Taralyn Tan Ph.D. Curriculum Fellow Harvard Medical
Check out these websites from GEN’s Best of the Web.
The Internet is a big place; when you’re looking for biotech-related websites, where should you start? At GEN’s Best of the Web, of course! Every other issue, we bring you a list of biotech- and biopharma-related websites we think you, GEN reader, would find useful and/or interesting. Here is our most recent list of the Best of the Web. Enjoy!
Key:
Four stars: Excellent
Three stars: Very Good
Two stars: Good
+ Strong points
– Weak points
Ant Genomes Portal ★★★
+ Browsable and downloadable genomes
– Not much information on wiki page
Ants are quickly becoming stars of genetic research, with their interesting and varied social behaviors, ease of maintenance in the lab, and sequenced genomes. The Ant Genomes Portal—part of the Hymenoptera Genome Database (HGD) project—is a great website that provides researchers with access to the seven sequenced ant genomes. The colorful cast of characters whose genomes have been sequenced includes a fungus eating ant, a leaf cutter ant, a Florida carpenter ant, Jerdon’s jumping ant, an Argentine ant, a red harvester ant, and a fire ant. The website includes genome browsers for each of these species, as well as an in-page BLAST search. Each genome assembly is available for download. In addition, the website contains a link to an Ant Genomes Wiki page, although the wiki does not yet contain much information.
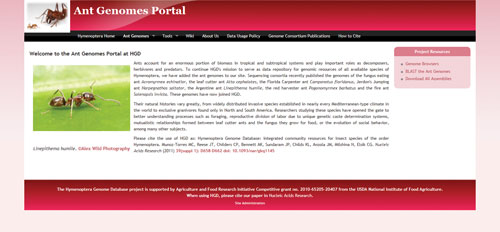
hymenopteragenome.org/ant_genomes
NetPrimer ★★★
+ Easy to use, adjustable parameters
– Need to register for free account to access
NetPrimer is not one of those primer tools that will do all of the work for you and design the primers; however, if you have primer sequences, NetPrimer has the analytical power to tell you how good your primers are. Simply enter a sequence into the NetPrimer Java applet and hit “analyze” to identify any palindromic, repeat, or run sequences within your primer, and to reveal whether your primer will form hairpins or dimers. NetPrimer will also return a number of calculated values for the inputted sequence, including melting temperature, GC composition, and various energy values. These values are calculated from a set of default parameters (ion concentrations, temperature, etc.) that can be modified by the user. The NetPrimer tool allows users to enter both a sense and antisense primer sequence to also assess interprimer interactions. Users can choose to enter names and descriptions for their primer sequences, which is useful given that users can print the resulting analyses.
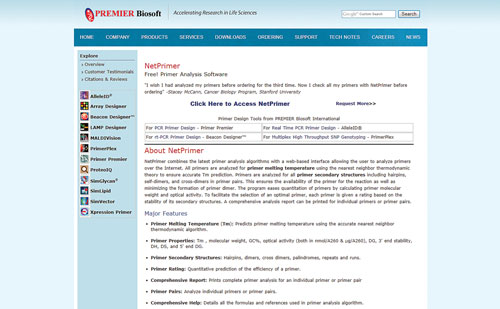
www.premierbiosoft.com/netprimer/index.html
bioRxiv ★★★★
+ Sleek site design, easy to browse contents
– Relatively new, not many manuscripts yet
In its purest form, the goal of scientific research is to discover new knowledge and share that knowledge with other scientists and the general public. However, sharing one’s research discoveries in the biological sciences via peer-reviewed journals is made difficult by the publishing process—it often takes months or even years for submitted manuscripts to be accepted for publication, as multiple rounds of revisions (and time-consuming additional experiments) may be requested. Now, however, scientists have a way to fight back against the lengthy publishing process, thanks to bioRxiv (pronounced “bio-archive”). Operated by the Cold Spring Harbor Laboratory, bioRxiv is an online repository for unpublished manuscripts in the life sciences. The preprints are published online on the bioRxiv site, where other scientists (and the general public) can read, comment on, and discuss the work. There is no paywall and no lengthy review process—just science, shared with the community.
