Surface plasmon resonance, or SPR, is used to measure changes in molecular weight. Thus, it has long been used to study protein-protein interactions by registering the association and dissociation of a ligand (prey) binding to an immobilized protein (bait).
SPR has recently been revitalized by its aptitude for fragment-based lead discovery. Fragments are low-molecular-weight compounds (<350 kD) that bind to targets with low affinity (µM–mM range), yet they exhibit high ligand efficiency—each atom contributes by directly contacting the target’s binding site.
Once a hit is identified, the fragment is grown or combined with other fragments, usually by structure-guided design and synthesis, to quickly generate a much more potent lead. Fragments are simple, but libraries composed of them can represent many diverse compounds and different chemistries. It is hoped that discovering drugs in this piecemeal manner will shortly yield therapeutic breakthroughs.
Biacore (now part of GE Healthcare) claims to be the first company to apply SPR to label-free protein-protein interaction analysis with its Biacore™ systems, according to Stefan Lofas, Ph.D., principle R&D scientist. The system’s various applications venture beyond protein-protein interactions to include protein-DNA interactions, peptide interactions, the binding properties of antibodies, drugs, and small molecules, and even interaction of viruses with whole cells, he adds.
Biacore makes different machines for different purposes, including screening and drug discovery. Moreover, GE Healthcare recently acquired Microcal, a market leader in calorimetry, whose instruments can detect the number of binding sites and measure the enthalpy (DH) and entropy (DS) of binding. This is a “perfect complementary technique to Biacore systems,” Dr. Lofas explains.
Tony Giannetti, Ph.D., research scientist at Genentech, uses Biacore systems to identify promiscuous binders and remove them from high-throughput screens where they would create false positives. This saves valuable time, money, and effort that might otherwise be spent crystallizing them, he notes.
Biacore systems have the ability to recognize all of the hallmarks of promiscuous binders, he insists: greater than 1:1 stoichiometry, a high Hill slope, irreversible binding, aggregation, and sensitivity to detergent. Dr. Giannetti utilized the latest methods in biosensor operation to develop a high-throughput procedure for hit identification from fragment libraries all the way to lead-generation chemistry. Key features include the ability to screen and verify thousands of compounds in a short time, the large dynamic range of the assay (kd from 200 pM to 20 mM), and the small amount of protein necessary for fragment screening (<0.5 mg protein from assay development through hit validation).
He warns of the “need to align crystallographic and SPR conditions,” lest the PEG often present in crystallography buffer destroy the binding observed in Biacore experiments; PEG should be included in the initial Biacore screen. Dr. Giannetti points out that Biacore systems can also be used to complement enzyme assays by confirming potencies. Using Biacore systems, he has identified binders to baits as varied as kinases, polymerases, cytokines, receptors, proteases, and oxidases.
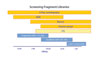
Vernalis’ approach to fragment-based drug discovery is called SeeDs, for structural exploitation of experimental drug startpoints. It includes the design of a fragment library, identification of fragments that bind competitively to a target, and evolution of these hits into leads.
Unique to the SeeDs strategy is displacement of bound fragments by a potent competitor, notes Roderick Hubbard, Ph.D., of the University of York and Vernalis. This feature ensures that the fragment is binding to the target’s active site and dramatically increases the quality of the initial hits, he says.
SPR has been successfully used in two phases of the SeeDs protocol. First, it has been used to identify compounds that bind, or if NMR was used for that, to validate binding. Since it is rapid and sensitive, SPR is an especially effective way to prescreen, or triage, a library before crystallization is attempted. It also allows for the screening of more hydrophobic fragments than NMR does.
Once binding has been established, the SeeDs protocol calls for SPR to characterize binding kinetics and thermodynamics. Applying SPR to the SeeDs method has successfully ascertained why isoxazole is a much more potent growth inhibitor than pyrazole, although they both have the same IC50 against Hsp90 (pyrazole is twofold faster on, but isoxazole is 15-fold slower off).
The FujiFilm Life Sciences Affinity Screening System, or AP-3000, was specifically designed and implemented for high throughput, sensitive analysis of small molecule binding, according to the company. The target protein is immobilized on disposable sensor sticks, with a short flow path to diminish reagent use but a wide bore to alleviate clogging.
There are six parallel channels, so six compounds and their corresponding blanks are analyzed simultaneously. It offers high-throughput (3,840 compounds in 24 hours) and high sensitivity (it can detect interactions as weak as 10 mM), and it is fully automated, from protein immobilization all the way through to data analysis, according to Don Janezic, the SPR business development manager. In addition, it can find hits, run dose response curves, and obtain affinity constants in less than two weeks, he says.
Conventional SPR Shake-Up
Graffinity’s screening method literally turns conventional SPR methodologies upside down. Rather than immobilizing the target protein on the chip and running each compound in the library over it, it immobilizes each compound in its own sensor field and runs the target protein over them, explains Mathias Woker, CBO, who adds that this method has a number of advantages.
First, he says, it can vary the point at which the compound binds to the chip, presenting different surfaces to the target protein.

Renate Sekul, Ph.D., head of research and development at Graffinity, thinks that “it is highly important to screen an active protein,” which is feasible in this system since the protein is in solution. This also makes standardizing the binding conditions across the library much easier.
Second, it is fast, Woker notes, making it well suited to high-throughput screens. This is partially because chip regeneration is not as much of an issue, and partially because they measure light spectrum shifts rather than light angle shifts to minimize errors, Dr. Sekul said.
Woker claims that “a Graffinity screen of 110,000 compounds takes 10 days where usually an experiment would take more than three months with strong limitations concerning the solubility of the compounds screened and the conformity of the results concerning consistent protein quality over the experiment time frame.”
Since 2002, Graffinity has screened over 80 targets, including kinases, phospodiesterases, proteases, nuclear hormone receptors, and even RNAs, and found hits for all of them, Woker concludes.
Protein Interaction Studies
Bernhard Geierstanger, Ph.D., group leader for NMR at the Genomics Institute of the Novartis Research Foundation, incorporates unnatural amino acids into proteins as site-specific NMR-active labels. This is “a unique way of putting an NMR active label at a chosen site in a protein,” he says, which is essential because in a “reasonable size protein—30 kD, for example—the number of signals is tremendous and impossible to resolve.”
Dr. Geierstanger, in collaboration with Peter Schultz, Ph.D., head of the institute, has created over 50 different unnatural amino acids with different purposes, along with their cognate orthogonal tRNA/aminoacyl-tRNA synthetase pairs. Some are fluorescent, some are photoactive, and some are labeled with fluorine or 15N or 13C. They can be used in lieu of any natural amino acid and placed exactly where the investigator chooses, which is a particular boon in large systems.
A potential limitation is that only one label can be incorporated into each sample. Monitoring the chemical shift change in a series of such single resonances, however, allows site-directed screening for binders. This can reduce the number of false positives currently seen in drug development by ensuring that the drug is binding to the correct site.
In an exciting twist, Dr. Geierstanger has made photocaged serine, cysteine, and tyrosine. Once incorporated at the desired site, the NMR label can be cut off of these residues. Thus, for the first time, individual amino acids can be labeled without altering the protein’s primary sequence, which alleviates any fears about changing the protein’s conformation or interactions. This allows him to use NMR for label-free binding, much like the other investigators use SPR.

