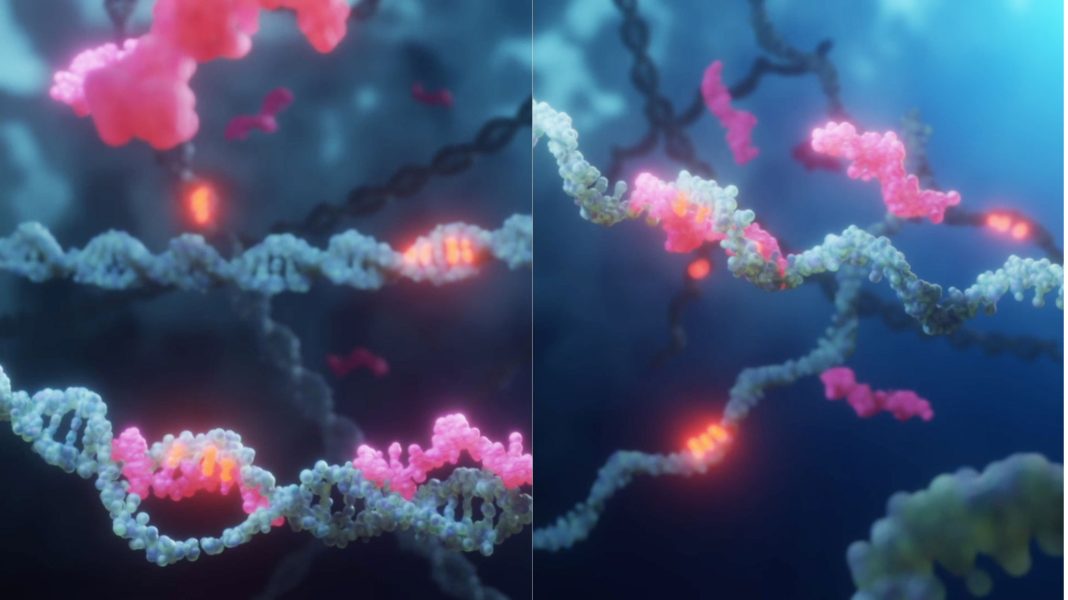
The pharmaceutical industry is equipping itself to target genetic defects directly. The pioneer generation of genetic medicines demonstrated the viability of the approach, but their applicability was restricted by their inability to target the genome at the base level. Also, they had limited biodistribution, tolerability, selectivity, and durability.

NeuBase Therapeutics
“If I were to encapsulate my sense of first-generation precision genetic medicines, I would say that they’re exciting because they work,” says Dietrich Stephan, PhD, CEO at NeuBase Therapeutics. However, he adds that they have a few downsides: “We’re left with treating the liver after systemic administration or injecting drugs locally. Early genetic medicines are often large, negatively charged, and trigger the immune system, limiting tolerability. They are usually not selective enough to see point mutations.”
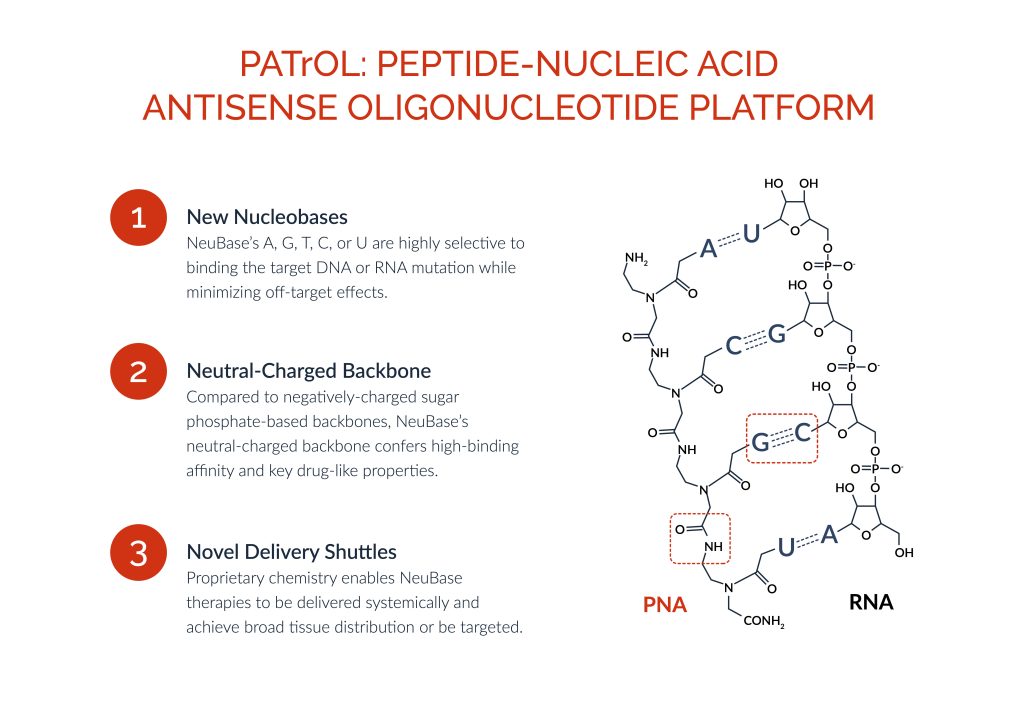
NeuBase is using a proprietary drug discovery platform—the Peptide-nucleic acid AnTisense OLigonucleotide (PATrOL) platform—to develop a class of small synthetic genetic medicines that address gain-of-function, loss-of-function, and change-of-function mutations in the human genome. “In one technology platform, we believe we have the capability to consolidate the nascent and fragmented genetic medicines industry,” Stephan declares.