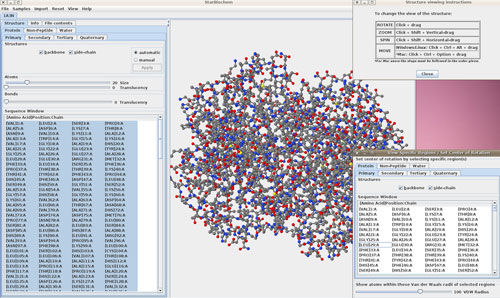
StarBiochem ★★★★
+ Intuitive to use, shows different structural levels
– Sometimes slow to render
There exist a number of high-quality 3D molecule viewers; however, the StarBiochem program is different in that it is specifically designed for students of biochemistry. In biochemistry courses, protein structure is introduced sequentially: primary, secondary, tertiary, and quaternary structure. Similarly, the StarBiochem Java applet allows users to explore 3D protein structures at these four levels of molecular interactions. For each structural level, users can adjust the size and transparency of the atoms and bonds (or larger structural motifs), which allows users to simultaneously view primary-through-quaternary structural features. Additionally, the program allows users to display or hide specific amino acid residues or other features of the 3D model. Users can open any of the included sample files or import any PDB file. The program is accompanied by a concise but thorough user manual and is overall quite intuitive to use.
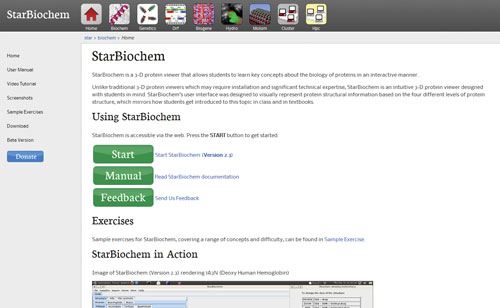
star.mit.edu/biochem/index.html
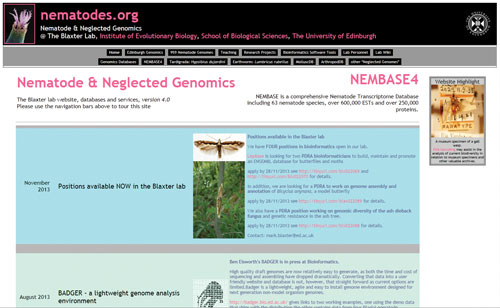
Nematode and Selected Genomics ★★★★
+ Many sequence databases and bioinformatics tools
The website for the Blaxter lab at the University of Edinburgh—which is subtitled, “Nematode and Neglected Genomics”—is the perfect place to visit if you are searching for genetic information for “misfit” organisms of the current genomics era. You won’t find DNA sequence information for Drosophila melanogaster or Mus musculus here; instead, you can browse sequence databases for 63 species of nematodes, 1 tardigrade species (or “water bear”), 9 species of molluscs, and 2 axolotl species, among others. In addition to the multiple sequence databases, the website includes a number of downloadable bioinformatics tools. These include transcriptome analysis tools such as peptide predictors and sequence annotators; tools for phylogenomics analyses such as a taxonomy database manager; DNA barcode analysis tools; and general genomics tools such as quality assurance tools for next-gen sequencing datasets.
University of Colorado Bioinformatics Homepage ★★
+ Tutorials for many basic online bioinformatics tools
– Many broken links
I don’t know if the stark University of Colorado Bioinformatics Homepage website will necessarily succeed in its goal of showing people “why they should be interested” in bioinformatics, but the site does include some useful information for people who already are interested in the subject (but perhaps are not yet experts in the field). A strength of the site is its collection of tutorials for basic bioinformatics tools; for instance, site visitors are taught how to search on PubMed or OMIM, how to perform sequence alignments using a BLAST tool hosted by Washington University, and how to perform multiple sequence alignments using ClustalW. In addition, the website contains a number of useful links to bioinformatics-related resources. The site would be improved if some of the broken links were removed (there are many…). In any case, as long as a couple of “page not found” errors don’t scare you away, there is good information to be found on this site.

www.colorado.edu/chemistry/bioinfo
Want more Best of the Web? Click here! Also, to suggest a website for Best of the Web, please send the URL to Taralyn Tan ([email protected]).