cancer and neurological, immunological, and metabolic disorders.
Because glycans affect protein stability, binding, and immunogenicity, they are also of critical importance for glycoprotein therapeutics such as monoclonal antibodies. Glycan analysis discriminates among therapies that, on the basis of their peptide constituents alone, might otherwise be considered identical. The extent, type, and location of glycan structures can impact the mechanism of action of these therapies, their clearance rate, their immunogenicity, and their efficacy.
For all the promise of glycomics, this research has traditionally been laborious and time-consuming and thus lagged behind the rapid adoption of genomics and proteomics. Unlike genes and proteins, glycans are not template-determined and adopt a branched, rather than linear, design. And there is no applicable method like PCR or in vitro protein expression systems to amplify glycans.
Recognizing the need to accelerate glycomics research, Ezose Sciences is using its GlycanMap® technology in collaborations with academic institutions and pharmaceutical, biotechnology, and diagnostics companies. The GlycanMap platform is a high-throughput method to deliver glycan data from either purified glycoproteins or crude biological materials. With a current capacity of two 96-well plates per day, which is fully scalable with the addition of more robotics platforms, the GlycanMap platform provides both the throughput and repeatability required for robust biomarker studies and glycoprotein-therapeutics analysis.
Automating Glycan Analysis
The GlycanMap platform combines automated chemoselective, bead-based, glycan enrichment with quantitative MALDI-TOF mass spectrometry coupled to custom bioinformatics. The platform is applicable to analysis of glycans whether they are N-linked (attached to the peptide backbone through an asparagine) or O-linked (attached through a serine or a threonine). It has yielded good reproducibility, linearity, and sensitivity compared to conventional methods in which glycans are fluorescently labeled and detected by high-performance liquid chromatography—but with clear advantages in throughput.
Figure 1 illustrates the GlycanMap workflow. Crude biological samples (serum, cerebrospinal fluid, tissue lysates, cell culture media, etc.) or glycoprotein preparations are denatured, digested with trypsin, and heat-inactivated. The resulting mixture is enzymatically or chemically treated to release the glycans, which are then captured through a bead-based method called glycoblotting. After on-bead washing to remove nonglycan components (peptides, lipids, salts, etc.), labile sialic acid residues are stabilized by methylesterification. Glycans are labeled and released from the beads prior to spotting on a MALDI target plate.
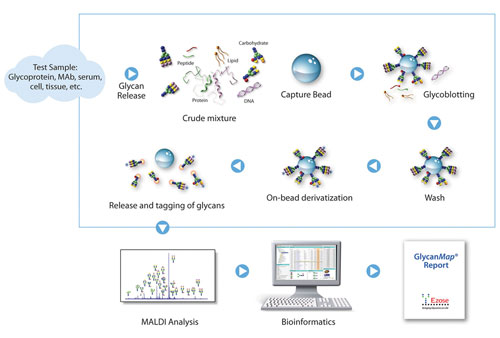
Figure 1. GlycanMap workflow
All steps from initial aliquotting through spotting are integrated into a 96-well robotic system. Following MALDI MS analysis, proprietary bioinformatics programs identify glycan compositions based on molecular weight, and determine glycan quantities by comparing peak heights to internal standards.
The GlycanMap platform has been applied to various biological samples and glycoprotein therapeutics.
In Figure 2, Panel A shows the N-glycan profiles of rat brain lysate, human serum, and human cerebrospinal fluid. Panel B shows the N-glycan profile of three biotherapeutics: the Fc fusion protein Enbrel®, the monoclonal antibody Erbitux®, and the complex glycoprotein Epogen®, which were purchased commercially from a pharmacy for this study.
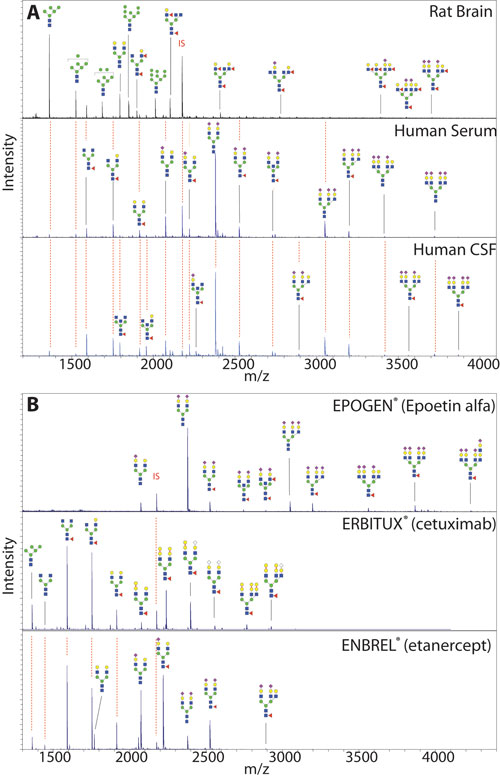
Figure 2. Glycan profiles of biological fluids (Panel A) and biotherapeutics (Panel B). Selected glycan peaks are annotated using the nomenclature of the Consortium for Functional Glycomics. When a glycan was detected in more than one sample type, a red dashed line extends from the annotation to so indicate.
