July 1, 2014 (Vol. 34, No. 13)
Kathy Liszewski
With the rapid rise of next-generation sequencing (NGS), one of its technologies, RNA sequencing (RNA-Seq), has taken center stage for analyzing whole transcriptomes. Although RNA-Seq is still the new kid on the block, this technology, its proponents say, could revolutionize transcriptomics, revealing the architecture of gene expression in unprecedented detail.
RNA-Seq applications are proliferating and include the elucidation of disease processes, targeted drug development, and personalized medicine. RNA-Seq also has veterinary and agricultural applications. RNA-Seq, however, will realize its promise only if certain impediments are overcome. These include consistency of methodology, cost, and optimization of data analysis.
To orient researchers who are unfamiliar with the differences between RNA-Seq platforms, Kelli Bramlett, R&D scientist, Life Technologies, poses two key questions:
1. Are you interested in pure discovery, in a nonguided fashion, of every RNA species present in your test samples?
2. Are you mainly focused on measuring expression levels of well-annotated coding RNA transcripts?
“You might have a set of genes crucial to identifying a disease state, or profiling the stage of a specific type of cancer, or monitoring development in your experimental system,” elaborates Dr. Bramlett. “You then would want to employ a system that allows you to quickly and efficiently focus on just your genes of interest and screen through many different samples in a short amount of time.”
Researchers who are clear about their experimental goals are better able to choose the technology that best suits their needs.
“Some RNA-Seq allows for true discovery but will require greater sequencing depth to get all the information. Oftentimes, this type of deep sequencing requires significant additional time for analysis,” continues Dr. Bramlett. “If a focused panel targeting specific RNAs will better meet your needs, this can be accomplished on systems with much faster turnaround time and less sequencing depth. Also, the analysis of a less complex targeted library is much more straightforward.”
Enhancing Sensitivity
RNA-Seq has advanced our ability to characterize transcriptomes at high resolution, and the laboratory and data analysis techniques used for this NGS application continue to mature, notes John Tan, Ph.D., senior scientist, Roche NimbleGen. “High sequencing costs combined with the omnipresence of pervasive, abundant transcripts decrease our power to study rare transcripts, decrease throughput, and limit the routine use of this technology.”
For example, notes Dr. Tan, a small number of highly expressed housekeeping genes can be responsible for a large fraction of total sequence reads in an experiment, thus increasing the amount of sequencing required to characterize less abundant transcripts of interest.
To improve the cost-effectiveness, throughput, and sensitivity of RNA-Seq, Dr. Tan and colleagues are developing methods to perform targeted RNA-Seq. “Targeted enrichment of transcripts of interest circumvents the need to perform separate rRNA depletion or polyA enrichment steps on input RNA,” explains Dr. Tan. “By targeting their sequencing, researchers can avoid wasting resources on housekeeping transcripts and focus instead on genes or genomic regions of interest.”
Targeted RNA-Seq can allow deeper sequence coverage, increased sensitivity for low-abundance transcripts, less total sequencing per sample, and more samples processed per sequencing instrument run. “Significantly, we observe that the enrichment step also preserves quantitative information very well,” adds Dr. Tan. “These advances will facilitate a more routine use of RNA-Seq technology.”
Sample Integrity Issues
Some samples present challenges for accurate RNA-Seq. “Formalin-fixed, paraffin-embedded (FFPE) patient tissue archives and the clinical data associated with them can be invaluable when dissecting the etiology and prognosis of disease. However, these may provide only limited amounts of sample that may also be degraded,” comments Gary Schroth, Ph.D., distinguished scientist, Illumina. “Standard techniques for RNA-Seq often don’t work.”
Dr. Schroth says that most labs currently gauge RNA integrity via the RIN (RNA integrity number). “Labs often will look at the RIN as a measure of quality control prior to working with sample. We found that the RIN number from FFPE samples is not a sensitive measure of RNA quality or a good predictor for library preparation. A better predictor is RNA fragment size. We developed the DV200 metric, the percentage of RNA fragments greater than 200 nucleotides, a size needed for accurate construction of libraries.”
