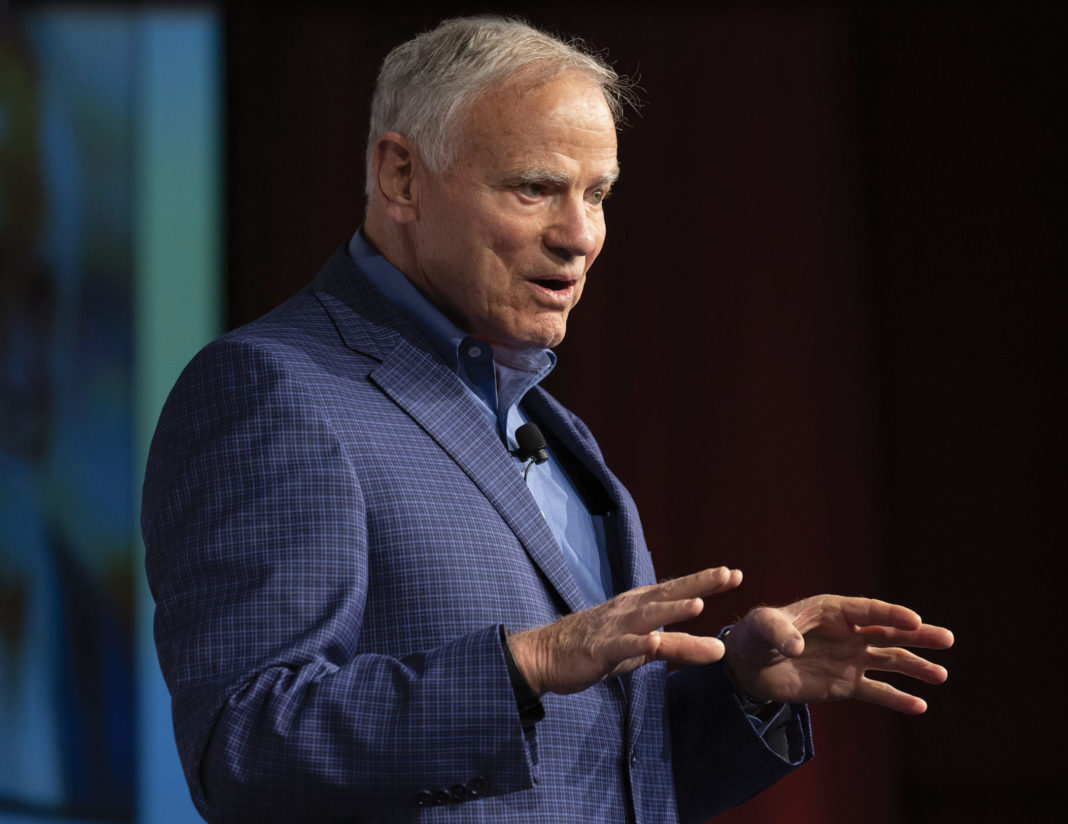
Leroy Hood is the perfect scientist to write the lead article in the first issue celebrating the 40th anniversary of GEN. Lee’s career essentially parallels that of the biotech industry, and as you will see from his article, he is hugely responsible for many of the essential tools and technologies that have driven much life sciences research and clinical product development over the past 40 years. We sincerely thank him for his special contribution to our January issue. —John Sterling, Editor in Chief

Indeed, these paradigm changes did begin to decipher human complexity in quite different and powerful ways, but they also framed my vision of 21st century medicine—my current focus. I will briefly describe these paradigm changes and then consider some of the impacts they had on my thinking about science and how scientific careers may be formed.
Seven paradigm-shifting developments
1. Confluence of engineering and biology
Engineering has been brought to biology over the past 40 years through the development of six instruments that allow the sequencing and synthesis of DNA and proteins.1–6 These tools gave us the ability to generate high-throughput data from human samples and also allowed us to begin accumulating the data needed to realize big data analytics.
2. Success of the Human Genome Project
I was privileged to participate in the conceptualization, initiation, and execution of the Human Genome Project.7–10 I was particularly excited by the way genome technology advanced our ability to analyze human genetic variation in the context of wellness and disease phenotypes.
