April 15, 2014 (Vol. 34, No. 8)
With the high cost of protein A capture resins for monoclonal antibodies, considerable development work has focused on alternative capture resins. Biopharmaceutical developer Synthon, for example, has been looking into selective precipitation of impurities preceding cation exchange (CEX) capture.
Precipitation removes host cell proteins, DNA, lipids, and polysaccharides, which are filtered off before the capture step.
As the flocculating agent, Synthon selected calcium chloride for its low cost, low toxicity, and lack of intellectual property constraints. “Its activity is relatively pH-independent,” explains lead scientist Guy de Roo, Ph.D. “After filtering away the precipitates, we acidify to pH 5 and load the material onto the AEX column.”
Synthon’s model system is a CHO cell culture producing a monoclonal antibody. The company has also used selective precipitation on duckweed extracts, but believes the approach has even broader applicability. “It should work with any eukaryotic cells used for biomanufacturing, since they all have the same types of DNA and host cell impurities,” Dr. de Roo says.
Companies considering a similar approach need to consider the economics, which are based on several factors. Binding capacity, for example, is about 70 g/L for Poros AEX (Thermo Fisher Scientific), the resin Synthon employed; protein A captures approximately 30 g/L. And AEX recovers approximately 90% of the product compared with protein A resins. “We lose about 10% in the flocculation,” notes Dr. de Roo.
Losses may be due to simple precipitation, binding to the flocculant, or deamidated species coming out of solution due to the increased number of negative charges. The question is whether the loss of yield is accompanied by lower or higher quality for recovered product. “We’re looking into that very question,” Dr. de Roo assures.
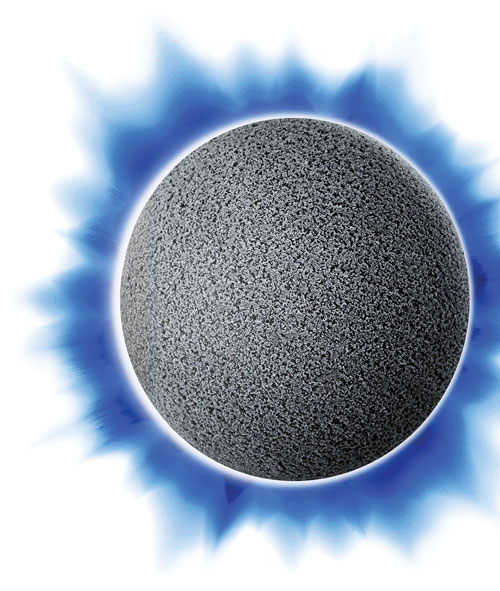

Thermo Fisher Scientific researchers maintain that in contrast to conventional chromatography particles, POROS® particles have large throughpores, which reduce diffusion time and allow increased flow rates without compromising capacity or resolution.
Using Superpro Designer (Intelligen), Dr. de Roo’s group calculated overall savings of 25% for downstream operations. One would expect that number to be higher based on the cost differential between protein A and AEX resins. However, capture is not the only expensive protein purification operation. Processors must still deal with a second column, filtration, virus removal, and membrane treatments.
Dr. de Roo attributes the relative lack of interest in AEX to “conservatism” on the part of biotech companies. “Why change something that’s working so well? Companies using platform technologies will be reluctant to abandon them,” Dr. de Roo explains.
Also, scaling up precipitation is more troublesome than switching to a larger column. “With protein A, you just load it, and it always works, regardless of the antibody type. AEX requires more fine-tuning,” comments Dr. de Roo. “What works for one antibody may not work for another. Nowadays, however, with the emergence of high-throughput process development tools, arriving at an acceptable capture step based on AEX should not be too difficult.”
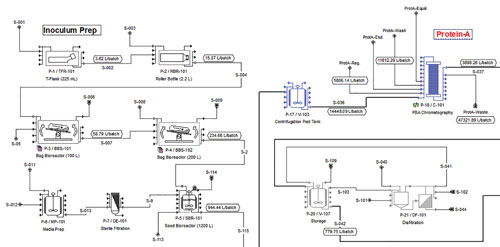
Intelligen’s SuperPro Designer was designed to facilitate modeling, evaluation, and the optimization of integrated processes in the production and purification of monoclonal antibodies.
Modeling and Multimodal Resins
Multimodal chromatography has immense potential for purifying many types of therapeutic proteins. Combining hydrophobic interaction and ion exchange in one resin creates exciting possibilities, but the complexity of multimodal interactions with diverse protein topographies has hindered new resin development and thwarted attempts at arriving at general rules for what works and what does not.
