September 1, 2010 (Vol. 30, No. 15)
Innovative Tools for Screening Hundreds of miRNAs and Subsequent Functional Studies
MiRNAs (microRNAs) are 20–23 nucleotide small RNA molecules that play an important role in post-transcriptional gene regulation in a wide variety of biological processes including development, disease, and apoptosis.
Scientists must screen hundreds of miRNAs to identify those involved in their pathway or disease of interest. This can be achieved using PCR arrays that allow quantification of hundreds of miRNAs in one experiment. Once screened, further analysis can be performed to verify the results, perform more in-depth functional studies, or get a better understanding of miRNA biogenesis.
RT2 miRNA PCR Arrays from Qiagen are assays in a 96- or 384-well plate format. Each well of these arrays contains a specific and sensitive primer for one miRNA. This allows high-throughput quantification and profiling of multiple miRNAs in a single RT-PCR run. Data from different disease states, development stages, or tissue types can be compared to identify up- or downregulated molecules.
RT2 miRNA PCR Arrays target the entire miRNome or subsets of miRNAs that are involved in biological pathways, processes such as inflammation and cell differentiation, or diseases such as cancer. The assays can distinguish between miRNA family members with single nucleotide mismatches as well as differentiate mature miRNAs from their precursors.
Screening of hundreds of miRNAs can be performed in a single experiment. As the first step, the miRNAs are converted to cDNA by reverse transcription and universal tailing. The master mix with the components for the RT-PCR reaction is then mixed with the sample and added into the well. After mixing, the PCR run is performed in a real-time PCR instrument and data is collected.
Integrated, web-based software performs gene-expression analysis from uploaded raw data and delivers results in multiple formats, enabling easy interpretation. Differences between different states can be calculated from raw CT values normalized to a panel of housekeeping small nuclear RNAs.
Quantification of the expression of multiple miRNAs in a cell or tissue type under defined conditions provides an miRNA profile or a snapshot of miRNA expression that can be used to elucidate the role of miRNAs in regulatory pathways or their misregulation in disease. A profile for colon cancer is shown in Figure 1. In this example, many of the miRNAs profiled appear to be upregulated in cancer cells.
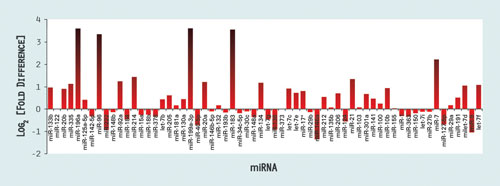
Figure 1. Example of an RT2 miRNA PCR Array assay panel that is specific for 88 cancer-related human miRNAs: The profile shows the expression changes in colon cancer in comparison to normal tissue.
Focusing on miRNA Biogenesis
miRNA-profiling experiments can give a high-level overview of miRNA expression status under defined conditions. Following these experiments, researchers may wish to proceed by selecting smaller numbers of miRNAs with interesting expression profiles for further study of biogenesis and function.
Quantification of miRNA precursors alongside mature miRNAs can provide important insights into the pathways from transcription to maturation. Real-time PCR can be used to perform this type of analysis (e.g., using the miScript PCR System with assays specific for either mature or precursor miRNA).
Comparison of mature and precursor miRNA levels can reveal variations in relative levels of mature and precursor miRNA for different miRNAs, or mechanisms whereby mature miRNAs are processed from different precursor miRNAs under different conditions.
The Next Step: Functional Studies
After miRNAs involved in a disease or pathway of interest have been identified by a PCR array, functional studies can provide further insights, such as identifying miRNA targets and binding sites as well as pinpointing the role of a particular miRNA in a pathway or process.
This type of investigation can be performed using strategies that involve transfection of functional molecules, such as synthetic miRNA mimics, miRNA inhibitors, or miRNA target protectors. A mimic behaves like a mature endogenous miRNA after its transfection, while miRNA inhibitors bind to an miRNA and inhibit its function.
Transfection of a mimic or inhibitor can be followed by phenotype or expression analysis. Target protectors allow the researcher to interfere with the interaction between a given miRNA and its specific target, leaving other targets of the miRNA unaffected. This is achieved because the target protector binds specifically to the miRNA-binding site in the 3´ UTR of an mRNA of one gene preventing miRNA binding and thereby preventing miRNA-mediated regulation (Figure 2).
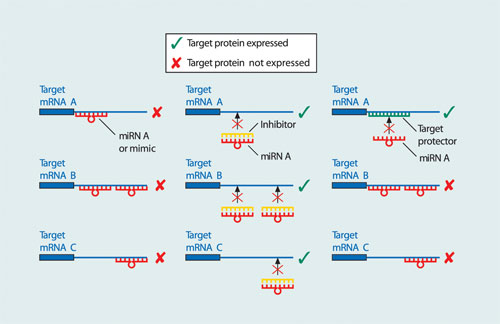
Figure 2. Mimics, inhibitors, and target protectors principle: (A) miRNAs or mimics bind to their target mRNAs, causing downregulation of expression of all the targets. (B) miRNA inhibitors bind to the miRNA of interest, resulting in the expression of all gene targets. (C) In contrast, target protectors bind to a specific miRNA binding site, resulting in expression of that miRNA target only, while any other targets remain downregulated.
miRNA Target Verification
Target prediction algorithms allow identification of putative miRNA-binding sites for genes of interest (e.g., TargetScan). This can be verified in vitro using target protectors.
An example is the DNMT3A gene that encodes DNA methyltransferase 3, which is involved in development and differentiation. TargetScan predicted a single binding site for the miRNA miR-29a in the 3´ UTR of DNMT3A. Transfection of a miR-29a mimic led to downregulation of DNMT3A as shown by real-time RT-PCR, indicating that miR-29a downregulates DNMT3A expression at the transcriptional level.
Co-transfection of a target protector designed for the miR-29a binding site resulted in an increase in DNMT3A expression. These results provided convincing evidence that DNMT3A is negatively regulated by miR-29a via the binding site predicted by TargetScan (Figure 3).
Innovative tools such as the RT2 miRNA PCR arrays from Qiagen enable scientists to screen hundreds of miRNAs in one experiment, facilitating the understanding of the role of miRNAs in their area of interest. The experiment is simple to perform and provides sensitive, reproducible, and reliable results to accurately profile multiple genes simultaneously.
Verification of the results and in-depth functional analysis can then be undertaken by other techniques such as transfection with miRNA mimics and inhibitors and subsequent expression analysis.
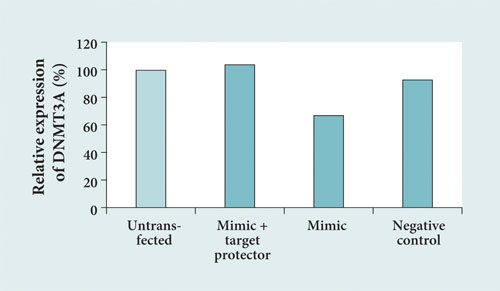
Figure 3. Target verification: MCF-7 cells were co-transfected with an miR-29a mimic plus a nonbinding target protector as negative control or miR-29a plus a target protector specific for the putative miR-29a binding site of DNMT3A (mimic + target protector). After transfection, expression analysis of DNMT3A was performed by real-time RT-PCR with untransfected cells also being analyzed. Negative control: control siRNA and control target protector.
Subu Yerramilli, Ph.D. ([email protected]), is associate director of R&D, Martin Kreutz is research scientist, Yexun Wang, Ph.D., is associate director of R&D, and Elizabeth Scanlan, Ph.D., is technical and marketing writer at Qiagen.



