The cell is the basic functional unit of biology, and mechanisms of disease are rooted in cellular dysfunction. To understand the complex biology of disease, we need to understand biology at its fundamental unit of resolution. However, current approaches to diagnose and treat disease have been based upon low-resolution data. Thus, a new foundation is needed.
The right way to understand biology is cell by cell

We currently understand only a small fraction of complex biology. This existing state of knowledge has not provided an adequate foundation to understand the underlying mechanisms of disease, and thus, the discovery of ways to diagnose and treat complex diseases has been limited. Exponential advancements are needed to fuel the kinds of scientific discovery that will dramatically improve human health.
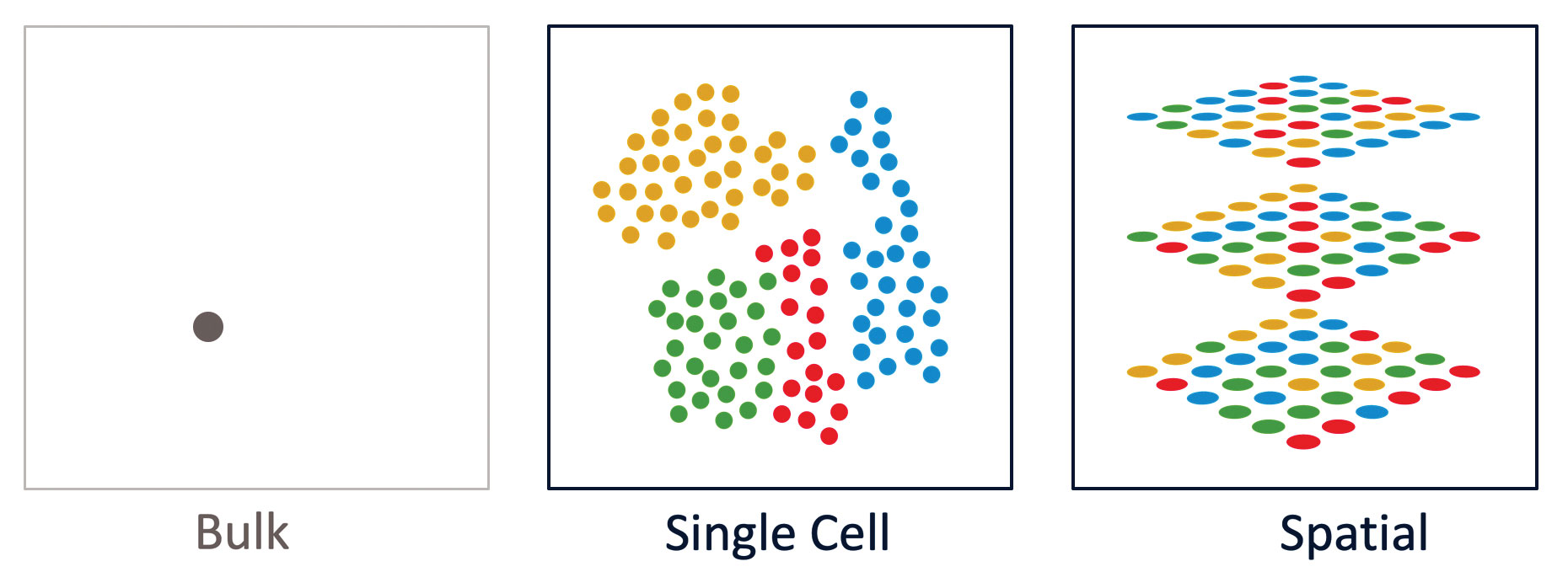
At 10x Genomics, our mission is to accelerate the mastery of biology to improve human health, and we believe that technological innovations are critical to driving this acceleration. Breakthrough technologies such as single-cell analysis, and newly emergent technologies such as spatial analysis, allow scientists to observe and measure complex biological processes, signaling pathways, and phenotypes at unprecedented scale and resolution. With this new information, we are starting to understand the differences between cell types—differences reflecting the influence of cell lineages, disease states, and individual traits—while gaining insights that may have clinical impact.
The single-cell revolution is just starting
In just a few years, single-cell analysis has grown from a niche technique used by experts into a widespread technique used across many areas of biological research, including cancer, immunology, immuno-oncology, neuroscience, and cell and developmental biology. New discoveries are being made—even in areas that had been well studied—because of the novel perspective that single-cell analysis provides.
Last year, Science and the American Association for the Advancement of Science named Single-Cell Analysis as the 2018 Breakthrough of the Year for its ability to track development of organisms and organs in stunning detail, cell by cell and through time, and for its overall potential for spurring advances in basic research and medicine that were not previously possible.1 These and other newer methods for single-cell genomics, transcriptomics, and proteomics will enable significant improvements in how we approach the clinical research, diagnosis, prognosis, and treatment of a broad range of human diseases.
Single-cell analysis has seen a tremendous growth in scale. In just a few short years, we have seen the number of single cells analyzed jump from a few thousand cells to hundreds of millions of single cells. Now, technologies having requisite scale for massively parallel and comprehensive characterization of millions of cells are enabling mapping projects to characterize all cell types of an organism, and ultimately the many trillions of cells found in the human body.