Steven Cramer, Ph.D., a professor of chemical and biological engineering at Rensselaer Polytechnic Institute, is using biophysical tools (NMR, SPR, and AFM), molecular simulations, coarse-grain modeling, and “a lot of chromatography” to develop models that predict relevant interactions between resins and proteins.
What’s interesting about multimode resins is that they invalidate many preconceptions regarding what works in chromatography. Dr. Cramer has recently used Capto™ adhere resin (GE Healthcare) as the capture step for a growth hormone analog, and achieved “exquisite capture and very high purification in one step.”
Even monoclonal antibodies, which are captured by cation exchange resins (in addition to protein A), might be purified by anion exchange multimode chromatography. “Even though the net charge on the monoclonal antibody may be positive, there are still hydrophobic regions, and negative patches on the protein surface,” explains Dr. Cramer. “So, depending on the protein, you can still get capture with a multimodal anion exchanger.”
Under the right conditions, anion change-based resins could also be used in flow-through mode to remove impurities that might otherwise be removed by precipitation.
Dr. Cramer has collaborated with several large resin and biomanufacturing companies on his approach. He found that they need to appreciate that multimodal interactions are not as straightforward as protein A binding for monoclonal antibodies. For example, many companies that express interest in the multimodal approach mistakenly believe that a resin capable of acting in either HIC or ion exchange mode should automatically bind to molecules with known affinity to these modes. It’s not that simple.
“It really depends,” Dr. Cramer says. “It’s not simply the sum of hydrophobic or electrostatic groups. Rather, it’s a matter of the patches on protein surface. How big are they, and how close together?” Ideally, the interaction should involve synergistic multimodal binding, not just a souped-up version of an ion exchanger.
That is why some resins behave like a multimodal ligand for one protein and like a unimodal ligand on another. It depends on the distribution of “patches” on the protein surface, their chemical nature, and their proximity.
Using simulation and biophysical data, Dr. Cramer has devised a way to quantify a protein’s affinity behavior and predict, through a “supermodel,” the binding of any protein with any multimodal system, at any pH.
Marcel Ottens, Ph.D., an assistant professor of microbiosystems technology and process chromotography at Delft University of Technology, uses computational and modeling tools with “biothermodynamic” inputs to predict the behavior of therapeutic proteins in common purification settings. Biothermodynamics is a term borrowed from the study of energetic characteristics of living systems and is equivalent to “thermodynamics.”
Dr. Ottens applies this modeling approach to chromatography, but it is also relevant to protein phase behavior as it relates to adsorption, extraction, solubility, partitioning, precipitation, and crystallization. This kind of approach, says Dr. Ottens, “paves the way for in silico process development, providing better process understanding.”
Normally, high-throughput process development is 100% experimental, for example, through high-throughput, automated design-of-experiment. Here Dr. Ottens uses the experiments to determine the parameters of thermodynamic or mechanistic models. “Once we have these parameters, we have what we need to perform scenario analyses on the computer. We can optimize configurations or operational conditions for a particular unit operation.”
Think of this approach as human-assisted in silico modeling where the person sets reasonable constraints and the computer works through data from actual high-throughput, automated experiments. Without human input, the computational problem becomes too complex to solve practically.
Dr. Ottens describes these inputs as “rules of thumb, heuristics, and common sense that limit the combinations you need to run in silico.” Without the computer, the design of experiment possibilities become too numerous for actual experimentation. Dr. Ottens has formed a company, Marlin Biopurification, to commercialize his technology. At present, the company is operating as a service center at Delft University of Technology.
Platform Viral Clearance
Michael Clark, Ph.D., senior scientist at AbbVie Bioresearch Center, believes that downstream processing will enjoy greater efficiencies through robust, predictable virus clearance during antibody purification based on platform processes.
While the evolution of viral clearance has been influenced by regulators, its ultimate driver is research and development toward greater patient safety from within the biopharmaceutical industry itself.
“Regulatory challenges associated with viral safety, which include prevention, detection, and mitigation, are the result of scientific progress that reveals the need for more sensitive detection of adventitious agents by exposing the limitations of existing techniques,” contends Dr. Clark.
“Platform process,” within the context of viral clearance, refers to standardized, predictable methodologies applicable to most monoclonal antibody purifications. The term may refer to a single process step such as low pH inactivation of viruses, or to an entire manufacturing process from the cell culture seed train to the final packaged product.
