May 15, 2012 (Vol. 32, No. 10)
Richard A. A. Stein M.D., Ph.D.
In Max Delbrück’s words, “Any living cell carries with it the experience of a billion years of experimentation by its ancestors.” This process of experimentation is reflected in the incredible complexity established by intracellular and intercellular networks. While a plethora of information has become available on the individual constituents building these networks, much less is known about their complex organization that results from modular interactions at different hierarchical levels.
To characterize existing systems and build novel ones, a new discipline, synthetic biology, has materialized at the interface between engineering and biology. While theoretical and practical developments that emerged over the past few decades catalyzed the establishment of this field, two recent advances in engineered gene networks brought an unprecedented expansion.
One of these, the “genetic toggle switch”, a synthetic bistable gene regulatory network built from two repressors and two constitutive promoters arranged in a mutually inhibitory network, allow independent, transient stimuli to switch between its two possible stable states.
In the second system, known as the “repressilator”, and comprising three transcriptional repressors that are not encountered together naturally, each repressor inhibits the transcription of the next one, establishing an oscillatory network among the three proteins. These landmark studies illustrated the possibility to build novel synthetic circuits that do not exist naturally, by using components that individually are encountered in separate biological contexts.
“We are trying to develop new parts and libraries of parts that others can use to drive the next stage of synthetic biology, and help implement industrial applications in biotechnology,” says Thomas Ellis, Ph.D., lecturer in synthetic biology at the Centre for Synthetic Biology and Innovation and the Department of Bioengineering, Imperial College London.
Dr. Ellis and colleagues are focusing on the budding yeast, an organism that is safe to work with, has been employed in biotechnology for thousands of years, and can easily be used worldwide. “The idea of switching from E. coli to yeast in synthetic biology brings us closer to realizable industrial outputs such as the production of high-value compounds,” says Dr. Ellis.
While the current, general view of synthetic biology is that of modular parts being assembled to provide new functions and rewiring the cells, this promises to change. “At some point in the future, we will be at a stage where we want to build an entire eukaryotic genome from simple parts, and we need to have some sort of understanding of how these parts behave and how to put all of them together,” explains Dr. Ellis.
A major effort in Dr. Ellis’ lab focuses on engineering promoters from the bottom up, to achieve precise gene-expression outputs and controllable regulation. Previously, Dr. Ellis and colleagues used a constitutive budding yeast promoter to illustrate that existing parts can be re-engineered and rationally diversified to design new parts and provide novel functions.
The investigators generated a synthetic promoter library, and subsequently illustrated the possibility of rationally constructing transcription activator-like protein effectors that can bind wild-type and synthetic promoters, a fundamental milestone toward generating gene networks.
Controlling the Metabolic Flux
Many bioindustrial applications need the metabolic flux to be redirected to optimize the production of the products of interest. This has been typically accomplished by gene knockouts, an approach that often may lead to suboptimal metabolic cellular states.
“We showed that gene knockouts are not required but we can, instead, dynamically modulate the genes of interest in order to appropriately redirect the metabolic flux between different pathways,” explains James J. Collins, Ph.D., professor of biomedical engineering at Boston University.
Dr. Collins and colleagues created a genetic switchboard, a panel of RNA switches that enable the simultaneous and independent regulation of multiple genes within a cell. This approach relies on an RNA-based riboregulator system that post-trascriptionally controls bacterial gene expression. In this system, two distinct promoters control the transcription of two RNA species, a cis-repressed RNA and a transactivating RNA.
The cis-repressed RNA sequence is engineered in front of the mRNA ribosome-binding site. When the mRNA is transcribed and produced, this cis-repressed sequence forms a stem and loop structure with the ribosome-binding site, and prevents ribosome docking onto the mRNA, inhibiting protein synthesis.
“We have shown that we can get 96% to 98% repression inside bacteria by using this system,” says Dr. Collins. A second switch can turn on the short, noncoding transactivating RNA that is designed to interact with the cis-repressed element, and this destabilizes the stem-and-loop structure, exposing the ribosome-binding site and enabling ribosome binding and protein synthesis.
“Because these are sequence-based elements, this approach affords an incredible flexibility for producing a large number of such systems, which can operate simultaneously and independently of each other inside the cell,” explains Dr. Collins.
These synthetic devices are tightly regulated, allow fast response times, and their modularity facilitates the independent control of multiple genes. “These developments open a number of biotechnology applications in terms of opportunities to effectively rewire and reprogram cells, to dynamically modulate gene expression,” explains Dr. Collins.
Using the same approach, investigators in Dr. Collins’ lab recently produced a programmable synthetic RNA-based kill switch in bacteria, in which the riboregulator system was used to deliver signals that affect inner membrane permeability and outer membrane integrity to combinatorially lyse the cells.
This switch was highlighted in the President’s Bioethics Commission report on synthetic biology. “This strategy is providing a much-needed safeguard for work with engineered organisms,” emphasizes Dr. Collins.
