March 1, 2012 (Vol. 32, No. 5)
Statistical and Biological Review of Transcriptome Data Made Possible by Interactive Report
Although bench scientists understand the value of using gene-expression microarray technology, the full value of this technology is frequently inaccessible due to persistent roadblocks in microarray data analysis.
Ingenuity® iReport™ from Ingenuity Systems is an interactive visual report for researchers who need to quickly understand gene-expression data, identify novel biological insights, and generate testable hypotheses to drive the experiment-to-experiment cycle.
This tutorial describes the best practices used by iReport for statistical and biological analysis of genomics data analysis, including novel tools for improved data visualization and insight discovery, resulting in an easy to use, one-step solution for statistical and biological analysis of gene expression data.
Pipeline for Statistical Analysis and Quality Control of Gene-Expression Data
Statistical analysis reduces instrument-level, per-sample measurements down to a set of significantly differentially expressed genes (DEGs). iReport includes a robust, fully automated statistical analysis pipeline for microarray data, based on industry-standard open-source components, primarily the widely used Bioconductor software for the R statistical programming language. It employs quantile normalization, RMA summarization and background correction, empirical Bayes methods for batch effect correction (ComBat), and empirical Bayes linear models for statistical analysis (Limma), which maximize its ability to detect differentially expressed genes. The pipeline controls type-I error using Benjamini and Hochberg’s False Discovery Rate when sufficient experimental replicates are available.
The statistical analysis pipeline helps the researcher control data by identifying outlier arrays, recognizing and evaluating the impact of batch effects, and alerting researchers to potential experimental design and statistical power problems (Figure 1).
The statistical pipeline currently supports gene-expression data from most human, mouse, and rat microarrays from Affymetrix, Illumina, and Agilent, as well as RNA-Seq data. It is extensible to other omics platforms, and support for qPCR is currently being developed.
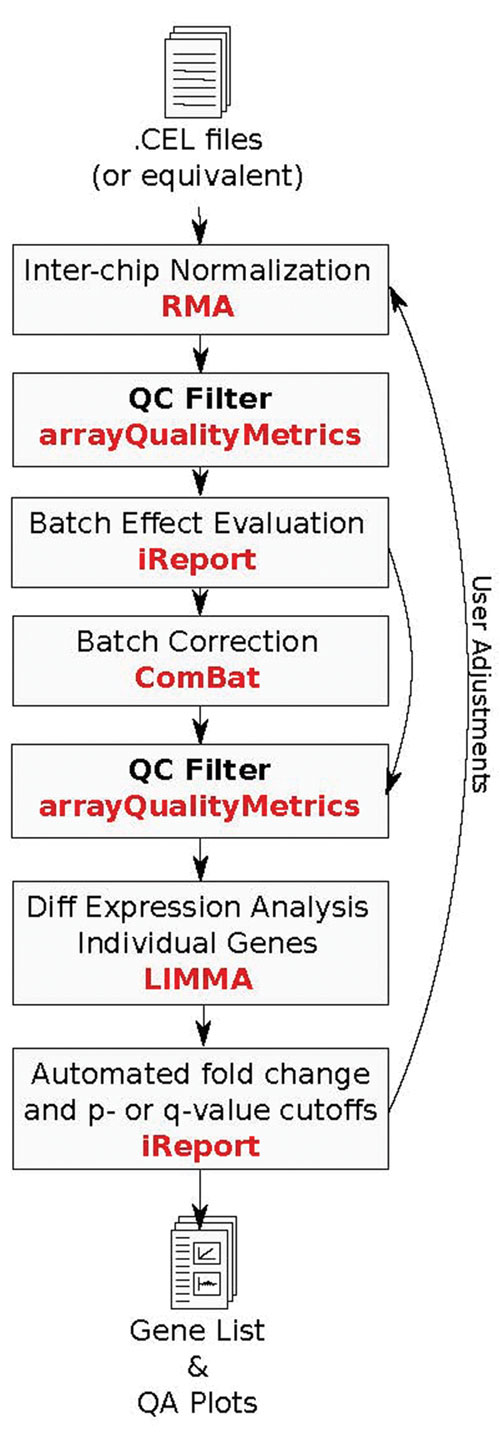

Figure 1. Schematic of statistical and quality control pipeline
Content and Algorithms for Biological Analysis of DEGs
Biological analysis of DEGs is often overlooked but is the critical step in getting rapid, complete value from microarray data and identifying insights for validation. Once a list of DEGs is identified by the iReport statistical pipeline, the DEGs are sent through a series of automated biological analyses, whose output forms the basis of iReport.