“Each process parameter may not be exactly the same from one project to another, but would be within well-defined ranges that have been demonstrated to deliver comparable product quality and process performance,” Dr. Clark adds.
The degree to which bioprocessors control parameters within a platform process depends on the mechanism of action for that process step. For example, tightly controlled parameters for low pH inactivation may include pH adjustments, hold duration, and buffer composition. Parameters with somewhat wider operating ranges include antibody concentration and temperature.
When a new therapeutic protein purification process is generated from scratch, testing viral clearance during development is most appropriate. According to Dr. Clark, however, clearance is highly predictable when using platform processes.
“In this scenario, the first GMP clinical batch for a new antibody is commonly manufactured without prior viral clearance testing. Following manufacturing, the capability of the process to remove viruses for that specific project is demonstrated as part of the required approval process prior to proceeding to clinical trials,” explains Dr. Clark. “Waiting to evaluate viral clearance until after the GMP campaign helps ensure that the studies faithfully represent actual manufacturing conditions.”
Platforming across Molecule Types
Common antibody purification techniques also apply to Fcabs, an emerging molecular class structurally related to monoclonal antibodies. An Fcab is derived from the Fc (constant) region of an antibody containing binding sites. Conventional antibodies bind in the variable (Fab) region. Vienna, Austria-based F-star uses plug-and-play technology to create bispecific antibodies with affinity built in from a library of 10 billion uniquely engineered Fc-binding domains. In addition to binding, Fcabs also mobilize immune effector functions.
“The molecules contain binding sites for Fc receptors, and also for c1Q, the first protein in the complement system that interacts with IgGs,” says Florian Rueker, Ph.D., an F-star advisor and faculty member at Vienna’s Universität für Bodenkultur (University of Agricultural Sciences). “It also contains a binding site for the neonatal Fc receptor, which mediates the long half-life of antibodies, and a protein A binding site.”
Fcabs’ resemblance to monoclonal antibodies means that protein A resins capture both molecules. “There is no need for affinity tags or novel separation methods to purify Fcabs,” Dr. Rueker adds.
Fcabs’ ease of manufacture compared with monoclonal antibodies also aids in purification. Unlike antibodies, which are constructed by co-expression of heavy and light chains, conventional monoclonal antibody manufacturing seeks to balance heavy and light chain expression. Overexpressed moieties usually wash through during protein A capture, but they nevertheless represent a downstream burden.
Fcabs are homodimeric—both subunits are identical, so all molecules end up in the product. “As with antibodies, Fcabs are more or less pure after the capture step,” Dr. Rueker states.
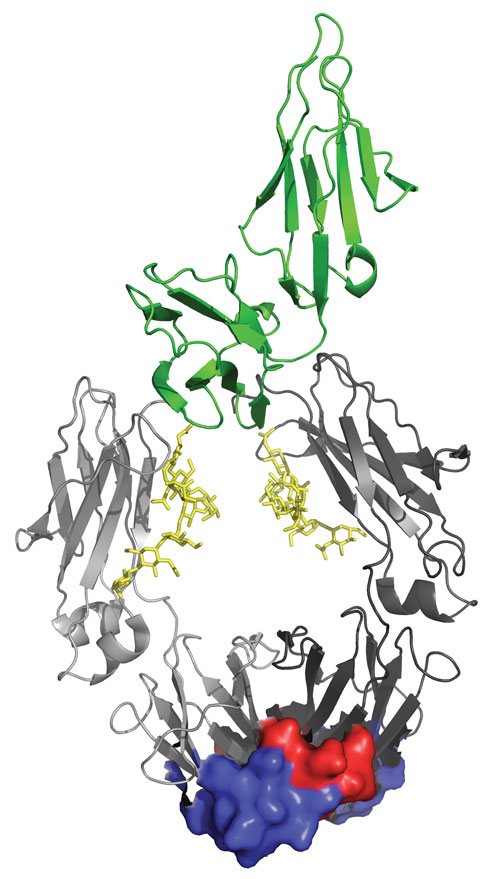
Ribbon presentation of a human antigen-binding Fc fragment of IgG1 (grey) in complex with CD16 (green). Each of the two antigen-binding sites in the Fc homodimer (dark and light grey indicate the two chains) is composed of residues in the AB (red) and in the EF (blue) loop. Sugar residues at the N-linked glycosylation sites are indicated in yellow. Image was made on the basis of Protein Data Bank entry 1T83 [32]. [Florian Rueker, Ph.D.]



