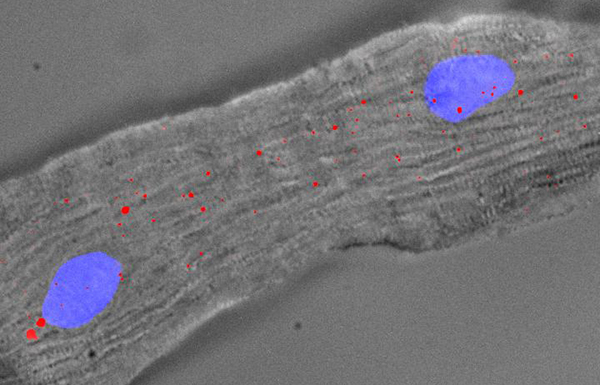
Heart tissue is notoriously poor at self-repair, but it isn’t entirely helpless. It’s just super-slow because mature, developed hearts retain very few heart cells that hold regeneration potential. For example, in adult mouse or human heart, new cardiomyocytes (CMs) are generated at rates of about 1%—not enough to repair damage from cardiovascular disease. But what if those rates could be increased? To explore this possibility, scientists based at A*STAR's Genome Institute of Singapore (GIS) and the National University Health System (NUHS) looked at cardiac cell subpopulations and how they differed in gene expression.
The scientists examined CMs from mouse and human hearts—from both failing and nonfailing adult hearts—using single nuclear RNA-sequencing. The scientists found that in cardiac subpopulations, the upregulation of cell cycle activators and inhibitors consequent to the stress-response varied. In particular, the scientists found that two long noncoding ribonucleic acids (ncRNAs) appear to regulate genes controlling the ability of heart cells to undergo repair or regeneration.
Details of this work appeared August 9 in the journal Nature Communications, in an article entitled “Single Cardiomyocyte Nuclear Transcriptomes Reveal a lincRNA-Regulated De-Differentiation and Cell Cycle Stress-Response In Vivo.” The article describes how the GIS and NUHS scientists used weighted gene co-expression network analysis to explore whether heterogeneity of the stress-response gene expression changes among the larger majority of cell cycle-arrested CMs would uncover a subpopulation that could be motivated to re-enter cell cycle.
“We noted distinct sub-populations of CMs and uncovered gene regulatory networks specific for each sub-population, displaying specific sub-group upregulation of cell cycle, and de-differentiation genes in the disease stress response,” wrote the article’s authors. “We further identified LINCMs [long intergenic noncoding RNA in nuclei of CMs] that occupy key nodal hubs in gene regulatory networks, and validated that knockdown of nodal LINCMs (namely, Gas5 and Sghrt) leads to corresponding changes in the expression of co-regulated network genes, including those known to control CM cell cycle.”
Essentially, the scientists discovered that a unique subpopulation of heart cells in diseased hearts activates gene programs related to heart cell division, uncovering the gene expression heterogeneity of diseased heart cells for the first time. In addition, they also found the “brakes” that prevent heart cells from dividing and thus self-healing. Targeting these “brakes” could help trigger the repair and regeneration of heart cells.
“There has always been a suspicion that the heart holds the key to its own healing, regenerative and repair capability. But that ability seems to become blocked as soon as the heart is past its developmental stage,” explained Prof. Roger Foo, the study's senior author and a principal investigator at both the GIS and the NUHS. “Our findings point to this potential block that when lifted, may allow the heart to heal itself.”
“In contrast to a skin wound where the scab falls off and new skin grows over, the heart lacks such a capability to self-heal, and suffers a permanent scar instead. If the heart can be motivated to heal like the skin, consequences of a heart attack would be banished forever.”
“We confirmed that LINCM3 (also called Gas5) and LINCM9 (previously annotated 1810058i24Rik, which we now call “Singheart,” Sghrt) were upregulated in TAC CMs [transverse aortic constriction cardiomyocytes],” detailed the article’s authors, “while LINCM5 was downregulated in TAC CMs as compared to Sham CMs [i.e., in sham-operated mice].”
“The gene regulatory networks and LINCMs derived from our single nuclear RNA-seq now serve as an invaluable resource for identifying key endogenous regulators of cardiac regeneration. Meanwhile, the mitotic potential found in a substantial subset of adult CM in vivo raises the hope that targeting negative regulators of CM proliferation may one day lead to successful cardiac regenerative therapy.”