October 1, 2016 (Vol. 36, No. 17)
Practical Solutions to Expression Problems Include RNA Interference and Glycoengineering of CHO Systems
Like a train that consists of a particular sequence of cars, from locomotive to caboose, proteins can only “leave the station” after each of its segments, amino acids, are hooked together.
Actually, the shuttling of cars in a rail yard is a lot simpler than the processes that make up protein expression—processes that begin with transcription, culminate in translation, and are even elaborated in post-translational modification. It is possible, however, to take control of these processes and freight valuable cargo such as complex biopharmaceuticals.
Aiming to provide practical solutions for today’s challenges, the recent Bioprocessing Summit focused one segment of its conference program to advances in protein expression. This segment covered new approaches such as utilizing genome-scale RNA interference (RNAi), improving glycoengineering of the ever-popular Chinese hamster ovary (CHO) cell expression systems, and harmonizing transient and stable expression to enhance drug development.
Presentations were also devoted to insect cells. Like other expression systems, insect cells can be used to produce biotherapeutics with careful optimization of manufacturing-compatible processes. Finally, some presentations touched on outsourcing issues, such as the relative advantages of using one or several companies for the multiple processes involved in protein expression.
Multidisciplinary Approach
Given the myriad challenges that can compromise efficient protein expression, it sometimes makes sense to consult specialists. For example, outsourcing issues have prompted many companies to work with Ingenza, which says it aims to be the partner of choice in industrial biotechnology and synthetic biology.
“There are many unseen as well as known facets to effective protein expression that require a high level of expertise to simultaneously optimize,” stated Ian Fotheringham, Ph.D., Ingenza’s managing director. “Some laboratories have knowledge in one aspect or another, but often it is more efficient to work with someone with the capabilities to address the multiple aspects of effective protein expression. We are distinctive in that respect.”
Dr. Fotheringham asserted that outsourcing is now a well-established option in the field of protein expression: “Customers want someone who can produce, innovate, troubleshoot, and continually dialog with them to increase their chances for success. We are a multidisciplinary company for a reason.”
As an example, Dr. Fotheringham described working with a leading U.S. biopharma company whose expressed target protein was truncated in the recombinant host. “First, we optimized the target gene for expression in Escherichia coli using a library of ~50,000 coding region sequence variants. We devised a simple petri plate screen that rapidly identified the best full-length performers.
“Prompted by such successes, we subsequently provided GMP-compliant production strains for many of our customer’s targets. Other projects have successfully involved dual cassette vectors (such as target and chaperone), alternate hosts, novel induction strategies, and targeted genome integration to optimize recombinant protein stability, production, and purification.”
Ingenza also works with academic institutions and emerging spin-outs to translate innovative research. Dr. Fotheringham cited the example of epidermicin, a potent antimicrobial that was discovered by one of the company’s university partners. Epidermicin can combat methicillin-resistant Staphylococcus aureus (MRSA) infection, but initially it was expressible only in low amounts due to host toxicity.
“We utilized our proprietary inABLE® combinatorial assembly platform to prepare and screen cleavable fusion constructs which, combined with design-of-experiment (DOE)-driven fermentation, permitted over-production of epidermicin and its derivatives,” detailed Dr. Fotheringham. “Success is enabling a university spin-off to exploit the outcome.”
The company is now moving further into the GMP biologics arena with key clients, using optimized microbial and mammalian production systems and modular single-use fermentation approaches to provide drug substances for clinical evaluation and manufacturing.
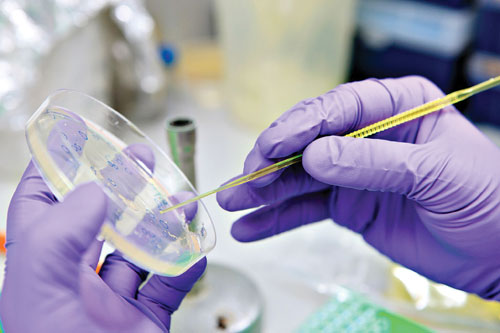
Ingenza utilizes simple petri dish screens to optimize target gene expression in Escherichia coli using a library of ~50,000 or more variants.
Harmonizing Transient and Stable CHO Expression
For those working in-house, expression of protein products in CHO cells remains one of the most popular expression platforms for a variety of reasons, said Yashas Rajendra, Ph.D., research scientist, Eli Lilly and Company. “CHO systems,” he explained, “are inherently adaptable and easy to scale up to industrial bioreactors”
However, for early-phase drug discovery of complex biopharmaceuticals, companies often rely on HEK293 transient expression platforms. “This can increase the risk of surprises when molecules are advanced and eventually expressed in CHO,” warned Dr. Rajendra. “To reduce this risk, we developed a transient CHO platform that is based on the same cell line, media package, and DNA expression cassette used for our stable CHO platform.”