3. Adoption of cross-disciplinary approaches
In 1992, I founded the first cross-disciplinary biology department, the Department of Molecular Biotechnology at the University of Washington. It brought together astronomers, biologists, chemists, computer scientists, engineers, mathematicians, and physicians. During the eight years of its existence, it made seminal technological innovations to genomics, the newly emerging field of proteomics, cell biology, and computational analyses.11–15 This paradigm-shifting development facilitated the transition from genotype to phenotype, and from technology to biology.
4. Establishment of systems biology
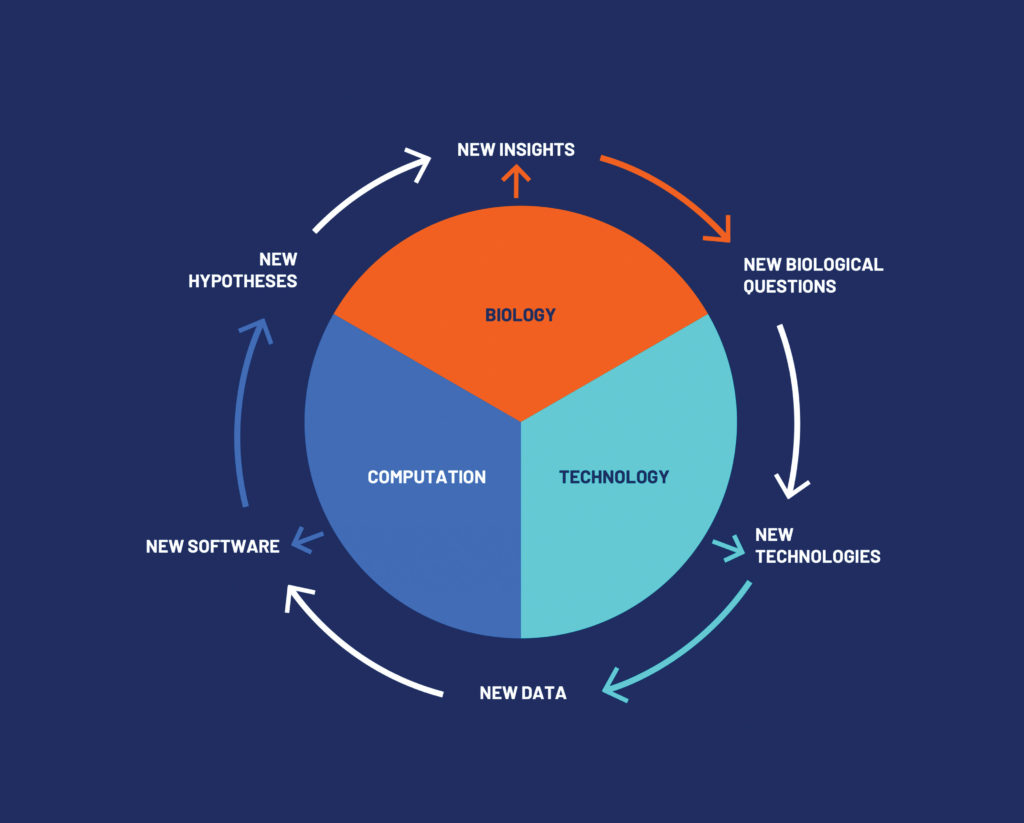
In 2000, I co-founded the first institution dedicated to systems biology, the Institute for Systems Biology (ISB), which pioneers global and holistic approaches to biology and disease. The ISB drives the application of genomics and computational technologies, including technologies that I helped develop in past decades, to the scientific problem of biocomplexity: moving biology from studying genes and proteins one at a time to analyzing entire systems.16–19 Essentially, the ISB works to understand what makes the whole more than the sum of its parts.
At the ISB, we learned to unite traditional holistic organismal biology with modern molecular biology first on model organisms, but we soon applied lessons learned to the study of human health and disease. The new systems paradigm led to the formulation in the early 2000s of my view of 21st century medicine. (I will refer to the 21st century medicine concept throughout this article).
