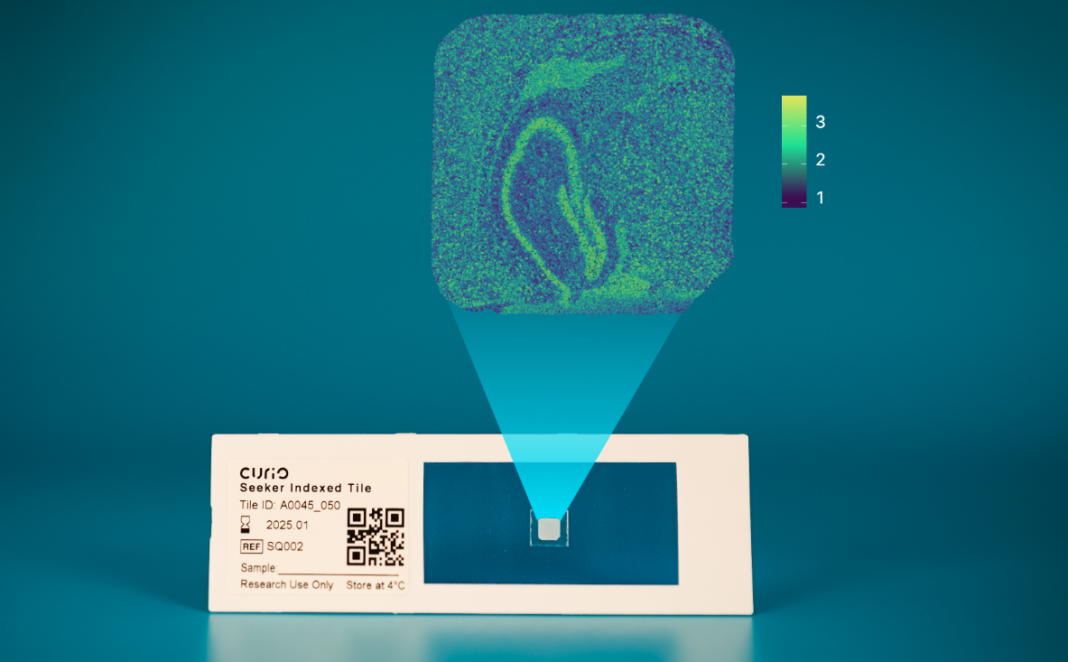
In 2019, spatial transcriptomics was creating a lot of buzz in the genomics world. Technologies like MERFISH and FISSEQ were the foundation of new companies. That same year, Fei Chen, PhD, Sam Rodriques, PhD, and Evan Macosko, MD, PhD, colleagues at the Broad Institute, published a new spatial technology that doesn’t need any equipment, other than a sequencer, called Slide-seq. In addition, Slide-seq offers high resolution due to the platform’s minute (10 microns) barcoded beads and limited diffusion.
Immediately after the paper was published, calls and requests from collaborators came rolling in. Chen told GEN that they were overwhelmed supporting other folks, some of whom were far outside of Chen’s core scientific focus.
Chen’s goal was to get the tool into the hands of other researchers. The impact of developing technologies, he says, is when people use them. But they needed a company to do it. They tried to do it themselves and sought interest from venture capital firms. But selling tools to researchers is a somewhat non-traditional business model for a VC. They were more interested in using it internally to discover drugs. And that wasn’t what Chen had in mind.
Then the pandemic hit, other priorities took over, and Slide-seq stayed in the lab. All the while, Chen’s group continued to work on Slide-seq, making improvements along the way.
In early 2021, Chen, Macosko, and Rodriques were connected to Steve Fodor, PhD, genomics pioneer and co-founder of Affymetrix, through a VC incubator. Chen called it a “very chance meeting.” Fodor liked the technology and liked the idea of distributing it to the world.
The partnership, which includes co-founders Ari Chaney (COO) and Christina Fan, PhD (CTO), was formed in early 2021 and today, less than two years later, Curio Bioscience is launching the Slide-seq technology under a new name, Seeker. The kit includes eight Seeker tiles (which are pucks in Slide-seq), reagents, and text files with bead barcodes and spatial locations.
Chen said that, in retrospect, 2019 probably would have been a little early to the game. The additional two years meant that the technology matured to a place where it was easy to manufacture and turn into a tool. And the excitement for spatial grew considerably over those two years. All this probably served the technology’s developers well in the end.
Two other non-imaging-based spatial methods have been developed. The highest resolution is Stereo-seq technology, developed by BGI-Research, which is not commercially available yet. One complication with Stereo-seq is that it requires an MGI sequencer. The other technology is 10X Genomics’s Visium—the most widely used spatial technology currently.
