May 15, 2011 (Vol. 31, No. 10)
Taking a Constructivist View, Engineering Starts with the Basics and Progresses toward Complexity
The philosophical theory of reductionism pursues the idea that biological systems are most fruitfully investigated at the lowest possible, most fundamental level. For many years, philosophy of science concerned itself with whether it is adequate to analyze a system by dissecting it into the component parts and whether molecular biology can explain all biological facts, obviating higher-level biological theories.
Can biology really be reduced to physics, chemistry, or, perhaps, simple engineering concepts? Synthetic biologists bring this question into sharp focus by reasoning that the function of complex systems can be elucidated by reconstruction of the whole from the basic parts. “However, as opposed to reductionism, synthetic biologists take a view of learning as a constructivism,” argues George Church, Ph.D., department of genetics, Harvard Medical School.
“Synthetic biology is about systems-level planning and testing. It is akin to systems engineering from bottom up as opposed to reduction from top down. We design and construct high order biological devices out of the biological building parts.”
The BioBricks Foundation’s international conference next month will discuss synthetic biology in the context of technology, bioethics, biosafety, and intellectual property.
Analysis & Insight: IP Rules for Synthetic Bio Generate More Questions than Answers
Biologists, bioethicists, and politicians are all weighing in on what guidelines should govern syn bio. Until there is a standardization of synthetic biology parts and devices, intellectual property issues remain up in the air and will follow similar ups and downs as the larger biotech field. Among unresolved issues are gene patentability, patent protection vs. open access, and the effect of patent reform. You can get all the details by reading our story.
For synthetic biologists, DNA and RNA sequences are receivers, senders, reporters, and actuators. These bioblocks make devices, and devices are connected to make systems. Parts, devices, and systems can be combined in self-contained modular units.
The BioBricks Foundation maintains the shared repository of standardized biological parts that can be assembled in a standardized way. Thus created, synthetic biological systems can be used for the purposes of capturing and processing the information, producing energy or manufacturing biomaterials.
The emerging field of functional RNA engineering offers considerable advantages over the engineering of more common protein-based devices, as RNA-based devices are nonimmunogenic and more readily programmed. Its versatile sensing and actuation functions make RNA a powerful design substrate in synthetic biology.
Christina D. Smolke, Ph.D., department of bioengineering, Stanford University, has designed RNA devices that sense increases in target protein signals that trigger certain cellular behaviors, such as the uncontrolled cell growth associated with cancer. Next, sensing of disease-related signaling by the RNA devices can be coupled to new cellular behaviors, such as cellular apoptosis.
“We envision that our devices will be delivered via viral vectors or nanoparticles to cells and can actuate responses specifically in diseased cells based on molecular markers of disease. Another application of these genetic devices is modification of stem cells and immune cells to improve their therapeutic potential. Our RNA devices can regulate various cellular functions, including proliferation and therapeutic efficacy.”
Dr. Smolke’s RNA device is a synthetic gene with built-in sequences for RNA aptamers. Aptamers are short noncoding RNAs that are able to recognize specific ligands such as proteins; essentially acting as the RNA analog of antibodies.
When the gene module is transcribed into RNA, the RNA aptamers bind to the target proteins. This binding event in turn hinders or disrupts a potential splicing event, resulting in the binary regulation of the RNA splicing event (spliced/not-spliced), which can be coupled to any type of genetic output, like a fluorescent signal or an apoptotic event.
The device’s specificity is enhanced by incorporating additional aptamer sensors against other signaling proteins. “Both the input processing function and genetic output are modular and can be altered to suit multiple applications,” says Dr. Smolke. “One can extend these systems to create quite sophisticated RNA-processing devices that are able to differentiate signals from complex regulatory networks.”
Controlling Disease Organisms
Synthetic biology could also be deployed to introduce a predictable control over higher-order organisms. Oxitec uses synthetic components to create sensors, actuators, and effectors that regulate functions of insects. These regulatory devices are deployed to generate insects whose offspring are not able to survive in the wild.
Termed RIDL (release of insects carrying a dominant lethal), the insects carry a synthetic construct that is not active under the laboratory conditions, when the insects’ feed is supplemented with tetracycline. When tetracycline is withdrawn, the genetic device is activated, resulting in accumulation of a recombinant marker protein, eventually interfering with the normal insect functions. Only females are affected, because the actuator is triggered by a sex-specific alternative splicing mechanism.
“We used a naturally occurring phenomenon found in Aeges aegypti mosquitos, a host of Dengue fever,” says Luke Alphey, Ph.D., CEO. “In these insects, actin-4 gene is predominantly expressed in the flight muscles of females. It is practically absent in males due to an alternative splicing event. We genetically tightened the event, further improving sex specificity. Then we coupled this actuator with the marker protein.”
Accumulation of the marker protein causes loss of the ability to fly in females, leading to population collapse within three generations. “Importantly, the effect is delayed until the adult stage, which means that the genetically modified larvae can effectively compete with the wild type,” adds Dr. Alphey.
