Doug Auld, Ph.D. Novartis Institutes for BioMedical Research
CRISPR Slips into LACE for Better Light
The CRISPR-Cas9 system continues to show rapid expansion in applications to gene editing and control of transcription. This paper couples the CRISPR-Cas9 complex to light-activated domains (so-called LACE system) to control gene expression dynamically by exposing cells to blue light. Light activated domains from Arabidopsis were used, which included cryptochrome (CRY) domains that act as blue-light receptors, which can modulate gene expression through binding to a cryptochrome-interacting basic-helix-loop-helix protein (CIB1). In the LACE system, a nuclease inactive form of Cas9 (dCas9) was fused at either its N- or C-terminal to CIB1, and a CRY2-VP6 fusion was co-expressed along with four guide RNAs to target IL1RN promoter (figure). Exposure of cells to blue light showed expression of IL1RN mRNA in HEK293 cells (figure). The LACE system is reversible upon removal of blue-light exposure, and the authors show that the system is dynamic, as one can cycle gene expression on or off on successive days by intermittent blue-light exposure. One can also spatially activate gene expression using a photomask. The LACE system described in this paper should be a useful tool in cell engineering providing both spatial and temporal control.
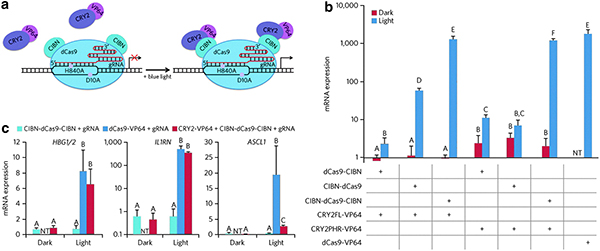
Light-inducible, RNA-guided activation of endogenous human genes by the LACE system. (a) When expressed by cells, CIBN-dCas9-CIBN localizes to the DNA sequence targeted by the gRNA. In the presence of blue light, CRY2 undergoes a conformational change that enables heterodimerization with CIBN, which causes translocation of CRY2FL-VP64 to the targeted DNA sequence and transcriptional activation of the downstream gene. (b) IL1RN activation in HEK293T cells using four targeting gRNAs; either CIBN-dCas9, dCas9-CIBN, or CIBN-dCas9-CIBN; and either CRY2FL-VP64 or CRY2PHR-VP64. Red bars, dark-incubated cells. Blue bars, cells illuminated with pulsing blue light (1 sec pulses at 0.067 Hz). Conditions not marked by the same letter gave results that were significantly different (P<0.05) as determined by ANOVA and Tukey's test. (c) Activation of multiple endogenous gene targets was achieved using different groups of gRNAs targeted to HBG1/2, IL1RN, or ASCL1 in HEK293T cells. Activation levels in illuminated cells that contained the LACE system were statistically similar to those in cells that expressed dCas9-VP64 and the same gRNAs when HBG1/2 or IL1RN were targeted. Conditions not marked by the same letter are significantly different (P<0.01) as determined by global ANOVA and Tukey's test. For b,c: NT, not tested. Data show combined replicates from two independent experiments (n=4±SD).
* Abstract from Nat Chem Biol 2015; Vol. 11:198–201
Optogenetic systems enable precise spatial and temporal control of cell behavior. We engineered a light-activated CRISPR-Cas9 effector (LACE) system that induces transcription of endogenous genes in the presence of blue light. This was accomplished by fusing the light-inducible heterodimerizing proteins CRY2 and CIB1 to a transactivation domain and the catalytically inactive dCas9, respectively. The versatile LACE system can be easily directed to new DNA sequences for the dynamic regulation of endogenous genes.
Doug Auld, Ph.D., is affiliated with the Novartis Institutes for BioMedical Research.
ASSAY & Drug Development Technologies, published by Mary Ann Liebert, Inc., offers a unique combination of original research and reports on the techniques and tools being used in cutting-edge drug development. The journal includes a "Literature Search and Review" column that identifies published papers of note and discusses their importance. GEN presents here one article that was analyzed in the "Literature Search and Review" column, a paper published in Nat Chem Bio titled "A light-inducible CRISPR-Cas9 system for control of endogenous gene activation." Authors of the paper are Polstein LR, Gersbach CA.



