June 1, 2009 (Vol. 29, No. 11)
Analyzing XpressWay Human Expression Data with OmniViz Visual Analytics Software
The sequencing of the human genome has identified a vast number of potentially interesting targets for drug development. The challenge is to evaluate these targets and select those that are relevant to human disease. An important step for the evaluation process is to identify where these targets are expressed, and to link this expression to potential therapeutic uses. This evaluation is greatly aided by the provision of high-quality gene-expression data and sophisticated software tools for identifying targets with interesting expression patterns.
In this article, we describe the combination of Asterand’s XpressWay® human gene-expression data with BioWisdom’s visual analytics software, OmniViz. The XpressWay dataset consists of more than 2,000 gene-expression profiles, the majority representing proteins that are potentially tractable drug targets. The OmniViz software provides easily interpreted visualizations and a suite of interactive tools that allows scientists to relate divergent data to uncover previously unknown associations and answer key scientific questions.
By analyzing XpressWay data using OmniViz, the user can quickly select and stratify genes of interest, providing a range of benefits including: increasing confidence in a therapeutic approach, selecting the best targets for drug development, identifying new targets for an indication, exploiting opportunities to switch indications, identifying potential side effect liabilities, predicting translation of preclinical to clinical drug effects, and gaining mechanistic insight by looking at the association of gene-expression profiles.
A Whole-Body Scan
XpressWay profiles consist of target expression in 72 different human tissues from three different donors (216 samples in total). These tissues are chosen to be representative of the major organ systems of the body, and comprise many sub-dissected regions. All the tissues are assessed as pathologically normal by pathologists. The breadth of tissues used for XpressWay profiles holds advantages over other gene-expression datasets that often consist of smaller sets of tissue.
Total RNA is isolated from the tissues using standard methodologies, and has to pass several QC criteria before being considered suitable for expression profiling.
Sensitive and Standardized Methodology
The XpressWay gene-expression profiles are generated using quantitative real-time PCR, a method that allows detection of absolute mRNA levels in tissues. This technique is different from other methodologies such as chip-based assays, which measure relative abundance of mRNA.
The target gene is multiplexed in the same well as GAPDH, which is used to confirm successful amplification. PCR amplification curves are analyzed to yield threshold values, and these are used to determine the starting mRNA copy number of both target and GAPDH genes by interpolation from a global standard curve. Rigorous pass/fail criteria are applied in order to ensure the quality of the data.
Gene Annotation
XpressWay gene-expression data is provided with BioWisdom annotation around gene synonyms, disease, and process information. The provision of gene synonyms ensures that the user can find the gene of interest, and also facilitates connections to other data sources. For example, it is possible to send the synonyms out to PubMed in order to retrieve all the relevant publications around the target of interest. These documents can easily be imported into OmniViz, and, by employing the analytical tools available, the important themes of the documents, and hence the nature of the target, can be revealed.
The inclusion of disease and process information provides context to the gene-expression data, and allows the user to search for interesting targets by virtue of their functional role.
View the Expression Landscape
Using OmniViz to analyze XpressWay data, it is possible to get an overview of the gene-expression patterns of all ~2,000 genes (Figure 1), and then to quickly identify those gene-expression profiles that possess features of interest. For example, if you are interested in developing a drug for a CNS disorder, you may be interested in the targets (see orange arrows on the left hand side of Figure 1) that have high expression (red) in nervous tissue and low expression (blue) in peripheral tissues.
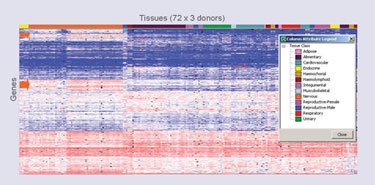
Figure 1. OmniViz treescape visualization provides an overview of all XpressWay gene-expression patterns (~2,000 genes).
Assess Individual Targets
Individual gene-expression profiles across the panel of 72 tissues can provide valuable information about potential drug action in those tissues. Within OmniViz, users can retrieve the expression profile for a drug target with a text query for the target name. The example target expression profile shown (Figure 2, ATP1A3) highlights high copy numbers in CNS and heart samples. Depending on the approach being used, this expression pattern may represent opportunities for the development of drugs for CNS or cardiac disorders, or may suggest potential side-effect liabilities in these areas. Knowledge of these data will help users to make rational decisions on the future of a drug developed against this target.