According to Dr. Rajendra, harmonization of transient and stable expression approaches can provide better predictability of protein quality and expression when transitioning the molecules from discovery to development and ultimately manufacturing. “This transient CHO system can rapidly (within seven days) generate high titers,” asserted Dr. Rajendra. “The system is scalable to 10 L.”
Dr. Rajendra’s group introduced another modification: the use of stable CHO pools (instead of master wells or clones) to generate gram quantities of therapeutic protein. “It takes 2–4 weeks to generate a stable CHO pool,” noted Dr. Rajendra. “In contrast, it usually takes several months to generate clonal CHO cell lines. Hence, speed is one major advantage of stable pools.
“Additionally, advantages over transient expression include use of smaller amounts of plasmid DNA for transfection, ease of volumetric scale-up, and the flexibility of performing multiple production runs over an extended period of time using a frozen cell bank. We recently published CHO pool titers ranging from 2 to 7.6 g/L.”
Dr. Rajendra indicated that at present, stable pools are primarily used for preclinical material generation due to concerns of clonal heterogeneity, genetic and expression stability, and product quality consistency. In the future, however, the use of stable pools may extend to the generation of toxicology lots and the first human dose studies.
“Although this is going to take some time,” Dr. Rajendra concluded, “given the advancements in host cell engineering, transposon-mediated approaches, and analytical tools that allow for in-depth product quality assessment, I am hopeful that we will see a shift from the ‘clonality’ paradigm to a “product quality consistency’ paradigm.”
Glycoengineering in CHO
Although they are widely used, CHO expression systems present challenges similar to those posed by other mammalian cells. In particular, CHO cells produce recombinant proteins that are heterogeneous in complex-type N-glycans.
“Glycosylation is one the most important post-translational modifications of proteins,” stated Andrew (Cheng-Yu) Chung, researcher, department of chemical and biomolecular engineering, Johns Hopkins University (JHU). “Glycans on proteins play a key role, particularly in protein folding. Thus, they significantly impact protein stability and protein-protein interactions.”
Chung, who is in the laboratory of Michael J. Betenbaugh, Ph.D., is studying sialylation of engineered proteins. “Because of the negative charge, size, and hydrophilic characteristics of sialic acid groups, sialylation is one of the very critical modifications on the glycan terminus,” Chung explained. These properties allowed sialic acid to have a substantial influence on protein-protein interactions. Further, sialylation can affect biotherapeutics’ efficiencies.”
Chung and colleagues examined the application of genetic engineering strategies to modulate the level of sialylation content on various potential biotherapeutics. “One common way to manipulate the glycan structure is via genetic engineering tools such as CRISPR/Cas9 to knockin/knockout or overexpress certain glycoenzymes,” he noted. “For example, overexpressing glycoenzymes involved in the sialylation pathway is a strategy to produce biologics with increased sialic acid content that may affect circulation retention time (CRT). Increased CRT may mean lower doses for patients.”
Another strategy to make glycosylation in CHO cells more uniform and similar to the analogous process in human cells is to add specific chemical supplements to the cell culture medium.
Chung’s group decided to try both strategies—genetic engineering and the addition media supplements—at the same time. “The genetic engineering of CHO cells can be used to produce biologics with enhanced sialylation as well as higher protein titers,” Chung indicated. This approach, continued Chung, has been combined with the use of chemical supplements. The specific chemical analogs that are added to the feed were developed by Kevin Yarema, Ph.D, an associated professor of biomedical engineering at JHU.
Insect Cell Expression
While most recombinant biotherapeutics are produced in mammalian cells, others are produced in insect cells. For example, an insect cell platform was selected to express recombinant proteins for Genocea Biosciences.
“Our therapeutic vaccine (GEN-003) against genital herpes is comprised of two recombinant proteins combined with an adjuvant,” said Rajiv Gangurde, Ph.D., associate director of protein production. “Both proteins are expressed in insect cells.
“The choice of the expression platform was primarily driven by specific post-translational modification(s) required for the activity, that is, the ability to elicit an appropriate immunogenic response, by one of the antigens. These modifications are not just specific to the host cells in which the recombinant protein is expressed, but are also well-tolerated in humans, which is one of the biggest advantages of using insect cells.”
Genital herpes is a serious chronic infection affecting more than 400 million people worldwide and over 50 million in the United States alone. Dr. Rajiv pointed out that there has been no real innovation on behalf of patients for decades, not since oral antivirals were approved.
Prior to expressing the product in insect cells, Dr. Rajiv and colleagues needed to overcome some specific hurdles. “Early on, we faced several challenges in developing manufacturing-compatible processes,” he noted. “We have overcome those hurdles through extensive in-house process development work and choosing a contract manufacturing organization with insect cell production scale-up experience.