5. Development of the P4 concept
The systems approach has been adapted to medicine and healthcare, leading to the view that treatments can be predictive, preventive, personalized, and participatory (P4). Furthermore, P4 healthcare now encompasses two domains: wellness (which had received little attention in traditional mainstream medicine) and disease.20–23
6. Systematic quantitation of wellness data
In 2014, Nathan Price, PhD, an ISB colleague and good friend, and I persuaded 108 individuals to undergo complete genome sequence analysis—and longitudinal phenome analyses of proteins, metabolites, clinical chemistries, and the gut microbiome every three months for a year—in addition to analyses of data collected with self-tracking technologies such as Fitbit, that is, analyses of “quantified self” data clouds. When analyzed, these data clouds produce individualized recommendations for optimizing wellness and avoiding disease.
This approach relies on multidimensional measurements of personal health variables and on existing generic medical knowledge. We call this approach scientific (quantitative) wellness.24–27
We started the for-profit company called Arivale in 2015 to bring scientific wellness to 5,000 clients over four years, thus generating informative longitudinal data clouds that confirmed the power of scientific wellness and led to the concept of healthy aging. We also confirmed the use of genetic risk for diseases to determine therapy strategies; developed the ability to detect, at the earliest stages, almost 200 wellness-to-early disease transitions; and developed ways to identify systems-driven therapies to reverse chronic diseases before they manifest as disease phenotypes.28–30
7. Launch of combined genome/longitudinal phenome analyses
Paradigm changes 5 and 6 broadened my outlook. I had been focused on the complexities of human biology and how they could inform the targeting of disease phenotypes. But then I began to appreciate a larger concept, namely, 21st century medicine. Enabled by new technology and biological insights, I advocated that we should follow the multidimensional health trajectories of each individual with genome and longitudinal phenome monitoring.31
In 2016, Rod Hochman, MD, the visionary president and CEO of Providence St. Joseph Health (PSJH), asked me to become the CSO of PSJH, and for the ISB to introduce PSJH to genomics and scientific wellness. I agreed because this was a way to bring 21st century medicine to a major healthcare system.
In this context, I immediately began planning a bold proposal to carry out genome/longitudinal phenome analyses on one million PSJH patients over five years, studying both well and diseased patients—the Million Person Project (MPP). This proposal could bring 21st century medicine with scientific wellness, healthy aging, and the prediction and prevention of diseases to a major healthcare system.31
Bridging academic disciplines
My career has spanned biology (molecular immunology, molecular neurobiology, and systems biology), medicine (P4 healthcare, scientific wellness, healthy aging, Alzheimer’s disease, 21st century medicine), technology development, and education (K-12 students, undergraduates, and graduates, as well as scientists and physicians). In the following, I draw some personal observations from these areas and the seven paradigm changes described above.
Thomas Kuhn wrote a thoughtful book on paradigm changes, The Structure of Scientific Revolutions.32 It suggests that paradigm changes are fundamentally new ways of thinking about or practicing science, and that they are extremely difficult to achieve because many scientists have conservative natures and are skeptical about new ideas. My experience echoes his insights. Moreover, my familiarity with the seven paradigm changes described earlier leads me to offer the following observations:
- Each of the seven paradigm changes was met with enormous skepticism, with doubters saying, “It is not true” or “It is trivial” or “It is a pipe dream.”
- To overcome such skepticism—to carry an initiative forward against the opinions of a vast majority of one’s colleagues—one needs a quality I call “determined optimism.”
- New ideas need new organizational structures to move forward. The bureaucracies of an organization are honed by past experience, and they often just barely manage to deal effectively with the challenges of the present. They generally fail completely in dealing with challenges of a more distant future. This is why industry, on occasion, creates an independent entity called a “skunk works” to explore new ideas unconstrained by the parent company. (For example, Xerox founded the Palo Alto Research Company (PARC), which developed innovative computer technology. Although the technology wasn’t exploited to the full by the parent company, it inspired Steve Jobs and the engineers at Apple.)
- The only way to convince the skeptics of a paradigm change is to be successful—to create transformational ideas or tools that open new avenues of exploration.