Illumina offers its TruSeq® RNA Access Library Preparation Kit especially for FFPE samples. This kit, when used with the DV200 metric, provides cleaner and more accurate library preparation. This new approach allows researchers to start with five-to tenfold less material when making libraries from FFPE samples.
Strand Specificity
Most NGS requires initial construction of libraries that may not provide the specificity desired even when prepared from mRNA. “Traditional RNA-Seq library preparation loses the strandedness of transcripts—information that is critical in understanding cellular transcription,” says Jungsoo Park, senior marketing and sales manager, Lexogen.
According to Park, Lexogen tackled this problem by developing a method to generate libraries with greater than 99.9% strand specificity with a simplified process that takes 4.5 hours to complete. Lexogen’s SENSE mRNA-Seq library kit initially isolates mRNA via the poly A tail and utilizes random hybridization of the transcripts that are bound to the magnetic beads without transcript fragmentation. “This is a revolutionary method, which keeps high strandedness of the transcripts,” asserts Park. “And it takes only half a day to finish.”
One of the novel aspects of this approach is the use of starter/stopper heterodimers containing platform-specific linkers that hybridize to the mRNA. “The starters serve as primers for reverse transcription, which then terminates once the stopper from the next heterodimer is reached,” explains Park. “At this point, the newly synthesized cDNA and the stopper are ligated while still bound to the RNA template.” According to Park, there is no need for a time-consuming fragmentation step, and library size is determined simply by the protocol itself.
For researchers only intending to see the expression levels, sequencing of the entire mRNA transcript will require subsequent bioinformatics processes such as RPKM, a measure of relative molar RNA concentration. To meet this challenge, Lexogen developed a kit, QuantSeq, which counts the 3′ end of each transcript. “Sequencing mRNA at or close to the 3′ end and counting their reads,” says Park, “provides an economical alternative to microarrays for gene expression and other studies.”
Transcript Targeting
Cells possess many thousands of transcripts. Digging through them to find just the right one can be a challenge for sample preparation prior to RNA-Seq. “Human as well as nonhuman samples often have an abundance of uninformative transcript species that can compromise data quality and the cost-effectiveness of sequencing,” notes Steven R. Kain, Ph.D., director of product management, NuGEN Technologies.
The company has developed a method for targeted depletion of unwanted transcripts following construction of RNA-Seq libraries. The method, called Insert Dependent Adaptor Cleavage (InDA-C), employs customized primers that target specific transcripts, such as ribosomal and globin RNAs, to exclude from final RNA-Seq libraries.
“For example, hemoglobin RNA derived from blood accounts for at least 60% of transcripts,” explains Dr. Kain. “By depleting these two transcript classes, we were able to quadruple informative reads in some instances.”
Unlike other methods using hybridization-mediated pull-down strategies to deplete unwanted RNA species, InDA-C does not perturb the original total RNA input to the workflow and thus avoids off-target mRNA cross-hybridization events that can potentially introduce bias. A key aspect of the technology is the versatility of the primer design.
“The species and transcript specificity of the workflow relies on the design of InDA-C primers, which can be constructed to target virtually any class of unwanted transcripts for targeted depletion,” continues Dr. Kain. “We also provide a no-cost design service to assist researchers in choosing InDA-C primers for targeted depletion of virtually any unwanted transcript type from any species.”
For an even more targeted approach to RNA analysis, NuGEN has developed Single Primer Enrichment Technology, which can be used to prepare targeted NGS libraries from both gDNA or cDNA. The approach, which combines a rapid single-day workflow with flexible and customized target design, has been used identify gene fusion products and alternative splicing patterns from enriched cDNA libraries.
NuGEN Technologies offers its Ovation Human Blood RNA-Seq Library System as an end-to-end solution for strand-specific RNA-Seq library construction. As little as 100 ng of total RNA obtained from adult human whole blood may be needed. (The image here depicts a circulating tumor cell in whole blood.) NuGEN’s Insert Dependent Adaptor Cleavage (InDA-C) technology can provide targeted depletion of unwanted high-abundance transcripts and can result in a significant reduction in sequencing reads derived from human rRNA and adult human reticulocyte globin.