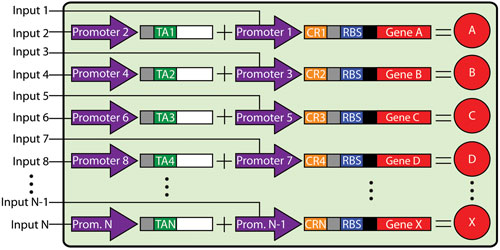
Genetic switchboard schematic: The higher-order synthetic device consists of modular, orthogonal riboregulators. The ellipses indicate the potential for further expansion. [Boston University]
For a different take on synthetic biology, click here.
“As we engineer biological systems to drive the production of high-value products such as biofuels, we want to be able to solve control problems and avoid the buildup of toxic products and unbalanced metabolites,” says James M. Carothers, Ph.D., research fellow at the University of California Berkeley and the DOE Joint BioEnergy Institute.
Recently, Dr. Carothers and colleagues reported the design of a dynamic sensor-regulatory system to produce fatty-acid-based products in E. coli. This system incorporated biosensors for fatty acid/acyl-CoA and hybrid promoters that responded to changes in fatty acid levels and to an exogenous inducer.
“We found that if we engineer genetic elements that we combine into the system, and couple the production of the intermediates to regulation in the pathway, we can increase production to almost 30% of the theoretical maximum,” says Dr. Carothers.
In addition to increasing the production of fatty acid ethyl esters as compared to a previous study, the host strain harboring this system was stabilized, a finding with important implications for the large-scale implementation of this approach. This represents an advantage over genetically modified microorganisms, in which the loss of gene function as a result of toxic metabolite accumulation opens ongoing technical and practical challenges.
Previously, Dr. Carothers and colleagues reported techniques to design RNA-based genetic control systems that could process cellular information and program the expression of large numbers of genes, potentially enabling the model-driven construction of metabolic pathways in various systems.
“Synthetic biology presents a lot of relevance for industrial biotechnology, for producing low- and high-value chemicals, and I think that in the future we will see new technologies and new approaches that will allow us to build larger and better systems,” says Dr. Carothers.
Researchers at the U.S. Department of Energy (DOE)’s Joint BioEnergy Institute have developed CAD-type tools for engineering RNA components that hold potential for microbial-based production of advanced biofuels and other goods now derived from petrochemicals. [Zosia Rostomian, Berkeley Lab]
Crossroads with Tumor Biology
Despite recent advances in tumor biology, molecular diagnostic and therapeutic approaches remain a challenging area. Recently, Yaakov Benenson, Ph.D., professor in the department of biosystems science and engineering at ETH Zürich, and colleagues designed a transcriptional/post-transcriptional synthetic regulatory circuit that can sense the expression pattern of a customizable battery of microRNAs, and illustrated its promise toward a new strategy with applicability in cancer diagnostics and therapeutics.
The investigators showed that this synthetic circuit can selectively trigger apoptosis in HeLa cells expressing specific combinations of markers expressed by cancer cells without affecting the surrounding, non-HeLa cell types.
“Our microRNA-based strategy provides a flexible approach that can use different marker combinations, and since different cancers may express very different combinations of markers, this approach can therefore be tailored to different types of tumors,” says Dr. Benenson.
This concept, enabling specific responses to be triggered based on defined and complex intracellular conditions, provides a therapeutic framework with applicability for other cellular states, and illustrates one of the applications of synthetic biology in cancer diagnosis and therapy. “Our long-term goal is to develop this into a therapy candidate but, like any potential treatment, it has to go first through all the stages, including cell culture and animal testing,” explains Dr. Benenson.
Optogenetics
“Our expansion of optogenetics as a field involved an approach to convert electromagnetic waves from the visible range of blue light into a sustained transcription response, which allows genes to be controlled by shining blue light onto them,” says Martin Fussenegger, Ph.D., professor of biotechnology and bioengineering at ETH Zürich.
The strategy used by Dr. Fussenegger and colleagues takes advantage of the signal cascade of melanopsin, the photopigment found in the photosensitive ganglion cells from the retina, which is most sensitive to blue light (~480 nm). Upon light exposure, melanopsin activates a G protein, triggering a subsequent signal transduction cascade that leads to an intracellular calcium surge.
This calcium surge was rewired to the nuclear factor of activated T-cells, which can initiate gene transcription from specific promoters. After demonstrating the functionality of this synthetic device in mammalian cells, Dr. Fussenegger and colleagues subsequently illustrated its use in a mouse model, where it enabled the light-induced expression of a transgene.
In a type 2 diabetes mouse model harboring these transgenic synthetic devices in intraperitoneal hollow-fiber or subcutaneous implants, the investigators reported that, upon exposure to light, the animals showed increased glucagon-like peptide 1 expression, reduced glycemic excursions, and decreased glycemic levels.
“This system is ready for clinical applications,” emphasizes Dr. Fussenegger. A key feature, fundamental for the clinical use of this synthetic device, is that it is fully humanized, and it does not contain any components that would elicit an immune response.
As individual parts, such as promoters, open reading frames, terminators, and transcription factors are combined to generate pathways and circuits, one of the goals of synthetic biology is to generate synthetic chromosomes and genomes, some of which have never existed before.
An important milestone was reached less than a year ago, with the publication of the first partially synthetic eukaryotic chromosome—that of the budding yeast. This advancement, along with other developments in the field, are signaling the beginning of a new era, one that provides a new level of scientific inquiry and promises to reshape medicine, biomedical research, and biotechnology.