Cell-mapping projects will generate new foundational knowledge
The most ambitious cell-mapping project is the Human Cell Atlas (HCA), an initiative that has global and generational implications. The HCA’s mission is to create a comprehensive reference map of all human cells as a basis for understanding human health and diagnosing, monitoring, and treating disease.2 The information generated during this multiyear effort will help scientists understand how genetic variants impact disease risk, define drug toxicities, discover better therapies, and advance regenerative medicine.
The HCA has identified six flagship areas of impact: immune system, brain and nervous system, epithelial tissue, cancer, development and pediatrics. The HCA consortium has built a global membership of scientists and forged partnerships with projects such as the Immunological Genome Project (ImmGen), the Allen Brain Atlas, and the NIH BRAIN Initiative. The importance of the HCA is underscored by the funding received from notable organizations such as the Chan Zuckerberg Intiative, which has also invested in nonhuman atlas initiatives such as the Tabula Muris Consortium.3
An important trend is the call from the scientific community for open access to data and data sharing resources. The advancement of this trend is a founding principle of the HCA, which demands an open, modular, and extensible approach to data coordination to support the scale and complexity of its work. These resources are shared with partners, members, and scientists to both deposit and access data through this resource and to ensure the creation of a globally accessible resource, so that the discoveries made through the program will establish a legacy for understanding disease mechanisms.
New insights for cancer and other diseases
Single-cell approaches to investigate biology have already begun to transform our understanding of disease, uncovering hidden phenomena in areas thought to be well understood. In the area of inherited disease, HCA researchers recognized the pulmonary ionocyte, a previously unknown cell type that could be the key to better treatment for cystic fibrosis,4 and unique foveal cell types in the nonhuman primate retina expressing gene variants associated with causes of human blindness.5
In neuroscience, a recent study analyzed 80,660 single-nucleus transcriptomes from the prefrontal cortex of 48 individuals with varying degrees of Alzheimer’s disease.6 The team was able to identify six major brain cell types with transcriptionally distinct subpopulations that were characterized by regulators of myelination, inflammation, and neuron survival, with results suggesting that myelination has a key role in Alzheimer’s disease pathophysiology. Importantly, this new single-cell transcriptomic resource provides a blueprint for interrogating the molecular and cellular basis of Alzheimer’s disease.
Cancer atlas studies have shown that pediatric brain tumors appear much earlier than thought in utero,7 identified potential genes for early detection of gastrointestinal cancers,8 and discovered unique and unexpected populations of inflammatory macrophages in the human liver cell atlas.9
Longitudinal single-cell studies have provided a more dynamic understanding of how cancer cells change over time, including tumor heterogeneity, cell types, transitional cell states, and cell-to-cell interactions. Last year, researchers identified discrete and dynamic cellular changes that had occurred in an archived patient specimen. The researchers were able to reconstruct the evolution of that patient’s tumor, revealing pathways leading to cancer recurrence and eventual evasion from targeted therapy (which occurred after the patient’s initial positive response),10 suggesting promising future approaches for improving patient management and care.
Mapping the immune system