Automation Advantages
Preparation of libraries for RNA-Seq entails an intensive workflow. When done manually, the process can be fraught with inconsistencies and prone to human error. “Automation can enhance sample quality and throughput and provide a more efficient and economical workflow,” notes Alisa Jackson, senior marketing manager, Genomic Solutions, Beckman Coulter.
According to Jackson, automation provides four key advantages:
1. Creation of high-quality mRNA libraries. Initial steps in this process include depleting samples of ribosomal RNA. Although it has the greatest abundance, rRNA gives the least amount of information.
“We’ve automated this process on our Biomek instruments using popular sample preparation kits from Illumina and New England Biolabs,” notes Jackson. “Accurate pipetting and thorough mixing are critical for this process. The Biomek liquid handler’s 96-channel pipetting head is used in combination with an on-deck orbital shaker to vigorously mix samples. Results show this ‘mix and shake’ approach works well.”
2. Limited exposure to RNAses from human contact. Every scientist’s nemesis when working with RNA is the universal presence of RNA-degrading RNAses. To help overcome this problem, says Jackson, “Biomek consumables such as pipette tips are DNase and RNase-free.”
3. Reduced exposure to toxic chemicals. “An instrument dispenses all reagents involved in the various steps of process.”
4. Enhanced reproducibility. “This is still a very expensive process,” asserts Jackson. “Obtaining accurate results the first time prevents costly repetitions. For this reason, we provide Biomek methods for many NGS library preparation kits. By fully testing these methods with real-life samples, we ensure reliable and repeatable creation of sequence-ready RNA libraries, whether stranded or nonstranded, mRNA or total RNA.”
Versatile liquid-handling platforms automate the RNA sequencing sample preparation pro- cess, increasing reproducibility and conserving valuable samples and reagents. [Beckman Coulter]
What’s Next?
“With RNA-Seq, we are closing in on personalized medicine,” suggests Qichao Zhu, Ph.D., principal scientist, Boehringer Ingelheim. “This technology allows more exact identification of patient subgroups. Instead of ‘one drug fits all,’ we can now begin to more appropriately define which drugs will work in which patients. Diseases such as cancer and cystic fibrosis as well as neurodegenerative illnesses have many patient subcategories. Future pharmaceutical drug discovery will be better able to develop targeted therapeutics with the help of RNA-Seq.”
There are still many challenges in the field, however. “A critical aspect is accuracy. Given the large scale set of RNA-Seq, even 99.99% accuracy is not good enough for diagnostics,” insists Dr. Zhu. “Further, as we move forward, we will need to improve many aspects of the technology including disease tissue sample isolation, library construction methodologies, as well as analysis of massive datasets.”
Dr. Zhu speculates that it’s only a matter of time until the technology is suitably refined: “In the future, a patient will go into the doctor’s office and have a whole transcriptome profile test performed.” Also, aside from personalized medicine, Dr. Zhu anticipates applications ranging from drug development to veterinary medicine and agriculture.
“The only limitation of RNA-Seq is brain power,” elaborates Dr. Zhu. “When PCR technology was discovered, no one knew just how powerful it would become or how many applications it would generate. Now, it is used everywhere. NGS technology and RNA-Seq have a similar potential. We are limited only by how much we can imagine.”
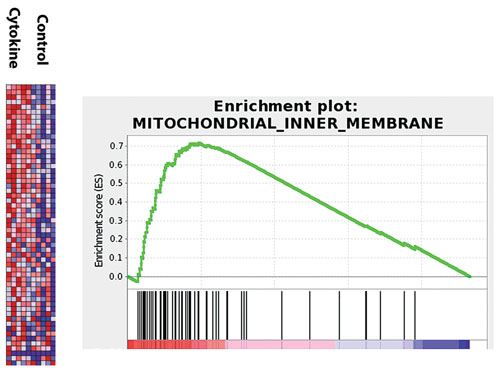
RNA-seq data analysis for target identification. [Boehringer Ingelheim]