In developing new tools or biological opportunities, academia and industry have distinct roles. Academia is good at conceptualizing new concepts and tools and performing proof-of-principle studies. Industry, with its larger resources and broader focus, excels at making new tools simple and robust, and at scaling performance to enable, for example, high-throughput data generation. This is exactly what happened with the development of automated DNA sequencing over the last 35 years.
Sequencing has now gone through three distinct stages: 1) the four-color Sanger chemistry of our first-stage instrument with up to several hundred sequence reads; 2) four-color reads with enormous parallelization to millions of short-read sequences; and 3) single-molecule sequence analyses generally with electronic detection and potentially long reads, allowing de novo assembly of each individual’s genome sequence independent of a relatively inaccurate standard prototype sequence. (I predict that this latter technology will, in time, bring the cost of individual genome sequences below $10.)
Bridging academic and commercial realms
I remember that late in 1979, I visited the president of Caltech, the physicist Marvin “Murph” Goldberger, PhD, and suggested that we should bring the tools we were developing to industry after we had proved their effectiveness. These tools included four automated instruments, one each for protein sequencing, DNA sequencing, peptide synthesis, and DNA synthesis.

Eventually, Bill Bowes, a San Francisco venture capitalist, offered to help fund the company that became Applied Biosystems and successfully commercialized the four instruments developed at Caltech. I subsequently helped co-found 15 additional companies. Each was generally successful in bringing our concepts or technologies to fruition.
One can create new companies to realize scientific objectives, or one can collaborate with them on science. Both approaches pose significant challenges. For example, in the early 1980s, I went to Edward Jefferson, PhD, then CEO of DuPont, and proposed that we collaborate on the protein-level sequencing of several interferons, which were then available only in vanishingly small quantities. Once this was accomplished, we could use integrative technology strategies to clone the corresponding genes (see the following).
Jefferson was enthusiastic, and he pointed me toward his director of life sciences, who proved to be an absolute roadblock. (The director had his own agenda and budget commitments and did not appreciate the intrusions of an academic.) I learned that although it is essential to get C-suite approval for a novel project, you must also persuade the operational scientists who direct the work that it is to their benefit. This was a lesson I never forgot.
Just as one can take a systems approach to biology and disease, it is also possible to do so with technology. I remember writing a paper on the integration of DNA and protein sequencers and synthesizers in early 1982 for Nature. The idea was that microsequencing vanishingly small quantities of protein would allow one to use the genetic code dictionary to synthesize degenerate oligonucleotide probes that could then be used to clone the corresponding gene.
We could then sequence the gene (and use it as a probe for expression analyses) and reverse-translate this sequence to peptide sequences. Then, these sequences could be used to synthesize peptide fragments, which could be used to generate antibodies to detect the protein. Thus, the microchemical facility gave one to ability to take a small amount of protein and generate the reagents necessary for determining its function—in a manner that had never before been possible.
The integration of these four technologies allowed us to look at biology through a fundamental new lens. This paper took more than two years to publish—because Nature demanded that we demonstrate the validity of each of the various components of these integrated approaches.33 We went on to microsequence more than 25 different proteins and often cloned their corresponding genes, which opened fascinating new areas for biological exploration.34–37
Let me add that several individuals played critical roles in developing these technologies: Michael W. Hunkapiller, PhD (automated protein and DNA sequencing); Lloyd M. Smith, PhD (automated DNA sequencing); Steven B.H. Kent, PhD (automated peptide synthesis); and Marvin H. Caruthers, PhD (pioneered the DNA synthesis chemistry and worked with Applied Biosystems on automated DNA synthesis). Applied Biosystems played a major role in catalyzing the maturation of these technologies.
My adventures in creating cross-disciplinary scientific environments led me to realize that, for future scientists, cross-disciplinary talents are essential. In addition to gaining enormous domain expertise in a particular area of biology, a young biologist should also develop the computational and statistical skills that will be necessary for dealing not only with big data but also with “big knowledge,” which will be central to all fields in biology and medicine.
A student should have some working understanding of not only data analytics, machine learning, and artificial intelligence, but also of complex systems theories and knowledge graphs. All are invaluable tools for managing and analyzing complex biological information and comprehending how the parts integrate to a whole.
