The neurotrauma research community is a well-established field that has developed and characterized preclinical animal models to better understand traumatic injury pathology and assess the efficacy of therapeutic interventions. Using a mouse model enabled a team to conduct biomarker discovery where the complexity and evolution of the injury pathology was progressing. The team plans to expand its studies to look for these same biomarkers in humans.
A scientific team led by Arizona State University (ASU) performed a biopanning search to reveal several biomarkers of acute and chronic traumatic brain injury.
The researchers reported the discovery of some of the first detailed molecular clues associated with traumatic brain injury (TBI). The disorder is a growing public health concern, affecting more than 1.7 million Americans at an estimated annual cost of $76.5 billion dollars. It is a leading cause of death and disability for children and young adults in industrialized countries, and people who experience TBI are more likely to develop severe, long-term cognitive and behavioral deficits.
“Unfortunately, the molecular and cellular mechanisms of TBI injury progression are multifaceted and have yet to be fully elucidated,” said Sarah Stabenfeldt, PhD, an ASU professor and the leader and corresponding author of the study (“Uncovering temporospatial sensitive TBI targeting strategies via in vivo phage display”), which appears in Science Advances. “Consequently, this complexity affects the development of diagnostic and treatment options for TBI; the goal of our research was to address these current limitations.”
Their approach was to perform a “biopanning” search to reveal several biomarkers, identified directly after immediately after the injury event (the acute phase), and also the long-term consequences (the chronic phase) of TBI.
“For TBI, the pathology evolves and changes over time, meaning that a single protein or receptor may be upregulated at one phase of the injury, but not two weeks later,” continued Stabenfeldt. “This dynamic environment makes developing a successful targeting strategy complicated.”
To overcome these limitations, the Stabenfeldt team used a mouse model to begin to study the root causes of TBI by identifying biomarkers.

“The heterogeneous pathophysiology of TBI is a barrier to advancing diagnostics and therapeutics, including targeted drug delivery. We used a unique discovery pipeline to identify novel targeting motifs that recognize specific temporal phases of TBI pathology. This pipeline combined in vivo biopanning with domain antibody (dAb) phage display, next-generation sequencing analysis, and peptide synthesis,” wrote the investigators.
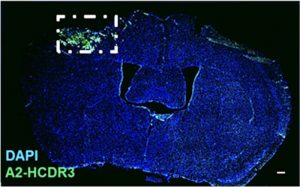
“We identified targeting motifs based on the complementarity-determining region 3 structure of dAbs for acute (1-day post-injury) and subacute (7 days post-injury) post-injury time points in a preclinical TBI model (controlled cortical impact). Bioreactivity and temporal sensitivity of the targeting motifs were validated via immunohistochemistry. Immunoprecipitation–mass spectrometry indicated that the acute TBI targeting motif recognized targets associated with metabolic and mitochondrial dysfunction, whereas the subacute TBI motif was largely associated with neurodegenerative processes.
“This pipeline successfully discovered temporally specific TBI targeting motif/epitope pairs that will serve as the foundation for the next-generation targeted TBI therapeutics and diagnostics.”
Taking a bottom-up approach
Scientists can often begin to design therapeutic agents or diagnostic devices based on biomarker discovery. Stabenfeldt’s team relied on a bottom-up approach.
“Top-down discovery methods are focused on assessing candidate biomarkers based on their known involvement in the condition of interest,” said study first author Briana Martinez, a recent PhD graduate in Stabenfeldt’s lab. “In contrast, a bottom-up method analyzes changes in tissue composition and finds a way to connect those changes to the condition. It’s a more unbiased approach but can be risky because you can possibly identify markers that are not specific to the condition or pathology of interest.”
Next, they utilized several biopanning tools and techniques to identify and capture molecules, including a bait technique for fishing out potential target molecules known as a phage-display system. In addition, the team used high-speed DNA sequencing to identify protein targets within the genome, and mass spectrometers to sequence the peptide fragments from the phase display experiments.
One further roadblock to discovery is the blood-brain barrier (BBB) between the vascular and brain tissue.
“In a healthy individual, the BBB tightly regulates nutrient and waste exchange from the blood to the brain and vice versa, essentially compartmentalizing the brain/central nervous system,” explained Stabenfeldt. “However, this barrier also complicates drug delivery to the brain so that most molecules/drugs do not passively cross this barrier; therefore, the drug delivery field has sought out ways to modulate both entry and delivery mechanisms. Similarly, for blood-based biomarkers for TBI or other neurodegenerative diseases, specificity to the pathology and transfer of the molecule (if it originates in the brain) from the brain to blood is a challenge.”
When a TBI occurs, the initial injury can disrupt the BBB, which triggers a cascade of cell death, torn, disrupted tissues, and debris. The long-term injury causes inflammation and swelling, and results in the immune response to spring into action, but also can lead to an impairment of the brain’s energy sources, or can choke off the brain’s blood supply, leading to more neuronal cell death and permanent disability.
One main advantage of their suite of experimental tools and techniques of the phage display system is that the molecules and potential biomarkers identified are small enough to slip through the tiny holes within the meshwork of the BBB—thus, opening the way to therapeutics based on these molecules.
“Our study leverages the sensitivity and specificity of phage to discover novel targeting motifs,” said Stabenfeldt. “The combination of phage and NGS [next-generation sequencing] has been used previously, thereby leveraging bioinformatic analysis. The unique contribution of our study is putting all of these tools together specifically for an in vivo model of TBI.”
They found a suite of unique biomarkers associated with only the acute or chronic phases of TBI. In the acute phase, the TBI targeting motif recognized targets associated with mainly metabolic and mitochondrial dysfunction, whereas, the chronic TBI motif was largely associated with neurodegenerative processes.
Next, the group plans to further its collaborations with ASU’s clinical partners and expand their studies to begin to look for these same molecules in human samples.
For additional information on traumatic brain injuries, see GEN: “Neuron Regeneration Following Brain Injury Activated by Neuron and Glia Connection” and “Machine Learning Optimizes Outcome Prediction in Traumatic Brain Injuries.” Also see: “Incidence, causes and consequences of moderate and severe traumatic brain injury as determined by Abbreviated Injury Score in the Netherlands” and “Traumatic brain injury patient characteristics and outcomes in Lebanon: a multicenter retrospective cohort study.”