In addition to searching for genes by their name or by their functional role, OmniViz provides the capability to make complex numeric queries of the data. For example, users can identify targets with a particular expression profile, or profiles that are most similar to a chosen target.
As an example, it is possible to search for genes with high expression in hippocampus as potential targets for Alzheimer’s disease (AD). It is known that the hippocampus atrophies in AD, and that it plays a role in processes that are affected in AD (e.g., memory). Within OmniViz, the user can to set up a search for genes that have greater than 10,000 mRNA copies in the hippocampus, but less than 100 mRNA copies in most of the other tissues. This particular query retrieves mGluR5-splice variant 2 (mGluR5-2). As an additional step, it is possible to retrieve other gene profiles with the closest similarity to mGluR5-2, by correlation or Euclidean matching (Figure 3).
Furthermore, it is possible to assess the relevance of the retrieved targets by viewing diseases and processes associated with them. As can be seen from the Table, mGluR5-2 and its closest neighbors play a role in several processes (e.g. synaptic transmission, synaptic plasticity) that may be relevant to AD. Therefore, these may be interesting targets to progress for this disorder.
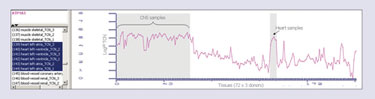
Figure 2. OmniViz profile plot shows expression of a single gene, ATP1A3, across the panel of tissues.
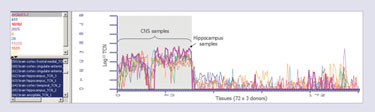
Figure 3. The expression profile of mGluR5-2 (bold pink line) and the 10 genes identified to have the most similar profiles
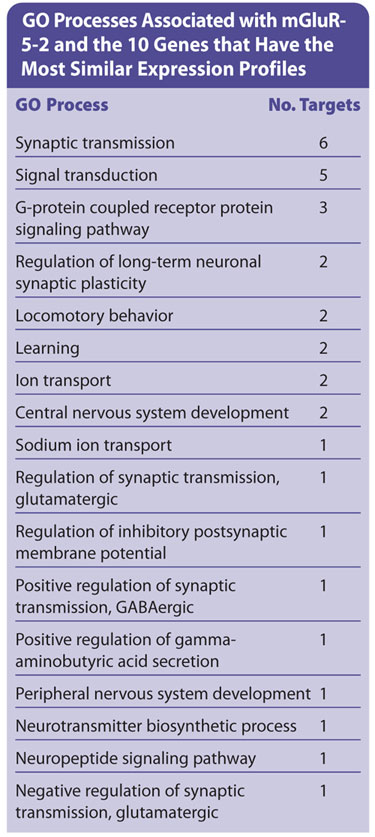
GO processes associated with mGluR5-2 and the 10 genes that have the most similar expression profiles
Compare with Other Datasets
With OmniViz, users can easily import other datasets (internal or publicly available) to compare with XpressWay gene-expression data. These datasets can be linked to or merged with the XpressWay dataset using common identifiers (e.g. gene ID).
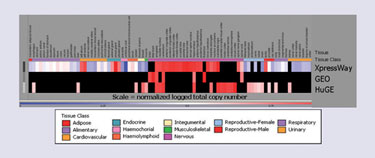
As an example, Figure 4 compares the expression profiles for the target, ATP1A3, in XpressWay, GEO, and HuGE datasets.

Figure 4. OmniViz treescape visualization compares expression patterns of ATP1A3 across XpressWay, GEO, and HuGE datasets.
Asset for Drug Development
The XpressWay dataset holds several advantages over other gene-expression datasets. First, the use of sensitive and standardized methodology, and the wide range of tissues used, allows users to have confidence in the presence (or not) of a target in a tissue, and the difference in expression between tissues.
Second, the enhanced interrogation provided by OmniViz allows users to see previously unknown expression patterns, to find associations between targets, and to find targets with particular features of interest. Finally, understanding of the data is augmented due to disease and process annotation, and the facility to bring related information (e.g., other gene expression data or PubMed articles on a target) into OmniViz for analysis.
Nicola C. Day, Ph.D. ([email protected]), is a healthcare
consultant at BioWisdom. Web: www.biowisdom.com. Sandra Williams, Ph.D., is department head, genomics, Asterand. Web: www.asterand.com.