Modulation of the immune system, one of the most complex systems in biology, is critical in the development of immunotherapies, such as adoptive T-cell transfer and therapeutic vaccines, as well as in the development of other kinds of treatments against infections, autoimmune disorders, and cancer. New multiomic single-cell technologies for immune-cell phenotyping and antigen mapping are enabling researchers to explore immunology with unprecedented breadth, depth, and precision.
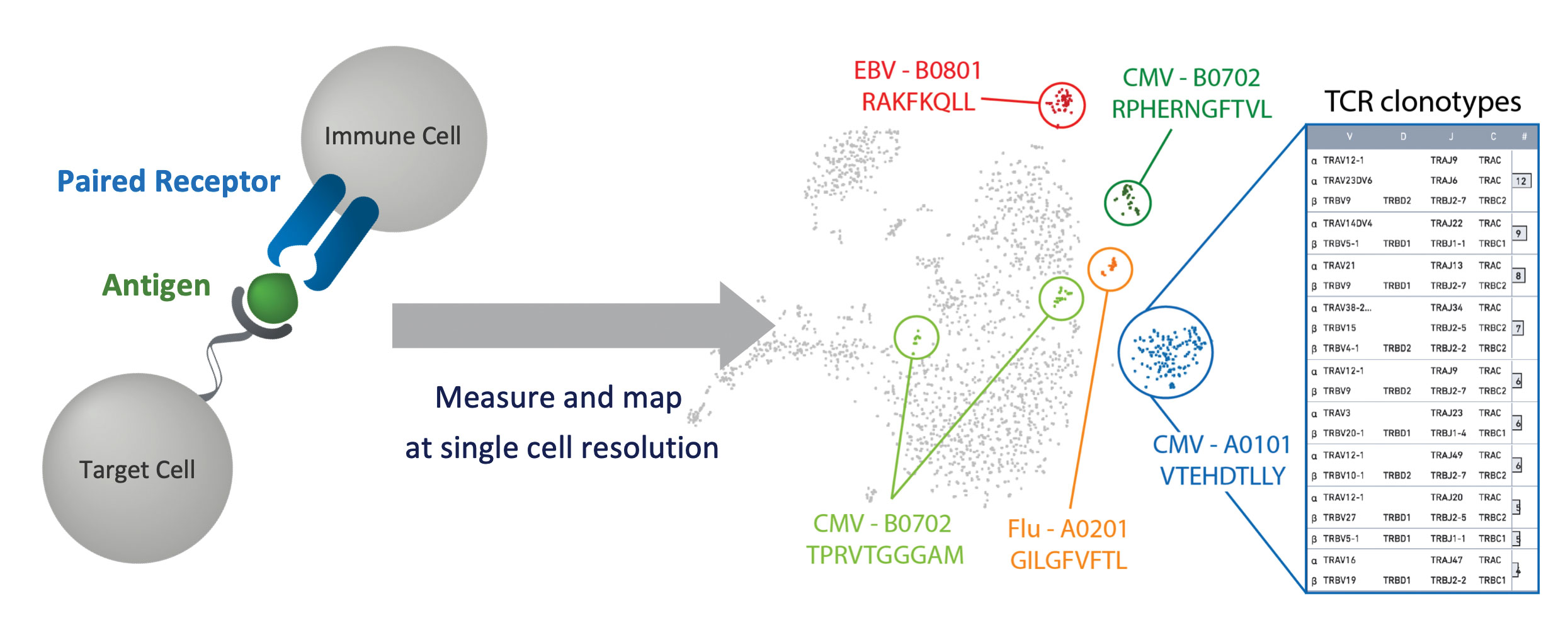
We can now define the full genetic repertoire of immune cells and reveal novel T-cell receptor (TCR)-antigen relationships, identify clonal expansions and peripheral blood markers, and measure T-cell exhaustion. Multiomic phenotyping is possible in tens of thousands of single T cells, including cell surface protein and antigen specificity characterization, and can be combined with immune repertoire and gene expression analysis with single-cell resolution.
Spatial resolution: Adding the “where” to “what”
New technologies are emerging for spatially resolved transcriptomics, which was selected as one of the eight “Methods to Watch” by Nature in 2017.13 Methods for spatial transcriptomics integrate information from applications such as gene expression and RNA sequencing with the contextual information provided from tissue analysis.
In late 2018, 10x Genomics acquired one of the innovators in this space, Spatial Transcriptomics, a company that helped pioneer this new technology.14 Our future efforts will further develop the platform to be compatible with our single-cell portfolio; improve performance; add the ability to interrogate difficult sample types, such as formalin-fixed paraffin-embedded (FFPE) tissues; and expand the technology to include new genomics applications.
These and other methods for spatial transcriptomics represent a groundbreaking way to measure all the gene activity in a tissue sample and map where that gene activity is occurring. Information that was previously out of reach is now helping scientists to make discoveries more quickly; new discoveries are already being made in neurodegenerative diseases, such as amyotrophic lateral sclerosis (ALS), as well as in breast and pancreatic cancer.
The next frontier will be to integrate these powerful new technologies for spatial and single-cell analysis to create new multidimensional modalities.
Ben Hindson, PhD, is a co-founder and the chief scientific officer of 10x Genomics.
References
1. Pennisi E. Development cell by cell. Science 2018; 362(6421): 1344–1345.
2. Human Cell Atlas.
3. Tabula Muris Consortium. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 2018; 562(7727): 367–372.
4. Montoro DT et al. A revised airway epithelial hierarchy includes CFTR-expressing ionocytes. Nature 2018; 560(7718): 319–324.
5. Peng YR et al. Molecular Classification and Comparative Taxonomics of Foveal and Peripheral Cells in Primate Retina. Cell 2019; 176(5): 1222–1237.
6. Mathys H et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 2019 (May 1): doi: 10.1038/s41586-019-1195-2. [Epub ahead of print]
7. Vladoiu MC et al. Childhood cerebellar tumours mirror conserved fetal transcriptional programs. Nature 2019 (May 1): doi: 10.1038/s41586-019-1158-7. [Epub ahead of print]
8. Zhang P et al. Dissecting the Single-Cell Transcriptome Network Underlying Gastric Premalignant Lesions and Early Gastric Cancer. Cell Reports 2019; 27(6): 1934–1947.
9. MacParland SA et al. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nature Communications 2018; 9(1): 4383.
10. Paulson KG et al. Acquired cancer resistance to combination immunotherapy from transcriptional loss of class I HLA. Nature Communications 2018; 9(1): 3868.
11. Azizi E et al. Single-Cell Map of Diverse Immune Phenotypes in the Breast Tumor Microenvironment. Cell 2018; 174(5): 1293–1308.
12. Tikhonova AN et al. The bone marrow microenvironment at single-cell resolution. Nature 2019; 569(7755): 222–228.
13. Nawy T. Spatial Transcriptomics [Method to Watch]. Nature Methods 2018; 15: 30.
14. Ståhl PL et al. Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 2016; 353(6294): 78–82.







Comments are closed.