Circulating tumor cells (CTCs) are cells that detach from primary or secondary tumors and enter the circulatory system. As such, they show promise as a noninvasive liquid biopsy technique. CTCs have been validated as prognostic and predictive biomarkers in many different cancers including breast, lung, prostate, and colorectal cancer. A number of technologies are now available to count, capture, and expand CTCs, and these methods are being developed to help predict and treat cancer patients. Through study of CTCs, the mechanisms of metastasis can be understood, and in some cases, the occurrence of metastasis can be predicted. Some of the newest developments in CTC research were presented at the Molecular Med Tri-Con event held recently in San Francisco.
Reproducible, deployable, density-based separation
With increasing interest in analyzing CTCs in a clinical setting, especially for monitoring studies in which it is impractical to routinely obtain serial biopsies from patients, researchers are looking to platform technologies that enable reliable CTC isolation, identification, and analysis. Such technologies must address the fact that CTCs are heterogeneous and extremely rare.
“You have a challenge [figuring out how to] detect CTCs with sensitivity and specificity in a reproducible manner,” said Tad George, PhD, vice president, scientific applications, RareCyte. His presentation, “An Open, End-to-End, and Flexible Platform for Scalable CTC Collection, Identification, and Analysis,” detailed how RareCyte’s platform enables a complete workflow for CTC-based liquid biopsy from blood draw to CTC enumeration and biomarker expression to single-cell isolation.
The RareCyte process begins when nucleated cells from a liquid biopsy are transferred to slides using a density-based sample preparation system. The slides can be banked for future studies, or stained with immunofluorescence panels designed to enable CTC identification and biomarker expression analysis. They are processed with six-channel imaging instruments with integrated machine learning (ML) algorithms to present rank-scored CTC candidates to a reviewer. The rank scoring system is informed by over 500 image-based parameters and thousands of clinical CTCs. Finally, individual CTCs can be retrieved for downstream molecular analysis with the integrated cell retrieval module.
According to George, one key advantage of RareCyte’s density-based approach to cell isolation is that it does not bias for or against isolation of CTCs on the basis of size or protein expression. This potentiates maximum sensitivity since CTCs are heterogeneous in both size and protein expression. The ability to sensitively and specifically detect CTCs is therefore dependent on the staining panel. Those include markers used for CTC identification, including antibodies to cytokeratin and EpCAM, as well as CD45 as an exclusionary marker. Additional markers can be added to identify other types of CTC or to measure CTC biomarker expression.
The Rarecyte platform includes a number of innovations that enhance reproducibility. First, its sample preparation method minimizes loss of CTCs. Loss of CTCs can lead to unreliable and unreproducible results. It also does not discriminate based on biomarker expression, cell compressibility, or cell size. As well, its slide-spreading process and ML algorithms contribute to reproducibility. Finally, the entire system is deployable, meaning that it can be set up in the customer’s lab, as opposed to being an off-site service that processes customer samples.
Understanding the mechanisms of metastasis
CTCs are implicated in the formation of metastases. One particular area of interest is the mechanisms by which cancer spreads through CTCs to different organs. Min Yu, PhD, assistant professor of stem biology and regenerative medicine at the University of Southern California, delivered a talk, “Patient-Derived Circulating Tumor Cells Inform Mechanisms of Metastasis,” describing her group’s progress in understanding metastasis.
“We were particularly interested in CTCs because they can be easily sampled compared to invasive biopsies,” said Yu. Using a mouse xenograft model with genetic studies to pin down how CTCs initiate brain metastasis, her group found that at least two different programs are required for CTCs to form brain metastases. One program helps the cells cross the blood-brain barrier, and another program allows the cells to resist the defenses of the local microenvironment to establish growth in the brain.
Using RNA sequencing, Yu compared metastases formed in the brain, lung, and bone to parental cells to identify genes highly expressed in brain metastasis. That allowed Yu’s team to narrow the field down to two gene candidates: a cell surface receptor that helps bind CTCs to endothelial cells, and a transcription factor that helps CTCs colonize the brain.
Yu said that these genes could be used to predict patient response to treatment, or the course of disease. “There’s still a lot more basic understanding we need to gain for patient-derived CTCs,” Yu suggested. “Only if we understand how they generate metastases can we design a better predictive measure or target therapies for the patient.”
Role of CTCs in immune system blockade
In his presentation, “Assessing PD-L1 Expression on CTCs and Correlation with Immunotherapy Response,” Rajan Kulkarni, MD, PhD, assistant professor, Oregon Health and Science University, discussed how CTCs can be used as a screening tool to supplement other testing methods in developing inhibitors for an emerging class of cancer therapy candidates. The immune system has a number of mechanisms for protecting the body from cancer. Aggressive tumors must evade those protections to grow and spread. One pathway that the immune system uses against cancer is the PD-1/PD-L1 checkpoint. The PD-1 receptor on the surface of activated T cells limits the development of a T-cell response when bound by the PD-L1 ligand. This ensures that the immune system does not mount an inappropriate response leading to harmful autoimmune inflammation.