He acknowledges the genetic medicine industry has made great strides but emphasizes these are mostly in drugging single-stranded RNA. DNA is a more difficult target. Companies like Alnylam and Ionis have pioneered degrading gain-of-function mRNAs while others like Moderna and BioNtech have taken the lead in replacing mRNAs.
Stephan believes NeuBase has found a way around the major limitations of first-generation drugs: “Modified peptide nucleic acids (PNAs) have the highest binding affinity per nucleobase of any technology combined with exquisite selectivity to uniquely drug the double-stranded genome. PNAs are broadly biodistributed after systemic injection, non-immunogenic, well tolerated, highly sequence selective, enduring, manufacturable, and scalable.”
Methods driving drug discovery
Whereas the availability of genome and variant databases provided the launchpad for genetic medicines, a battery of technical advances continues to propel the field forward.
NeuBase is interested in technical advances to enable the pharmaceutical application of PNAs. These advances include parallelized synthetic peptide production, computational modules to capture and analyze biological data, and high-throughput in vivo pharmacokinetics and pharmacodynamics assays. Stephan believes that these are the top methods facilitating drug discovery.
Computational tools to store and query big data and design medicines that have minimal off-target engagements across the genome and transcriptome are transforming drug discovery. “Deep learning illuminates structure-activity relationships,” Stephan says. But more important and direct, he adds, could be the effect of commoditized computational solutions that allow biologists to capture data in machine-readable formats, enabling iterative predictive learning.
Despite significant headway in high-throughput manufacturing, Stephan observes bottlenecks in products ranging from mRNA vaccines to antibody-drug conjugates, oligos, and peptides. He notes, “Improving coupling efficiencies of parallelized synthetic peptide synthesis without the need for purification and at a scale to enable in vitro screening will improve our ability to do genome-wide activity screens.”
Root of the problem
Traditionally, discovering new drugs has involved targeting the workhorses of the cell—proteins. However, proteins are large, dynamic, short lived, and steps removed from the genetic cause. “We start with root causality—sequence changes in genes,” Stephan emphasizes.
Stephan’s team addresses genetic overexpression by inhibiting transcription and loss-of-function mutations by increasing gene output. NeuBase’s platform is also adept at editing genes without double-stranded breaks and with layers of selectivity that minimize off-target edits. “Our technology,” he asserts, “has been used to perform in vivo editing with perfect fidelity and tolerability in preclinical models of cystic fibrosis and β-thalassemia.”
Stephan says that grappling with pathogenesis at the protein level is hard. “Proteins are complex structures, constantly moving and changing shapes,” he remarks. “Until a few months ago, we really didn’t know how to model them. Throwing small molecules at them in a high-throughput, random manner is expensive, time consuming, and has a low probability of success.”
Complications in targeting proteins not only escalate drug prices, since companies must recoup upfront costs, but low success rates have also created a burden of untreatable diseases. “Every disease is genetic,” Stephan insists. “The promise of sequencing the genome was to figure out all root causes, get causal variant information to patients, and drug at the base level to resolve the disease.”