Healthy aging
On a personal level, I was really excited when we showed that we could use the 5,000 Arivale longitudinal data clouds to determine for each individual their biological age—that is, the age your body says you are, rather than the age specified by your birthday.28 The younger the biological age is relative to your chronological age, the better you are aging. Three hundred Arivale individuals with diabetes, for example, had an average biological age six years older than their chronological ages.
I have a biological age 15 years younger than my chronological age, and I am a fanatic for exercise. Not only can one use biological age as a metric for wellness, one can also use out-of-range blood analyte concentrations to calculate a person’s biological age and optimize at least to some degree each individual’s healthy aging.
Nathan Price and I started a company called Aevum to commercialize this opportunity. (Aevum has recently been integrated into a health intelligence company called Onegevity.) My own belief is that scientific wellness plus healthy aging will go a long way to ensuring one can live mentally alert and physically agile well into your 90s or even 100s. I am living the experiment now.
Envisioning 21st century medicine
I am excited by a fundamental idea called 21st century medicine and what it will do for healthcare. The idea is one that we are already applying by exploting personal data clouds. Every one of us is different—not only with respect to the genome, but also in thousands of dimensions of the phenome.
Therefore, 21st century medicine is an N-of-1 medicine: We will be able to optimize the health trajectory for each individual through assessments of the genome and longitudinal phenome and interrogating the vast knowledge graphs that soon will encompass the entirety of our biomedical knowledge. The output for the individual is customized and concrete, and it offers actionable possibilities to influence the health trajectory in a desired way.38,39
Each individual’s health trajectory has three stages: 1) wellness, which often characterizes the early and middle years of life; 2) an occasional transition from wellness to disease; and 3) disease, which occurs if the second stage is not checked or naturally reversed. 21st century medicine will optimize wellness and extend it as long as possible for each life through scientific wellness and healthy aging.
We have shown that it is possible to detect wellness-to-disease transitions many years before any clinical disease phenotype is manifest for many chronic diseases (diabetes, cancer, Alzheimer’s, etc.).29 So, in the future, we will reverse diseases at this earliest and simplest stage when they are still reversible. Hopefully, we will do this for all chronic diseases that plague mankind and impart a heavy financial burden to society.
Thus, the vision of 21st century medicine is P4—predictive, preventive, personalized, and participatory. “Predictive” and “preventive” refer to the detection and reversal of early chronic diseases, for example, through the use of genetic risk assessments to define treatment approaches that distinguish between patients who are at high or low risk of having a given disease. “Personalized” refers to scientific wellness and healthy aging. “Participatory” refers to the ways in which patients may actively participate in 21st century medicine.40
Finally, we will use systems approaches to treat manifest disease more effectively. Despite the adage that prevention is better than treatment, contemporary medicine still deals with disease generally after it is clinically diagnosed. Scientific research has mostly been devoted to correcting the clinical disease which is generally not reversible by simple single-drug treatments once critical transitions in the underlying disease mechanisms have happened.
Profound applications of 21st century medicine
Optimizing wellness through scientific wellness and healthy aging and the detection and early reversal of chronic disease before the point of no return lies at the heart of optimizing health in 21st century medicine.31
One of the personal tragedies of my life was the diagnosis of my wife, Valerie, with Alzheimer’s disease (AD) almost 15 years ago. I know the heartbreak of seeing a loved one slowly fade away from the world of awareness and the agony of going through these transitions together as a caretaker. At the time of her diagnosis, all drugs were ineffective, and this prompted me to take a serious look at AD.
Over the last 12 years, 400 clinical trials have been ineffective. A totally new start is needed. We need to recognize that it is unlikely that one drug will cure AD.
I decided that AD would be the first chronic disease that we would approach using 21st century medicine for early detection and prevention. It is clear AD starts 4–10 years before it is clinically diagnosed (from metabolic PET scanning). To determine how well we can detect the earliest AD transitions, we are designing a clinical trial with genome/longitudinal phenome analyses to assess individuals at high risk for AD.
