October 15, 2011 (Vol. 31, No. 18)
Kathy Liszewski
Advances Unravel Complexities, Shed Light on Therapeutic Targets, and Move Molecule to Fore
RNA is beginning to take its place among the movers and shakers of biology. For decades, it was considered a background player in biology, a static carrier of genetic instructions from big daddy DNA. Now, however, RNA and its analysis are coming into the spotlight in an ever-expanding arena. New players bearing acronyms such as piRNA, siRNA, shRNA, and ncRNA are now regulars in scientific lexicons (and, perhaps more importantly, Wikipedia).
Emerging trends and new advances in the field are reviewed in this article, including revolutionary tools for transcriptomic analyses, deciphering how pathogens subvert immune systems, and new modeling methods to better delineate RNA structure/function.
Piwi-interacting RNAs (piRNAs) are a new chapter in the emerging story of RNA biology. These small, noncoding RNAs (ncRNA) were discovered from a search for ncRNAs that interacted with the Drosophila signaling protein called Piwi. Two classes of piRNAs have been discovered. One set is abundant in germ cells and a second set is present in somatic cells and has a role in regulation of mRNA from early embryos and gonads.
The recent discovery of piRNAs in the central nervous system (CNS) was a surprise, reported Kenneth S. Kosik, M.D., professor and director, Neuroscience Research Institute, University of California, Santa Barbara.
“We are interested in how synapses of the CNS are locally regulated,” he said. “Since their discovery in 2006, the roles of piRNAs have continued to expand. We performed deep sequencing as a nonbiased approach toward searching for any category of small RNAs in the hippocampus, a brain region commonly used in studies of plasticity and studies of local translation in neuronal dendrites, an important facet of plasticity. Among the small RNA sequences observed were a set of piRNAs.”
Dr. Kosik and colleagues prepared small RNA libraries made by extracting RNA from male mouse hippocampus tissue to capture a large number of RNAs.
“We found a set of small RNAs in the size range of 24–31 base pairs, which is characteristic of piRNAs and quite distinct from the smaller ~21 nt microRNAs. We searched the piRNA database and found annotated sequences that corresponded to those in our sample. Then we confirmed our results by co-immunoprecipitating the piRNAs with the protein they bind to in the mouse (termed MIWI).”
The scientists also validated the results by performing in situ hybridization for several of the piRNAs. “Interestingly, we found that one piRNA extended out to dendrites; it was not confined to the neuron body. The dendrite is a site for a great deal of control over RNA and protein synthesis. These results may help us understand the plasticity of the nervous system.
“While we don’t know if piRNAs will have a role here, it certainly opens up the doors to explore if they are players. Minimally these studies revealed that piRNAs should no longer be considered as limited to the germ line. It is becoming increasingly clear that we need to think more broadly about them.”
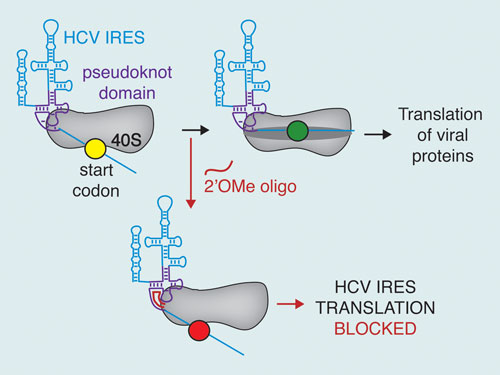
Hepatitis C virus RNA binds to 40S ribosomal subunits during viral infection. The start codon, depicted as a green, yellow, or red circle, is positioned in the mRNA binding cleft of the ribosome by the structure of the IRES pseudoknot domain. As this is a necessary step for viral protein synthesis, 2’OMe oligonucleotides were designed to disrupt the pseudoknot domain structure and effectively inhibited protein synthesis by the HCV IRES. [Jennifer Doudna, University of California, Berkeley]
RNA Sequencing
RNA sequencing is helping to revolutionize the characterization of gene expression. But, can the technology unmask the transcriptome of poorly understood and elusive pathogens such as Campylobacter jejuni? Yes, said Andrew J. Grant, Ph.D., senior research associate in the department of veterinary medicine, University of Cambridge, U.K.