“One protein is expressed as a soluble, secreted protein, while the other is an intracellular, insoluble protein. This fundamental difference in expression makes it easier to express the proteins separately. Moreover, there are notable differences in the levels of expression; hence, independent expression allows the required flexibility in downstream processing and drug product manufacture.”
The company, which is conducting its third clinical trial with GEN-003, expects to start Phase III in the second half of next year. Next, the company plans to file a Biologics License Application with the FDA in 2019. If the application is approved, the company will launch GEN-003 in 2020.
“We believe we have established GEN-003 as a highly attractive product that requires only a once-yearly injection offering similar efficacy to that of daily oral antiviral therapy,” asserts Dr. Rajiv. “Given that most patients choose not to take daily oral anti-viral pills and accounting for compliance challenges with daily therapy, we believe the real-world efficacy of GEN-003 from an annual injection could translate into significant patient benefits and a revenue opportunity in excess of $1 billion in the United States alone.”

Genocea Biosciences’ Rajiv Gurde, Ph.D., utilizes insect cells to separately express two key recombinant proteins for the company’s therapeutic vaccine, GEN-003, against genital herpes.
RNAi to Enhance Expression
Aside from its traditional use as a workhorse technique for basic research, RNAi also has applications for improving protein expression.
“RNAi has been often applied in the fields of medical and basic research, especially to understand disease mechanisms,” reports Joseph Shiloach, Ph.D., director of the Biotechnology Core Laboratory, Intramural Research at the National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health. “We decided to use it for identifying genes that are currently not known to be involved in protein expression, yet whose knockdown could improve recombinant protein production in mammalian cells.”
Dr. Shiloach and colleagues performed a genome-scale, high-throughput RNAi screen using HEK293 cells expressing firefly luciferase. “The basic idea was to individually knock down approximately 22,000 genes one at a time and look for those that affected expression, but did not affect cell growth,” explained Dr. Shiloach. “We utilized three different siRNAs for each gene and performed the work collaborating with the National Center for Advanced Translational Science.”
Among the “hits” identified was the gene for ornithine decarboxylase antizyme 1. The protein product helps regulate intracellular polyamines that are needed for cell growth and proliferation. “We also performed a detailed investigation to confirm the findings,” continued Dr. Shiloach. “We are now assessing the long-term effects of knockdown, the mechanisms involved in metabolism, and feasibility of extending this approach to other mammalian cells.”
According to Dr. Shiloach, the take-home message from these studies is that genome-scale RNAi screens can identify key pathways and genes whose modulation may help improve the expression and production of recombinant proteins. “These studies are more a beginning than an ending,” he stated. “But we have established a different foundation for the targeted design of more efficient mammalian cell expression platforms.”
Boosting Expression
Expression levels of recombinant proteins (simple proteins as well as complex biologics) can be unpredictable in mammalian cells, and this causes major headaches for target discovery and scale-up manufacture, according to Tom Payne, Ph.D., head of cell engineering at Oxford Genetics. Scientists are familiar with finding that sometimes proteins express at unexpectedly low levels, even when under the control of strong promoters (e.g., CMV).
“To minimize protein-to-protein variability and maximize yield, we adopted a high-throughput approach to screen and optimize >5,000 recombinant promoters by random shuffling, >30 5’ UTRs in different configurations and >15 poly-A signals,” said Dr. Payne. “We also developed Kozak and stop codon libraries and screened >1,000 genes from diverse organisms encoding putative ancillary protein to generate expression plasmids that consistently produce high yields of diverse proteins in HEK-293 and CHO systems, termed SnapFast Pro.”
Validating the System
To validate this vector system, Dr. Payne and his team compared protein yields to an industry-standard protein-expression vector. To maximize the relevance of the dataset they algorithmically determined the frequency with which each gene in the human genome had been cited in publications (PubMed hits, 2000–2016), identifying those genes consistently increasing in hits, as an indication of importance in biomedical research.
“The top 150 genes were batch codon-optimized, FLAG-tagged, and sub-cloned into the expression vector or SnapFast Pro,” continued Dr. Payne. “Following transient transfection into either CHO-K1 or HEK-293 cells, lysates and/or supernatants (depending on whether the protein is secreted) were subjected to high-throughput automated western blot. For both secreted and nonsecreted proteins, SnapFast Pro gave consistently high-level protein expression, in many cases showing high expression where no protein could be detected with the reference expression vector.”

Oxford Genetics is a specialist synthetic biology company focused on providing DNA, protein, virus, and cell-line design and development solutions. Two of the firm’s scientists can be seen working on a custom cell-line development project looking to optimize expression of a client’s gene of interest.