Once AD is detected, we believe that a multimodal approach (drugs, supplements, exercise, proper diet, removal of toxins, lifestyle modification, etc.), where each mode is rooted in a scientific rationale, is the key to treating AD.41 We have started three clinical trials using various combinations of multimodal therapies on early AD patients. (I recently presented details about the multimodal approach at an event organized by the Innovation Research Interchange.42)
My hope is that within five years, we will be able to reverse more than 80% of early AD, and that later, we will prevent it completely. Wouldn’t a world without AD be a wonderful tribute to the Valeries of the world and their caretakers?
Upon the outbreak of COVID-19 in Seattle in February of 2020, we started an observational study of 300 patients, taking blood samples at diagnosis, 10 days later, and 40 days later in convalescence. In addition to performing genome/deep phenome analyses, we carried out deep immune phenotyping (single-cell analyses of 5,000 white blood cells at each blood draw for each patient). We analyzed single-cell transcriptomes, relevant cell-surface and secreted molecules, the presentation of HLA alleles, and T-cell and B-cell receptor sequences.
This deep immune phenotyping was revelatory.43 We determined trajectories corresponding to patients who experienced disease with different degrees of severity. Specifically, we identified trajectories indicating the drugs that allowed us to design optimal clinical trials (for example, the optimal time point to give the drug to achieve maximal effect); we identified new states of immune cells and followed their responses to the virus (activation and evolution in maturity); and we made fundamental observations about the nature of viral epitopes that the immune systems employed.
These had profound implications for vaccines. What these studies told us clearly was that from just 100 or so patients, we could derive fundamental insights. We are confident that in the future, we will use deep phenotyping to probe the immune, hormone, metabolic, neurological, and other systems. Moreover, this work will allow us to gain new insights into any disease. We hope to go far beyond what large clinical trials have ever done before. Even trials that encompass large cohorts have confined themselves to addressing questions about a handful of variables. Deep phenotyping will spawn a host of new types of clinical trials that will be insightful and revelatory.
Sources of inspiration
Three people had an enormous impact on my career. Cliff Olsen, my high school chemistry teacher from Shelby, MT, showed me how exciting science could be, and he convinced me that I had the potential to do something significant in science. Perhaps most important, he persuaded me to go to Caltech as an undergraduate, where I received superb training to be a scientist.
Ray D. Owen, PhD, an immunologist at Caltech, was present during my entire 30-year career at Caltech. I was an undergraduate, a graduate student, a faculty member for 22 years, and chairman of the biology department for my last 10 years. He introduced me to the fun and excitement of science, and at several stages of my career, he gave me invaluable advice about how to be a professor or a chairman. He was a wonderful friend up until he died at 94.
My doctoral advisor at Caltech, William J. Dreyer, PhD, introduced me to the power of technology to transform biology and argued compellingly that one should always practice biology at its frontiers—dictums I have taken seriously.
A fascinating question is, what events had the most impact on my career? I would say without question it was the four major moves I made during my academic career. The first three moves took me from one institution to another—from the National Institutes of Health to Caltech; from Caltech to the University of Washington; and from the University of Washington to start the Institute for Systems Biology in Seattle. When I made my next move, I became chief science officer for PSJH while retaining my professorship at the ISB. With each move, I reset my scientific objectives, met fascinating new people, and organized new scientific entities. I effectively changed careers multiple times.

Indeed, if you challenge yourself in this way every 10–15 years, you can maintain a career with a continuously ascending positive success slope. I am testing the possibility that I will never want to retire if I employ this strategy. One must be certain to always explore new avenues.
The final point I would like to make is that systems thinking can be applied to virtually every problem in contemporary science—life sciences, medicine, agriculture, energy, healthcare, education, global warming, climate change, etc. It can also be applied to virtually every academic discipline, and it can be applied to our personal lives. We are surrounded in life by systems at every level that cry out for understanding and effective responses through systems approaches. All students should be taught systems thinking for health and life. We have designed (and continue to design) courses for K-12 students in this regard.
With the challenges of realizing 21st century medicine, I think I will have enough to do to keep me out of retirement for the rest of my functional life—through my 90s. Perhaps I will write another commentary in 20 years on our progress in P4 healthcare.