This biological analysis begins with a series of queries to the content in the Ingenuity Knowledge Base, a database of over 3.5 million highly descriptive findings curated from the biomedical literature and structured for computation. This content allows iReport to relate the expression of individual genes and gene sets to known and experimentally demonstrated information on signaling and metabolic pathways, biological processes, cellular functions, diseases, and experimentally demonstrated molecular interactions (both physical and functional).
iReport then identifies the subset of most significantly overrepresented biological and cellular functions, pathways, and diseases from those queries using standard bioinformatics enrichment tools, namely the Fisher’s Exact Test. (This process has been vetted in over 5,500 publications that cite IPA®, which uses the same statistical-enrichment process.) This helps researchers to understand putatively affected cellular functions and pathways and to identify potential markers of disease and key molecular interactions all without relying on bioinformatics support or labor-intensive literature searches.
Visualization Tools and Workflows for Exploration and Interpretation of Results
iReport removes data-analysis challenges by providing novel tools and workflows for visualizing, querying, and filtering the results. These tools help researchers perform downstream analysis based on biological criteria, interrogate their data from multiple angles, and quickly identify compelling genes for further study.
The Summary Page in iReport features “Top Results by Experimental Keyword” and an interactive volcano plot (Figure 2) that enables the user to quickly identify expected, biologically relevant sets of gene-expression changes. Clicking on one of the top results (for example, “Antiviral Response”) lets the user drill down in the volcano plot to see which DEGs are associated with that process and takes the user directly to the experimentally demonstrated evidence that implicates those DEGs in that biological process.
These features accomplish two critical goals of microarray data interpretation: 1) biologists can quickly find a set of DEGs that anchor on a relevant biological process or phenotype, and 2) biologists can clarify the strength of this association by accessing the literature evidence and experimental context substantiating that association in the Publications and Findings panel in each chapter of iReport.
By reducing the effort required to find an expected, relevant result, iReport Summary Page enables researchers to move on to explore unexpected, potentially novel insights about their samples.
To facilitate this exploration, we built The Wheel, which enables biologists to focus on individual or sets of DEGs based on their biological properties (such as molecular function, fold change, subcellular location, etc.) and known biological associations (pathways, diseases, cellular functions, etc.). The Wheel uses a biologist-friendly hierarchy of topics to quickly expand the investigation from a very specific result (e.g., antiviral response, 10 DEGs) to a broader topic within that same theme (e.g., regulation of immune response, 32 DEGs) (Figure 3).
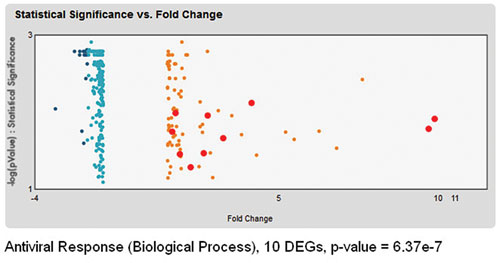
Figure 2. An interactive volcano plot of 300 DEGs, with 10 DEGs associated with the biological process antiviral response highlighted in red.
iReport was designed to enable researchers to find a compelling set of results and then follow that lead to understand whether a set of DEGs holds together in other biological contexts (pathways, interactions, etc). For example, iReport easily transitions this set of immune response genes into the molecular interactions chapter to understand whether, or how, these genes affect each other directly, either physically (e.g., protein interactions) or functionally (e.g., activation, inhibition, phosphorylation, etc.). This provides an opportunity to identify key regulatory points in the gene set.
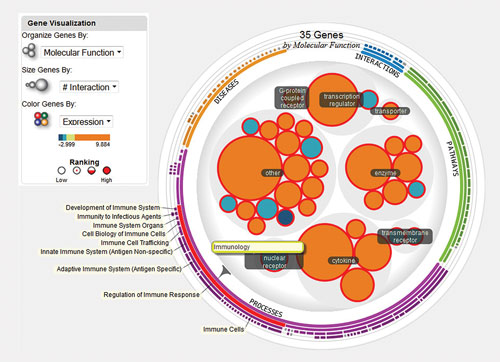
Figure 3. The Wheel, one of many organizational options in iReport, structures biological results into a hierarchy of topics, enabling users to quickly expand their view from a set of DEGs associated with a very specific result to a broader but related set of 32 DEGs involved in regulation of immune response. Here 32 DEGs involved in regulation of immune response are visually grouped by their molecular function, and sized by the number of molecular interactions with other DEGs.
Conclusions
By integrating an automated statistical and quality control pipeline, biological knowledge from the Ingenuity Knowledge Base, and novel workflow and user interface tools to interpret that knowledge, Ingenuity iReport for gene expression data makes best practices in statistical and biological analysis of microarray data accessible to bench scientists.
iReport identifies sets of gene-expression changes that are significantly different between samples, and maps those changes to cellular functions, phenotypes, pathways, and molecular interactions. This helps bench scientists find the biological stories that typically remain obscured when simply viewing lists of genes and expression changes, rapidly expand their knowledge of an experimental model, and identify a promising set of genes to interrogate with further experimentation.
Richard L. Halpert is associate staff software engineer and Megan E. Laurance, Ph.D. ([email protected]), is associate staff scientist product management at Ingenuity Systems. Web: www.ingenuity.com.