Neutral minimalism and delivery shuttles
Biodistribution is the first challenge that Stephan’s team took on. Size and charge limited the biodistribution of siRNA, ASO, and protein therapeutics. NeuBase’s genetic pharmacophores—the PNAs—are minimalistic structures about 5 kD in size that look like oligonucleotides but have a synthetic polyamide backbone. Furthermore, systemic administration of negatively charged genetic medicines cause aggregation and elimination through scavenger receptors in the liver. Designing for neutrality prevented NeuBase’s drugs from being prematurely excreted.

The small, neutral structure linked to the delivery shuttle forms an emulsion with phospholipid bilayers that diffuses down the membrane’s electrochemical gradient. “We developed a proprietary delivery shuttle that gets past the liver and sticks to cell membranes,” Stephan explains. This increases the drug’s half-life in circulation and enables it to bypass endosomal entrapment, an issue that plagued first-generation medicines.
Once in the cytoplasm, PNAs diffuse through nuclear pores, whether the delivery shuttle is jettisoned by means of a cleavable linker or left on, resulting in target engagement within 24 hours after systemic dosing. Nuclear localization tags don’t improve potency further.
Balancing biodistribution and selectivity
PNAs balance broad biodistribution with selective function in cells that express the disease gene. “Our strategy,” Stephan relates, “has been to get the drug into every cell and lean on improved selectivity of the polyamide backbone.”
The semirigid backbone of the drug prevents it from engaging with mismatched loci. “PNAs have exquisite sequence selectivity down to the single base,” he adds. This minimizes off-target bindings and adverse events.
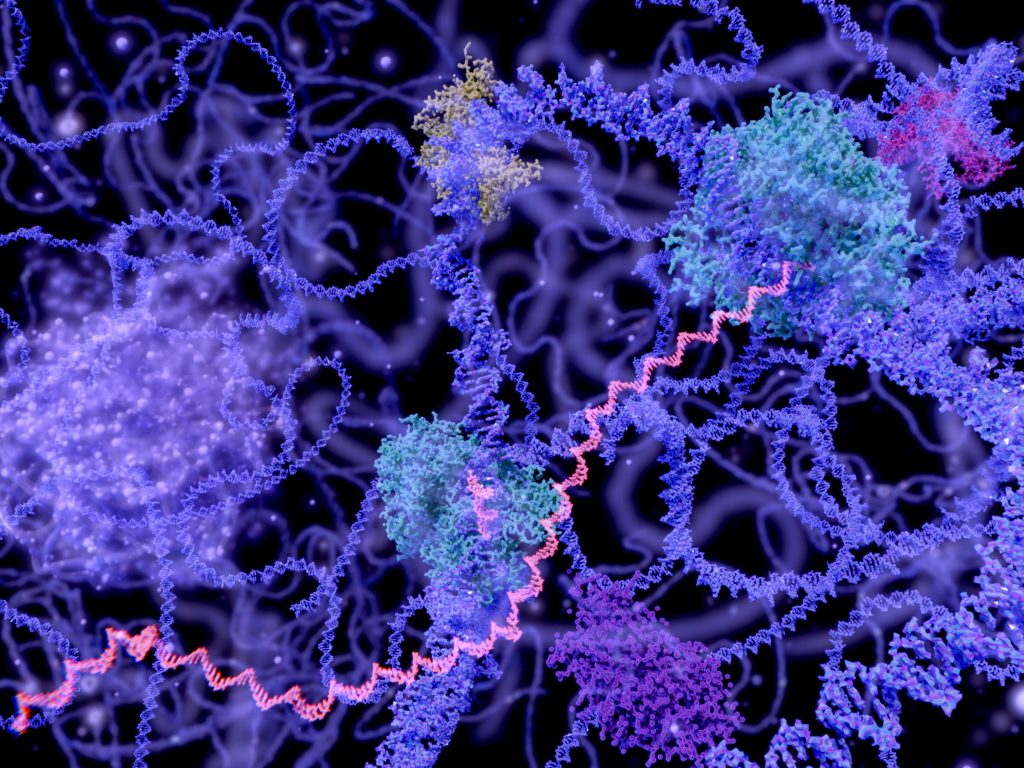
Nuclear chromatin is a dynamic protein-rich coil with inactive, densely packed regions that are more inaccessible than regions being transcribed. “As the double helix breathes and opens when it’s expressed and allows RNA polymerase to come through, that’s the opportunity our drugs take to invade and bind,” Stephan details. “When the DNA tries to close again, it can’t because the PNA binding affinity is so high.”