Chen is excited to see Slide-seq launching as a product, in particular for researchers in the single-cell community who want to ask questions about tissue biology. Another key group are researchers working in the tumor microenvironment, developmental biology, and non-model organisms—an area that is not as heavily focused on but is uniquely enabling.
Weird critters
“We work on weird critters,” said Anoja Perera, director, sequencing and discovery genomics at the Stowers Institute for Medical Research in Kansas City. Instead of human samples, they have model organisms like planaria and axolotls. “You name it,” said Perera. When they started using spatial technologies, about three years ago, they used Slide-seq (in collaboration with Chen) and Visium from 10X Genomics.
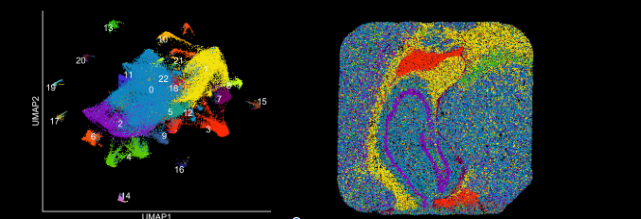
That said, they are considering all the spatial platforms in her facility. For example, they are currently testing samples using stereo-seq. The in situ-based spatial techniques are, Perera said, less applicable to the work at Stowers because they are geared to human and mouse. In addition, she likes the flexibility of Curio. One day, she can do an experiment on zebrafish, another she can do planaria. To use Seeker, all she needs is for the organism to have a genome sequence. And she doesn’t have to worry about highly expressed genes saturating the images. But she added that, as with any small company, she worries about the production—will they be able to keep up?
Because of their familiarity with Slide-seq, Perera said moving to Seeker was an easy transition. But in a lab that is not as familiar, she thinks that Curio may need to visit customer labs and do more training. Spatial is almost an art, she said; you need a great histology group and a skilled molecular biology group. If there is a good histology lab and someone who is skilled in molecular biology protocols, she said, someone who has never done spatial before could use Seeker.
Back to humans
Angela Pisco, PhD, associate director of data science, quantitative cell science at the Chan Zuckerberg Biohub, leads a project to build a reference atlas. The first version of Tabula-sapians, which was built on dissociated single cells—half a million cells in 23 tissues—was published last year.
Although the project started with dissociated samples, Pisco and colleagues have been obtaining spatial data since the beginning of 2021. They have been using Vizgen’s platform, the Merscope, but jumped at the chance to use the Seeker, primarily because they were interested in the unbiased approach it offered. Now, they are about to embark on building a reference using Curio. Pisco said that Curio’s signal-to-noise ratio is a key reason why she is using it. The quality of the signal is a priority because, she says, a reference that is going to be used by the community must be as high quality as possible.
One downside is that the current version of the Seeker works only on fresh frozen samples, not formalin-fixed paraffin-embedded (FFPE) samples. Moreover, unlike some of the other platforms, there is no point at which a picture of the tissue is taken. So, there is no way to recover the morphology, thus making a comparison to an H&E stained image, which may be important in a clinical setting, very difficult. Curio may need to reconcile the lack of a way to image the tissue. Without it, researchers need to place a lot of trust in the technology. And some scientists may be less willing to do that than others.
The only option
Kelvin Chen, PhD, is a researcher in Shimon Sakaguchi’s lab at Osaka University—an immunology lab that relies heavily on single-cell RNA-seq and ATAC data to profile human tissue samples. Spatial would be very useful to profile cell-to-cell interactions. But the previous spatial technologies were not options for Chen. Either the resolution was too low, or they were prohibitively expensive.
Although Chen wanted to use Slide-seq, he didn’t know how to access it. He doesn’t know Chen, Macosko, or Rodriques. But when Bertrand Yeung, PhD, director of business development at Curio, reached out to him about the Seeker, he was eager to try it.
Kelvin Chen said he finds the Seeker easy to use, but quickly added that he has a lot of experience in this type of work, especially in library prep. As an immunology lab, the histology has been the biggest challenge for him. And, he added, the tiles are small, which is tricky. It’s difficult to mount a large sample onto a small tile. And “it’s computationally heavy.” Indeed, the cell resolution was so high, he said, that the department’s supercomputer ran out of memory.
Fei Chen is excited to see what people do with the Seeker. It’s “a little bit into the unknown.” He’s also looking forward to redirecting the ongoing requests for Slide-seq to a company. As for himself, he’ll continue making arrays, and building on top of the Slide-seq technology, in his lab. And he’ll likely become a customer of Curio’s as well.
But he might have to get in line.