![Researchers have sequenced the human diploid genome without the need for any DNA amplification techniques.[Darryl Leja, NHGRI]](https://genengnews.com/wp-content/uploads/2018/08/June29_2015_Sequencing_DarrylLejaNHGRI1616234223-1.jpg)
Researchers have sequenced the human diploid genome without the need for any DNA amplification techniques.[Darryl Leja, NHGRI]
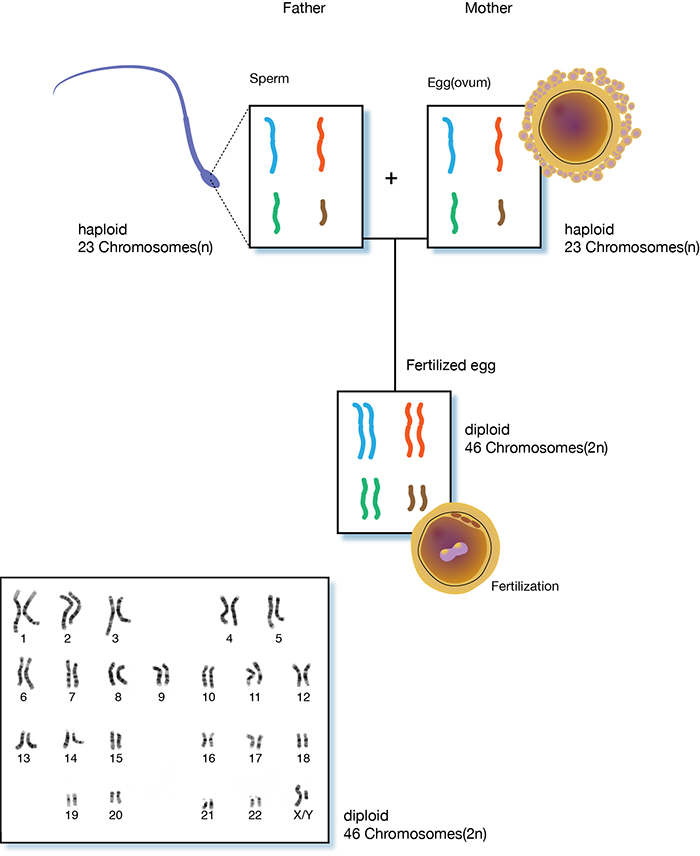
Completion of the human genome sequence in 2003 was a milestone in the biological sciences that can be compared to few other endeavors. However, the project wasn’t without its pitfalls and limitations. In particular, the final assembled sequence, often referred to as the reference genome, is composed of a haploid sequence from its human donor. Since human genomes are diploid, receiving one set of chromosomes from maternal DNA and the other set of paternal DNA, there are many advantages to sequencing the genome in its entirety, simultaneously.
Now, a collaboration of scientists from BioNano Genomics, Pacific Biosciences and led by researchers from Icahn School of Medicine at Mt. Sinai has created a comprehensive analysis of a diploid human genome using two complementary single DNA molecule methods for sequencing and genome mapping. Furthermore, the sequencing was accomplished using long sequencing reads without the need for any DNA amplification techniques—an essential step for all other large-scale sequencing projects that can often introduce replication errors and artifacts into the nascent strand.
The investigators sequenced, mapped, and analyzed a diploid human genome with the goal to integrate single-molecule sequence data and genome mapping data. This approach generated a de novo assembled genome that was reference quality and improved upon the contiguity observed from traditional sequencing methods. Moreover, the combination of BioNano genome mapping and Pacific Biosciences sequencing resulted in an improvement in the contiguity of the initial sequence assembly nearly 30-fold and the initial genome map assembly nearly 8-fold.
“This is the first study demonstrating that our genome mapping technology and single molecule sequencing technology complement each other to generate a reference quality whole genome assembly with haplotype blocks several hundreds of kilobases long,” explained Han Cao, Ph.D., founder and CSO of BioNano Genomics. “This is also the first full de novo assembly of a human genome leveraging intact long native DNA (> 150 kb), without any clone libraries and the artifacts that cloning can introduce.”
The results from this study were released today in Nature Methods through an article entitled “Assembly and Diploid Architecture of an Individual Human Genome via Single Molecule Technologies.”
The researchers’ initial objective was to investigate information often overlooked with sequencing, such as long range repeats and rearrangements, which can be clinically important in complex diseases such as cancer or cardiovascular disease.
Interestingly, as the research team was comparing their newly generated genomic sequence with the current reference genome they found an underrepresentation of lipoprotein A (LPA) gene tandem repeats in the reference sequence. The LPA gene is involved in regulating plasma lipid levels and has been shown to be associated with risk of cardiovascular disease. Quantifying these long 5.6 kb repeats over the span of hundreds of kilobases enables the researchers to assess health risk.
“Many large and complex forms of variation are missed by traditional next generation sequencing approaches,” said Ali Bashir, Ph.D. assistant professor of genetics and genomics at Mt. Sinai and senior author of the study. “Combining long read sequencing and BioNano genome mapping produces highly contiguous de novo assemblies, enabling unbiased comparison of nearly complete genomes—something we have been trying to do for years.”