That PNAs cannot lock onto a locus until its target has been unwound is not a negative. “If a gene is deactivated in a cell, it’s not able to cause the disease in that cell,” Stephan points out. “In most cases, that works to our advantage.”
Toxicity and tolerability
Nonhuman primate pharmacokinetic profiles of the new PNAs confirm a half-life of two hours in circulation after systemic administration. The drugs are taken up by all tissues—including neurons in deep brain structures—where they remain for a half-life of approximately three months and are renally excreted intact. They are resistant to protease and nuclease digestion. “The only thing that we worry about from a tolerability perspective is interaction of the drug with identical sequences elsewhere in the genome,” Stephan notes. “So, we spend a lot of time on the bioinformatics up front to design drugs with unique targets.”
Factors that enhance PNA tolerability include their inability to aggregate or trigger inflammation through an innate immune response (by activating TLR9 receptors resulting in a TH1 response or by activating the complement pathway) or an acquired B- or T-cell immune response. Considering their non-immunogenicity, their remaining in tissues for prolonged periods poses no problem. The literature suggests that the tolerable doses of first-generation PNAs are up to 100-fold greater than their effective doses.
“We’ve seen no tolerability issues at or above effective doses,” Stephan reports. “That and the pharmacology has given us the confidence to nominate our first clinical candidate scale-up for CMC manufacturing. We’ll file our first IND this year, and scale thereafter.”
Durability
First-generation viral gene therapies have limited durability of response and cannot be administered repeatedly due to their immunogenicity. In its initial attempts at targeting the genome, NeuBase is focusing on applications that require temporary drug exposure. The durability of the pharmacologic effect is on par with the drug’s half-life in tissues. Specific disease scenarios could call for monthly to quarterly doses to replenish the pharmacologic effect, but that is not a disadvantage, Stephan believes.
“We put a drug molecule on a DNA locus and address root causality, but at some point, either because of cell division or Koff rates, the drug will come off and be renally excreted,” he says. PNAs that execute gene editing, however, won’t require repeated dosing.
PNA-based gene editing is inherently different from CRISPR-based approaches. “Our drugs pry open the double helix to allow a DNA guide strand to engage on the mutation,” Stephan says. “Without double-strand breaks, we recruit endogenous mismatch repair enzymes that have evolved over millennia to have the requisite fidelity to repair DNA mutations.” The PNA in this case acts like a carjack. Once the mutation is fixed, the PNA comes off and is renally excreted, leaving the mutation permanently fixed.
Scalability
Stephan indicates that NeuBase has built scalability into a modular manufacturing pipeline: “Small molecules can be routinely manufactured at the ton scale using established CMOs across the globe in a GMP setting. Those then get snapped together like Lego blocks using standard synthetic peptide manufacturing capabilities. There are any number of CMOs that do GMP peptide synthesis at scale. When we combine them, we have manufacturing capability for a drug without having to build our own factory.”
Like Moderna and Pfizer, each of which succeeds in manufacturing at population scale by focusing on one vaccine, NeuBase banks on standard manufacturing of identical components of their drugs to allow it to optimize the single variable: the oligo. “We have true scalability because we use the same delivery shuttle for every medicine we build,” Stephan asserts. “We can put that on rails from a manufacturing perspective. The basic building blocks of the polyamide scaffold are identical. All we’re doing is shuffling nuclear bases on the oligo.”
The comparable pharmacokinetics and biodistribution of these drugs on account of their identical delivery shuttle will clarify their route, dose, and frequency of administration, eliminating considerable efforts in developing individual drugs. Stephan predicts, “At some point after our first few drugs, we’ll have a standard toxicology package that will allow us to streamline our preclinical and Phase I work with the only delta being the potential off-target engagements, which we believe we can engineer out by using computational methods and leveraging the exquisite selectivity of the chemistry.”
Expanding realms of application
Although NeuBase’s initial discoveries involve monogenic diseases, PNAs can be combined through cleavable links to address more complex scenarios. “We already know we can create molecules that engage two specific sequences within the genome,” Stephan notes. These bispecific drugs combine different PNAs that engage neighboring sequences and are held together by linkers. Ultimately, Stephan wants to steer NeuBase toward expanding the range of applications of genetic medicines to complex diseases such as cancers, infectious diseases, and chronic conditions where multiple mutations work in combination with environmental triggers.