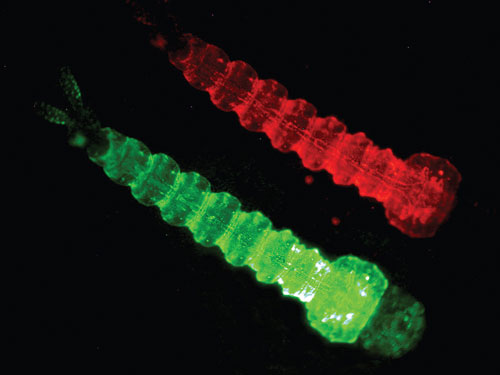
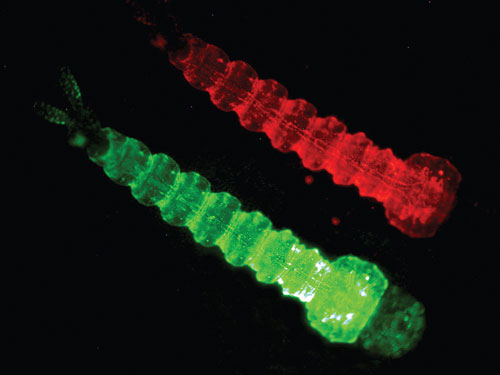
OX3604C is a female flightless RIDL strain from Oxitec. When mosquitoes are reared without the tetracycline antidote, adult females cannot fly or mate. Flightless females in the wild cannot seek hosts or mates, take a blood meal, or spread disease. They are likely to be rapidly eaten by predators, so this trait is equivalent functionally to a lethal condition. The strain contains the DsRed marker, which is clearly visible in larvae.
Measuring Output
Synthetic biology promises to generate microorganisms with novel or improved metabolic capabilities. The Fuels Synthesis Division of the Joint Bioenergy Institute uses the tools of synthetic biology to engineer new microbes that can quickly and efficiently ferment plant waste products into biofuels as well as into other valuable chemical products.
Since the fuel performance is highly dependent on the composition of fatty acids, many synthetic clones need to be screened to identify the top producer. Ideally, such analysis would capture all metabolites in a broad net, measure metabolites directly rather than via a surrogate marker, and allow for high-throughput format.
Analysis and comparison of the detailed metabolite profiles may also help to deduce why some clones perform better than others and to adjust the components to optimize production for a particular lipid. Mass spectrometry is ideally suited for such screening applications, providing the exact composition of analytes. Conventional techniques for metabolite profiling aren’t suitable for screening large libraries due to limitations on throughput.
“A high-sensitivity and -throughput technological solution was needed that could screen large libraries for key metabolite levels,” remarks Trent R. Northen, Ph.D., scientist at Lawrence Berkeley National Laboratory. He developed a nanotechnology for mass-spec analysis that his group used to develop a high-throughput fatty acid screen.
The silicon surface is composed of nanostructures with 10 nm pores. A liquid, termed “initiator”, is trapped in the pores. A sample is loaded on the substrate, spreading over the pores containing the trapped initiator. Rapid heating of the silicon substrate by the laser causes vaporization of the initiator, desorbing the analyte molecules. “The NIMS, or nanostructure-initiator mass spectrometry, is well positioned for screening of small molecules to support synthetic biology.”
Highly Parallel Gene Synthesis
The commercial cost of large-scale DNA synthesis makes it prohibitively expensive to make millions of genome-scale constructions as required for optimization of metabolic and genetic systems. To overcome this limiting factor, the team led by Dr. Church utilizes an unorthodox gene-synthesis method, which he claims reduces the cost at least 20-fold. First, a computer algorithm helps to break down the desired sequence into 40–150 bp pieces. Then these oligonucleotides are printed on custom microchips, which can pack as many as 55,000 individual sequences.
“The advantage of this method is in highly parallel synthesis of thousands of DNA fragments,” says Emily Leproust, Ph.D., director, applications and chemistry R&D, Agilent Technologies. “Any conceivable DNA variants can be all done at the same time. The resulting pool of oligos can serve as a starting point for many creative applications.”
Dr. Church’s team amplified the oligo pool, generated from Agilent’s Oligo Library Synthesis (OLS) platform, via several rounds of PCR reactions. The fragments were linked together, creating hundreds of genetic products of kilobases or more.
“This technique is a core technology at the Center for Causal Consequences of Variation,” comments Dr. Church. “We continue the optimization of the process in order to make writing of human genome accurate and efficient.” Writing, or editing, of the genome on the large scale will enable researchers to test what variations in the human genome really cause diseases.
“By engineering many combinations of such variations, and directly observing their impact on the cell, we will be able to establish cause and effect in a more direct way than genome-wide association studies allow for.”
“To test the genetic variations, we place our synthetic constructs into human cells by using synthetic proteins.” The team has generated artificial versions of TALEs (transcription activator-like effectors), DNA-binding proteins found in the plant pathogen Xanthomonas spp. Each monomeric domain of a TALE can be engineered to recognize just one specific nucleotide. A specific arrangement of TALE monomers, therefore, will recognize a specific DNA motif of 18–30 base pairs.
TALEs bring with them the enzymes for DNA cleavage and recombination, enabling the insertion of the synthetic DNA precisely in the desired location.
Creatively designed OLS pools have been used in generating synthetic bioblocks. Researchers from the University of California, San Francisco designed OLS microchips containing thousands of short hairpin RNAs (shRNAs). At approximately 30 shRNA/gene, the pool targeted ~600 genes.
“Without Agilent OLS technology, this would be cost-prohibitive,” adds Dr. Leproust. “We print oligos on the femtomole scale, >1,000 times lower output than the column-based method. And the library now can be created without the extensive cost of synthesizing and assaying the silencing efficacy of these oligos one at a time, but rather as a pool in a simple tube assay.”
The shRNA library is delivered into cells by lentiviruses. The outcome is followed by sorting of individual cells according to a desired phenotype. OLS strategy was also used to accelerate discovery of new bioblocks such as promoter elements. Synthetic promoters containing every possible combination of mutations were printed on the array and assayed as a pool off the chip for transcriptional efficiencies.
In a single experiment, researchers assayed the effects of all possible single-nucleotide mutations in a given promoter region. “Saturation mutagenesis on this scale is simply impossible by other methods,” concludes Dr. Leproust.