Doug Auld, Ph.D. Novartis Institutes for BioMedical Research
A new microfluidic approach can dramatically increase the number of data points per compound during screening.
ASSAY & Drug Development Technologies offers a unique combination of original research and reports on the techniques and tools being used in cutting-edge drug development. The journal includes a “Literature Search and Review” column that identifies published papers of note and discusses their importance. GEN presents one article that was analyzed in the “Literature Search and Review” column, a paper published in Proceedings of the National Academy of Sciences USA titled “High-resolution dose–response screening using droplet-based microfluidics.” Authors of the paper are Miller OJ, Harrak AE, Mangeat T, Baret J-C, Frenz L, Debs BE, Mayot E, Samuels ML, Rooney EK, Dieu P, Galvan M, Link DR, and Griffiths AD.
Abstract from PNAS USA
A critical early step in drug discovery is the screening of a chemical library. Typically, promising compounds are identified in a primary screen and then more fully characterized in a dose–response analysis with 7–10 data points per compound. Here, we describe a robust microfluidic approach that increases the number of data points to approximately 10,000 per compound. The system exploits Taylor–Aris dispersion to create concentration gradients, which are then segmented into picoliter microreactors by droplet-based microfluidics. The large number of data points results in IC50 values that are highly precise (±2.40% at 95% confidence) and highly reproducible (CV = 2.45%, n = 16). In addition, the high resolution of the data reveals complex dose–response relationships unambiguously. We used this system to screen a chemical library of 704 compounds against protein tyrosine phosphatase 1B, a diabetes, obesity, and cancer target. We identified a number of novel inhibitors, the most potent being sodium cefsulodine, which has an IC50 of 27 ± 0.83 μM.
Commentary
Understanding pharmacology stems from accurately measuring concentration–effect relationships. Generally, concentration response curves (CRCs) can be divided into three regions. These include a region of biological deficiency occurring at low doses at which no effect is observed (characterized by the “NOEL” point, the “no-effect-limit”), a region of biological efficiency at which a biological effect is observed, and a region of toxicity at which deleterious effects are observed at higher doses. Exactly how all three of these regions are modulated by a particular substance can lead to many types of complex pharmacology, including bell-shaped CRCs, partial agonism/antagonism, and steep or shallow Hill slopes—all of which can occur over several logs in concentration.
For compounds, several strategies are used to capture and measure the activity in bioassays. For large chemical libraries (>1 million compounds) high-throughput screening (HTS) is used in which libraries are screened at typically one concentration and “hits” are then followed up with 8- to 16-point CRCs. Another HTS approach (qHTS) uses a flexible plate archive allowing screening of medium-size libraries (up to ~350,000 compounds has been accomplished; see Michael et al., Assay Drug Dev Technol 2008;6:637–658) at up to seven concentrations which can capture complex pharmacological relationships during the screen. This article provides a system that uses droplet-based microfluidics to generate highly precise CRCs containing ~10,000 data points.
In the biochemical assays shown, these CRCs could be collected at rate of 1 every ~2.6 min. These high density data come from mixing the assay components with a compound concentration gradient that is formed inside the microfluidics (see Figure). An initial bolus of compound is injected that undergoes Taylor–Aris dispersion, which converts the rectangular concentration profile into a smooth Gaussian profile. To calculate the concentration in the gradient, an encoder fluorophore (a near-infrared dye, DY-682) is added to the compound sample. The detected concentration range can cover approximately 3 orders of magnitude, although the authors note that this could be increased to 4 or 5 orders of magnitude by using an improved fluorescence detector. The enzyme assay is dispersed into fluorinated oil, which forms small aqueous drops that are mixed with portions of the compound concentration gradient (see Figure). This procedure requires approximately 18 times less reagent than a traditional 8-point CRC. Therefore, this technology provides for highly precise measurements of CRCs with low reagent consumption.
The article shows collection of high resolution CRCs for β-galactosidase and PTP1B and the use of this technology to screen small libraries in a dose–response mode. The microfluidic system should be applicable to any fluorescent enzyme assay. In the article, a complex CRC was characterized for suramin against PTP1B in which inhibition and activation of enzyme activity occurred over a narrow range. The use of a concentration gradient as described here eliminates some issues with compound presentation such as carry-over on pintools. Adapting this to rapid response cell-based assays could enable the use of primary cells due to the low volumes involved. Increasing the throughput of the system would allow addressing larger libraries with very low reagent consumption, which would expand on the paradigm of qHTS that aims to reduce false positives and negatives by measuring the pharmacology of compounds at the level of the primary screen.
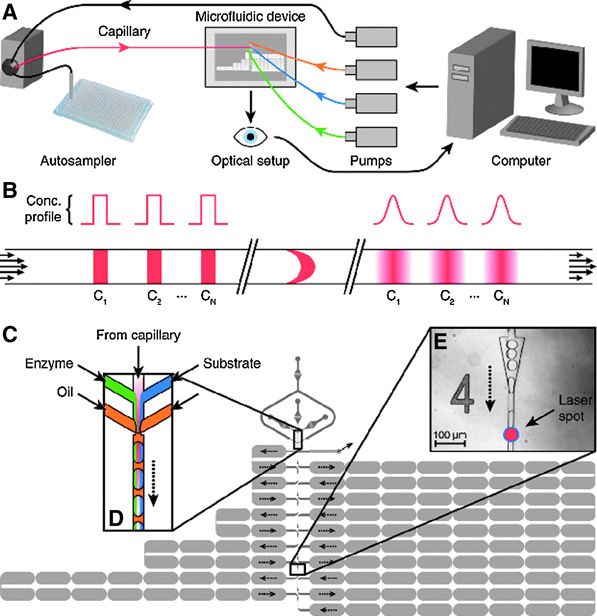
Figure. The microfluidic screening system. (A) Overview of the system. (B) Schematic showing how sequential injections of different compounds from the autosampler (C1 to CN) are transformed into smooth pulses by Taylor–Aris dispersion in the capillary. Each pulse gradually rises and then falls in concentration as it arrives at the subsequent microfluidic device. (C) Design of the microfluidic device (plan view) showing the two depths of channel: 25 µm (dark gray) and 75 µm (pale gray). Dotted black arrows show the route the droplets take through the device. (D) Schematic of the droplet production region of the device. Smoothed compound pulses from the capillary are combined with the enzyme (green) and the substrate (blue) and are then segmented into droplets by two streams of fluorinated oil (yellow). Each droplet contains a different concentration of the compound but constant concentrations of enzyme and substrate. (E) Light micrograph of one of the 10 analysis points with the position of the laser spot indicated just after the triangular droplet-respacing feature.
Doug Auld, Ph.D., is affiliated with the Novartis Institutes for BioMedical Research.