References
1. Blanchard AP, Kaiser RJ, Hood L. High-Density Oligonucleotide Arrays. Biosens. Bioelectron. 1996; 11: 687–690.
2. Geiss GK, Bumgarner RE, Birditt B, et al. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nat. Biotechnol. 2008; 26(3): 317–325.
3. Hewick RM, Hunkapiller MW, Hood LE, Dreyer WJ. A gas-liquid solid phase peptide and protein sequenator. J. Biol. Chem. 1981; 256(15): 7990–7997.
4. Horvath SJ, Firca JR, Hunkapiller T, et al. An automated DNA synthesizer employing deoxynucleoside 3′-phosphoramidites. Methods Enzymol. 1987; 154: 314–326.
5. Kent SB, Hood LE, Beilan H, et al. A novel approach to automated peptide synthesis based on new insights into solid phase chemistry. In: Isymiya N, ed. Proceedings of the Japanese Peptide Symposium. Osaka: Protein Research Foundation; 1984: 217–222.
6. Smith LM, Sanders JZ, Kaiser RJ, et al. Fluorescence detection in automated DNA sequence analysis. Nature 1986; 321(6071): 674–679.
7. Kevles DJ, Hood L, eds. The Code of Codes: Scientific and Social Issues in the Human Genome Project. Cambridge, MA: Harvard University Press; 1993.
8. Lander ES, Linton LM, Birren B, et al. Initial sequencing and analysis of the human genome. Nature 2001; 409(6822): 860–921.
9. Brüls T, Gyapay G, Petit JL, et al. A physical map of human chromosome 14. Nature 2001; 409(6822): 947–948.
10. Heilig R, Eckenberg R, Petit JL, et al. The DNA sequence and analysis of human chromosome 14. Nature 2003; 421(6923): 601–607.
11. Gordon D, Abajian C, Green P. Consed: a graphical tool for sequence finishing. Genome Res. 1998; 8(3): 195–202.
12. Gygi SP, Rist B, Gerber SA, et al. Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat. Biotechnol. 1999. 17(10): 994–999.
13. Hood L. A personal journey of discovery: developing technology and changing biology. Annu. Rev. Anal. Chem. (Palo Alto Calif) 2008; 1: 1–43.
14. Link AJ, Eng J, Schieltz DM, et al. Direct analysis of protein complexes using mass spectrometry. Nat. Biotechnol. 1999. 17(7): 676–682.
15. Ewing B, Hillier L, Wendl MC, Green P. Base-calling of automated sequencer traces using phred. I. Accuracy assessment. Genome Res. 1998. 8(3): 175–185.
16. Bonneau R, Facciotti MT, Reiss DJ, et al. A predictive model for transcriptional control of physiology in a free living cell. Cell 2007; 131(7): 1354–1365.
17. Hood L, Rowen L, Galas DJ, Aitchison JD. Systems biology at the Institute for Systems Biology. Brief. Funct. Genomics Proteomics 2008; 7(4): 239–248.
18. Ideker T, Galitski T, Hood, L. A new approach to decoding life: systems biology. Annu. Rev. Genomics Hum. Genet. 2001. 2: 343–372.
19. Ideker T, Thorsson V, Ranish JA,et al. Integrated genomic and proteomic analyses of a systematically perturbed metabolic network. Science 2001. 292(5518): 929–934.
20. Hood L, Flores M. A personal view on systems medicine and the emergence of proactive P4 medicine: predictive, preventive, personalized and participatory. N. Biotechnol. 2012; 29(6): 613–624.
21. Hood L, Friend SH. Predictive, personalized, preventive, participatory (P4) cancer medicine. Nat. Rev. Clin. Oncol. 2011; 8(3): 184–167.
22. Hood L, Heath JR, Phelps ME, Lin B. Systems biology and new technologies enable predictive and preventative medicine. Science 2004; 306(5696): 640–643.
23. Weston, AD, Hood L. Systems biology, proteomics, and the future of health care: toward predictive, preventative, and personalized medicine. J. Proteome. Res. 2004; 3(2): 179–196.
24. Hood, L, Lovejoy JC, Price ND. Integrating big data and actionable health coaching to optimize wellness. BMC Med. 2015; 13: 4.