Kulkarni’s lab developed an assay to quantify PD-L1 expression on CTCs using the Vortex chip. To separate CTCs based on size, the chip uses microfluidic channels that generate vortices at certain flow rates. The CTCs are then stained with an anti-PD-L1 antibody.
Kulkarni’s group tested the assay in a group of patients, comparing this group’s data with biopsy results where available. In patients with tissue biopsy results, Kulkarni said, there was “relatively good concordance of PD-L1 expression on CTCs compared to the tumor,” but it was in only four to five samples total.
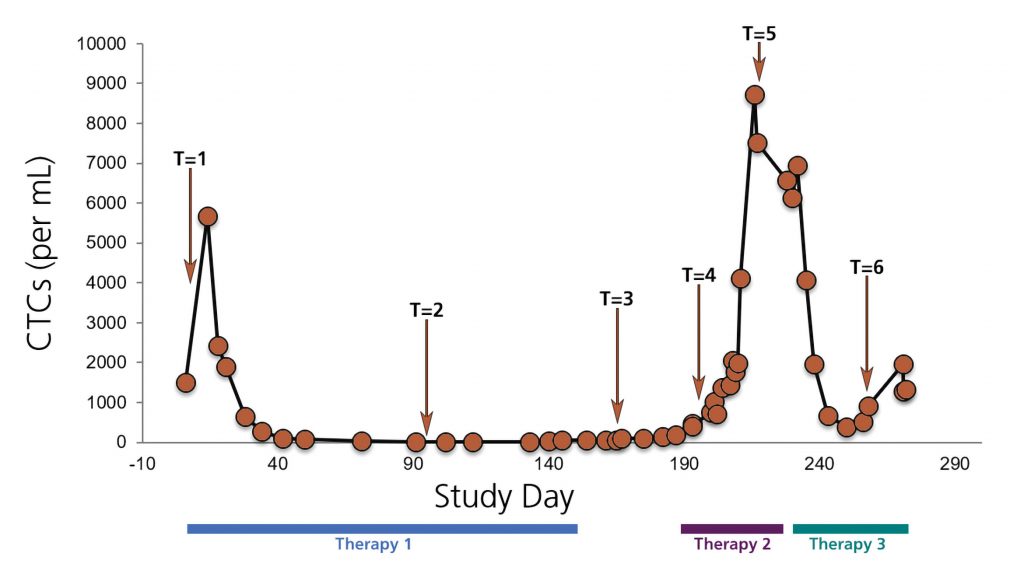
His group also assessed PD-L1 expression on CTCs prior to the first treatment cycle, and followed those patients over time to measure their response. In those patients—about 20 altogether—Kulkarni said there was a trend toward response, but there were not enough patients for statistical significance. Another finding was that the number of CTCs prior to starting immunotherapy was also correlated with response to therapy. “Those with more CTCs had better progression-free survival,” noted Kulkarni.
Many applications of acoustic microstreaming
Acoustic microstreaming is a method of focusing, sorting, and enriching CTCs within a microchannel. It works through oscillation of bubble surfaces by an acoustic energy source that creates a streaming flow in the channel. The instrument uses lateral cavity acoustic transducers (LCATs) to separate whole blood into its cellular constituents. It enriches rare cells based on size and enables fluorescent labeling of target cells. Cells can therefore be identified by size and surface biomarkers in a single device.
In his presentation, “From Sample to Single Cells to Answer: An Integrated Microfluidic Approach for Identifying Cancer Cells,” Abraham Lee, PhD, professor, mechanical and aerospace engineering, University of California, Irvine, described how a platform capable of multiplex analysis can be a powerful analytic tool: “I can take millions of cells and separate down to thousands of cells. And thousands of cells can be trapped into single-cell trapping arrays.”
Identification of critical subpopulations of cells is an important bottleneck when isolating and targeting CTCs in a liquid biopsy. To understand cellular heterogeneity, it is necessary to separate target cells from unwanted and contaminated cells. An integrated microfluidics platform provides a bridge from liquid biopsy to clinical applications.
Lee has investigated techniques for identifying cancer cells in a liquid biopsy, using both phenotype and genotype methods. One is a collaboration with Michelle Digman, PhD, assistant professor, department of biomedical engineering, UCI, using fluorescence lifetime imaging microscopy (FLIM) to measure changes between free and bound fractions of NADH to determine if a cell is in a state of oxidative phosphorylation or glycolysis. Lee maintained that metabolic state can indicate if cells are healthy or cancerous.
In another approach, Lee collaborates with UCI’s Kumar Wickramasinghe, PhD, chair, electrical engineering and computer science, to nondestructively probe separated single cells by dielectric nanotweezers, which are modified by AFM tips to extract mRNA. The mRNA is tested for genotypes linked to cancer to identify CTCs from healthy white blood cells.
Using the same platform, Lee can also analyze the antibody spectrum of cells. “We’re using these single-cell traps to trap double cells or different types of cells and putting them together,” explained Lee. “There’s a lot of these techniques that are originally distinct and separate from each other. Now with the microfluidic platform, they can be fused together and cross-fertilized so we understand the heterogeneity of the liquid biopsy sample and how it changes over time.”