May 15, 2009 (Vol. 29, No. 10)
Ruth A. VanBogelen
Richard C. Jones
M. Walid Qoronfleh
Multiple Reaction Monitoring Platform Addresses Obstacles in Multiplex Testing
In the post-genome era, the use of molecular biomarkers is becoming de rigueur for R&D programs. The biomarker testing market has a proven record of revenue generation ($612 million in 2007) and is estimated to have an annual growth rate of 23.5% based on currently available biomarker assays. Thus both need and return on investment are driving the development of new technology and faster, more cost-effective biomarker development workflows.
Proteomic technologies have, in the past, been used successfully for biomarker discovery projects, producing many candidate protein biomarkers. Further verification work, however, is typically limited by the small number of proteins for which there are commercially available assays (about 500 human proteins). Using traditional antibody-based approaches it could take years and cost millions of dollars to develop assays for all of the candidate protein markers.
NextGen Sciences devised a workflow for development of protein biomarker assays, overcoming the bottleneck with the antibody-based approach (Figure 1).
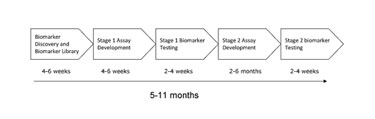
Figure 1. Workflow for biomarker assay development
Biomarker Discovery
Biomarker discovery platforms must be able to detect a large number of protein species in the sample type of interest and at the same time provide the protein information necessary for assay development. A platform that meets these criteria is commonly referred to as GeLC-MS (Figure 2); a combination of SDS-PAGE for protein separation and fractionation and liquid chromatography-tandem mass spectrometry (LC/MS/MS) for detection and quantitation of proteins via proteolytically derived peptides.
The process involves quantitative characterization of the proteome of a biological sample. Sample preparation methods vary depending on the nature of the sample. The proteins in each sample are then separated by SDS-PAGE. Each lane on the gel is then segmented (typically 24–40 segments) before enzymatic digestion. Peptides from each digest are analyzed by mass spectrometry (LC/MS/MS). The top graph of Figure 2C shows a typical chromatogram for a one-hour LC gradient, while the bottom graph shows an example of MS/MS (product ion) data for one peptide.
Approximately 50,000 spectra corresponding to 25,000 unique peptides are typically detected per lane. The data is then searched against appropriate protein databases; after false discovery rate analysis is performed a nonredundant list of proteins is compiled using commercially available tools. The end product includes a list of proteins identified with quantitative data for each protein in the sample. Statistical analysis is applied to the quantitative data providing fold changes across groups, coefficient of variance, p-values, principle component analysis, and hierarchal clustering.
The GeLC/MS output includes data about the peptides matched to each protein and their isoforms present in the sample. Peptide information includes amino acid sequence, product ion data, chromatographic performance, and ion intensity. The data from the GeLC-MS platform not only provides the quantitation needed to select putative biomarkers but also provides the empirical peptide information required for the assay development.

Figure 2 (A-F). Biomarker Discovery
Stage-One Assay Development
The objective of stage-one assay development is to develop an assay that will be used to confirm candidate biomarkers. Having the necessary data on the candidate proteins captured during biomarker discovery allows for quick assay development. This information may also be provided by a database of proteins previously detected by mass spectrometry. NextGen Sciences has such a database of protein biomarker information, called biomarkerlibrary™.
The assay platform used in this workflow is multiple reaction monitoring (MRM) for peptides (Figure 3). This is a mass spectrometry-based assay that targets specific surrogate peptides in a sample. The approach brings high specificity for each protein in the assay. MRM assays do not require antibodies and can be designed to monitor a specific protein, single or multiple isoforms of a protein, and even a single amino acid mutation within the protein(s) being measured.
Peptides for each protein are selected based on their known amino acid composition, response factor, hydrophobicity, chromatographic peak shape, and product ion distribution. A set of parent ion/product ion combinations (termed transitions) per peptide is tested; typically many more peptides and transitions are chosen than used in the final assay.
During assay development, positive correlation is required between the transition ratios obtained during MRM analysis and with the data from the discovery experiment or the data in the biomarker library. This ensures that a specific signal is unique to the biomarker protein. The peptide and transition list is refined at this point.
Next, analytical reproducibility of each peptide is assessed, by analyzing one preparation of the same sample many times in the mass spectrometer. Technical reproducibility of each peptide is then assessed by measuring the peptide performance from multiple preparations of the same sample. The peptide and transition list may again be refined at this point. Once all peptides have been selected they can be monitored in the assay.
A key feature of the MRM platform is that many peptides and their transitions can be multiplexed in a single chromatographic run. The ability to quickly develop a multiplexed assay for many proteins allows for facile initial biomarker testing. The timeline for this stage is typically four to six weeks.
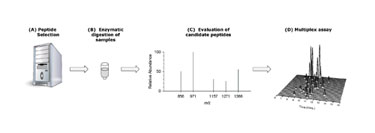

Figure 3. Stage-one assay development
Stage-One Biomarker Testing
The objective of stage-one biomarker testing is to confirm candidate biomarkers. The MRM assay is used to monitor the level of proteins in a sample set, often three to five times the size of the sample set used at the discovery stage. The assay is relative to a reference sample that is either a selected sample or a pooled sample. The level of each protein is calculated based on the response of the corresponding peptide relative to the level in the reference sample (Figure 4). Statistical analysis (p-value and false discovery rate) is applied to provide an assessment of the data. This will determine the significance of each biomarker. Ideally, many of the biomarkers would be confirmed after testing the larger sample set.
The data at this point can be used for internal decisions. If the requirement is for clinical validation or validation of clinical utility than stage-two assay development would be necessary. The timeline for this stage is typically two to four weeks.

Figure 4. Biomarker testing
Stage-Two Assay Development
While determination of the relative protein levels is sufficient for biomarker confirmation, protein concentration levels are required for biomarker validation. In order to achieve this, the relative quantitation assay from stage one is further developed to provide concentration values for each protein—often called absolute quantitation.
Isotopically labeled peptide standards (typically 13C or 15N labeled amino acids), that behave identically to the native peptide except for a shift in mass, are used to convert mass spectrometry data (analyte peak area) to protein concentrations. Separate calibration curves for each peptide included in the assay are established, each including six calibrators that cover the expected concentration range in the samples. The curves are run under the exact conditions established during assay development.
The MRM assay is validated on a fit-for-purpose basis; since small molecule MRM has been used for over 20 years for clinical samples, detailed and specific validation steps have been established that can be applied to MRM for peptides. The timeline for this stage is typically two to six months.
Stage-Two Biomarker Testing
The objective of stage-two testing is to validate the biomarkers. Data are reported as the concentration of each protein in each sample. This step of biomarker development involves hundreds of samples tested in batches. The assay is robust enough for the data to be highly reproducible over time.
NextGen Sciences is currently using the workflow described in this tutorial to develop protein biomarker assays for pharmaceutical and biotechnology companies. This has helped relieve the bottleneck of assay development. We have demonstrated that the MRM platform can be used to develop multiplex assays in the timelines presented in this workflow.
Ruth A. VanBogelen, Ph.D. ([email protected]), is the director of biomarkers and proteomics; Richard C. Jones, Ph.D.,
is head of mass spectrometry, and M. Walid Qoronfleh, Ph.D., is vp of business development at NextGen Sciences. Web: www.nextgensciences.com.