25. Hood L, Price ND. Demystifying disease, democratizing health care. Sci. Transl. Med. 2014; 6(225): 225ed5.
26. Price ND, Magis AT, Earls JC, et al. A wellness study of 108 individuals using personal, dense, dynamic data clouds. Nat. Biotechnol. 2017; 35(8): 747–756.
27. Zubair N, Conomos MP, Hood L, et al et al. Genetic Predisposition Impacts Clinical Changes in a Lifestyle Coaching Program. Sci. Rep. 2019; 9(1): 6805.
28. Earls JC, Rappaport N, Heath L, et al. Multi-Omic Biological Age Estimation and Its Correlation With Wellness and Disease Phenotypes: A Longitudinal Study of 3,558 Individuals. J. Gerontol. A Biol. Sci. Med. Sci. 2019; 74(Suppl. 1): S52–S60.
29. Magis AT, Rappaport N, Conomos MP, et al. Untargeted longitudinal analysis of a wellness cohort identifies markers of metastatic cancer years prior to diagnosis. Sci. Rep. 2020; 10(1): 16275.
30. Wainberg M, Magis AT, Earls JC, et al. Multiomic blood correlates of genetic risk identify presymptomatic disease alterations. Proc. Natl. Acad. Sci. U.S.A. 2020; 117(35): 21813–21820.
31. Hood, L, Price ND, Evans SJ. What 21st Century Medicine Should Be—History, Vision, Implementation and Opportunities. In: Barilan YM, Brusa BM, Ciechanover A, eds. Can Precision Medicine Be Personal? Can Personalized Medicine Be Precise? Oxford University Press. In press.
32. Kuhn, TS. The Structure of Scientific Revolutions. University of Chicago Press; 1962.
33. Hunkapiller M, Kent S, Caruthers M, et al. A microchemical facility for the analysis and synthesis of genes and proteins. Nature 1984; 310(5973): 105–111.
34. Doolittle RF, Hunkapiller MW, Hood LE, et al. Simian sarcoma virus onc gene, v-sis, is derived from the gene (or genes) encoding a platelet-derived growth factor. Science 1983; 221(4607): 275–257.
35. Oesch B, Westaway D, Wälchli M, et al. A cellular gene encodes scrapie PrP 27-30 protein. Cell 1985; 40(4): 735–746.
36. Prusiner SB, Groth DF, Bolton DC, et al. Purification and structural studies of a major scrapie prion protein. Cell 1984; 38(1): 127–134.
37. Raftery MA, Hunkapiller MW, Strader CD, Hood LE. Acetylcholine receptor: complex of homologous subunits. Science 1980; 208(4451): 1454–1456.
38. Huang S, Hood L. Personalized, Precision, and N-of-One Medicine: A Clarification of Terminology and Concepts. Perspect. Biol. Med. 2019; 62(4): 617–639.
39. Schork NJ. Personalized medicine: Time for one-person trials. Nature 2015; 520(7549): 609–611.
40. Hood L, Auffray C. Participatory medicine: a driving force for revolutionizing healthcare. Genome Med. 2013; 5(12): 110.
41. Bredesen DE. The End of Alzheimer’s: The First Pogram to Prevent and Reverse Cognitive Decline. New York, NY: Avery; 2017.
42. Hood L. How Technology, Big Data, and Systems Approaches Are Transforming Medicine. Paper presented at: Annual Conference of the Innovation Research Interchange—Innovation Unleashed; April 29–May 2, 2019.
43. Su Y, Chen D, Yuan D, et al. Multi-Omics Resolves a Sharp Disease-State Shift between Mild and Moderate COVID-19. Cell 2020; October 28. DOI: 0.1016/j.cell.2020.10.037.
To help us preview the future, GEN asked opinion leaders, all from outstanding technology companies, to discuss a range of new initiatives. View the rest of the articles below.
Uncharted Territory: Top Challenges Facing Gene Therapy Development
Envisioning Future Trends in Regenerative Medicine
Engineering Biology—Accelerating Transition
Bioprocessing in a Post-COVID-19 World
Sustainability and the Synthetic Biology Revolution
Sowing the Seeds of Agricultural Biotechnology
Neuroscience Widens Its Investigations of Disease Mechanisms