Connecting Glycan Biomarkers to the Underlying Biology
Once glycan patterns have been identified, several complementary approaches can elucidate mechanism and biological relevance. In one approach, parent proteins carrying the glycan modifications of interest are isolated by reverse glycoblotting, in which specific glycopeptides are purified from peptide mixtures by chemoselective enrichment and identified by MS/MS.
Glycan changes can also be analyzed with respect to structural classes and biosynthetic pathways. Figure 3 shows data from a feasibility study where the GlycanMap platform was used to investigate changes in N-glycosylation of cellular proteins accompanying endoplasmic reticulum (ER) stress. Severe ER stress has been implicated in cancer, myocardial infarction, and such neurodegenerative disorders as Parkinson and Alzheimer disease.
A colon cancer cell line (HCT116) was treated for 12 or 36 hours with 0.5 μg/mL Brefeldin A (BFA), a fungal derivative used to study protein transport. An untreated cell line served as the control. The heat map in Panel A compares the relative concentrations of each of the 34 glycans detected in this study in untreated cells and after BFA-treatment.
Panel B projects the BFA-induced glycan changes onto a pathway map of the endoplasmic reticulum and the Golgi apparatus. Each node on the map represents a glycan. The most pronounced concentration changes, indicated by the bright red nodes, were in glycans normally synthesized in the medial-compartment of the Golgi apparatus, suggesting impaired transport between the medial- and trans-Golgi. This observation is consistent with the known BFA-induced fusion of the ER with the cis- and medial-Golgi, but not trans-Golgi, compartments.
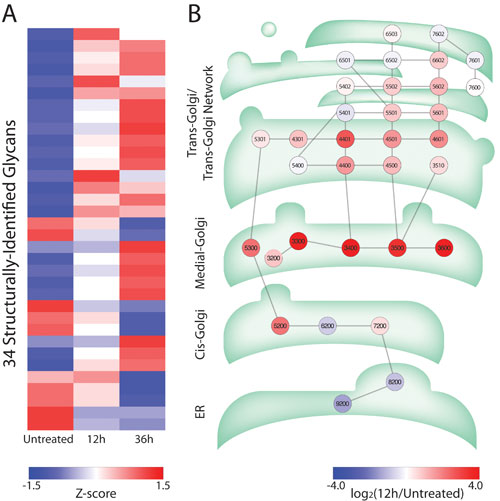
Figure 3. Treatment-induced glycan changes in a human colon cancer cell line: A heat map (Panel A) illustrates relative concentrations of the 34 glycans detected, with blue indicating lower concentrations and red higher concentrations. Panel B shows the projection of glycan changes against the compartments in which each glycan is synthesized: the endoplasmic reticulum and the cis-, medial-, and trans-compartments of the Golgi apparatus. Glycans are identified using a four-digit code, which represents the number of hexoses (Gal, Man, or Glc), N-acetylhexosamines (GlcNAc or GalNAc), deoxyhexoses (Fuc), and N-acetylneuraminic acid (Neu5Ac). [Data shown was generated in collaboration with the National University of Ireland, Galway.]
Moving Forward
The introduction of a practical, high-throughput method of glycan analysis opens many avenues of research—it facilitates an OMICS that until now has been difficult to apply to large-scale studies.
In collaborations with pharmaceutical companies, Ezose has put the GlycanMap platform to use in efforts to discover biomarkers relevant to profiling disease and disease progression and to measuring response to therapeutic interventions. Glycan biomarkers may thus serve as companion diagnostics, too.
Ezose is also partnering with biopharmaceutical companies to apply the GlycanMap platform to characterization of glycan heterogeneity in glycoprotein therapeutic agents. This application of glycan analysis is expanding with the growing interest in biologics, including not only first-generation products but also biosimilars and biobetters.
The glycome remains largely uncharted, and the size of the territory affords many opportunities for exploration.
Yoshiaki Miura, Ph.D., is senior director, research and development, Taku Nakahara, Ph.D., is director, bioinformatics, and Diane McCarthy, Ph.D. ([email protected]), is director, scientific affairs at Ezose Sciences.