“C. jejuni is the most common cause of bacterial food-borne diseases in the developed world. Its genome of ~1.6 megabases was sequenced around a decade ago, but we still understand very little about the pathogen.
“Our lab wants to better understand the roles of individual genes and proteins involved in its colonization, survival, and virulence as well as the general biochemistry and physiology of the bug. We utilized functional genomic approaches to do this, including sequencing the entire transcriptome of C. jejuni. At the time we began our studies, RNA sequencing of prokaryotes was very limited.”
According to Dr. Grant, high-throughput RNA sequencing can provide an unprecedented means to perform functional genomics. “Recently the transcriptomes of several eukaryotes and prokaryotes have been profiled using direct high-throughput Illumina sequencing of cDNAs, a process known as RNA-seq. We utilized Illumina high-throughput DNA sequencing to study mRNA expression levels of C. jejuni and then compared that to protein expression data.”
The Illumina platform has the capacity to generate tens of millions to billions of reads per run. The data can be aligned across splice junctions to assist characterizing novel transcripts, gene fusions, and isoforms. The technology is more sensitive than DNA microarrays that can miss ncRNAs.
To perform the studies, Dr. Grant and colleagues first isolated total RNA, subtracted out the 16 and 23 rRNA, and then reverse transcribed the resulting RNA into a cDNA library that could be sequenced on an Illumina Genome Analyzer.
“Our studies demonstrated the efficacy of high-throughput sequencing for profiling mRNA expression levels and for identifying genes that are differentially expressed. The technology also allowed us to identify a number of novel nonannotated ncRNA genes.”
Dr. Grant’s group is now developing the RNA-seq technology for use in mice infected with Salmonella enterica. “We hope to track expression changes in specific tissues or cells during active infection. Profiling both the transcriptome and the proteome is a powerful way to get at the bigger picture of what happens during infection by pathogens.”
Hepatitis C Therapeutics
The hepatitis C virus (HCV) genome consists of a small enveloped single-stranded RNA with a single open reading frame that produces one protein product, which is subsequently processed into active proteins. Its 5´ untranslated region (UTR) possesses an internal ribosome entry site (IRES) with a specific secondary structure that facilitates viral protein synthesis.
Jennifer A. Doudna, Ph.D., Howard Hughes Medical Institute investigator and professor of molecular and cell biology and chemistry at the University of California, Berkeley, studies HCV protein synthesis. “We have performed mutational analyses and determined crystallographic structures of functional parts of the IRES. We want to understand how the virus uses the IRES to control protein expression.
“The IRES contains a characteristic structure called a pseudoknot. If we can understand how this piece engages the ribosome, we may be able to develop a therapeutic to block that process.”
To better define the ribosomal binding site of the IRES, Dr. Doudna and colleagues created a series of RNA mutants of the IRES domain and tested their ability to interact with ribosomes.
“We utilized a rabbit reticulocyte lysate system to assess the effect of our mutants on translation.” Her laboratory also performed ribosomal toeprinting experiments that monitor how the viral RNA structure changes after binding the ribosome. The assay is based on inhibition of nucleotide elongation using reverse transcription.
“We found the pseudoknot conformation positions the RNA into the binding cleft of the ribosome and that the global structure contributes to its overall activity. Based on these findings, we next performed proof-of-principle experiments utilizing antisense 2´-O-methyl oligonucleotides to see if we could block this activity. Three of four 2´-O-methyl oligos showed potent inhibition of viral protein synthesis.”
Dr. Doudna’s laboratory is now also pursuing small molecule inhibitors that bind to the pseudoknot. “As we better understand how its structure contributes to its function, we will be looking at more targeted compounds to interfere with the pseudoknot.”
SHAPE
Most RNAs undergo extensive folding to form sophisticated based-paired secondary structures that become an integral part of cellular gene-expression machinery. Accurately deducing these structures is no small matter, yet this is a critical aspect of understanding structure/function.
“Structure is function,” noted Bruce A. Shapiro, Ph.D., principle investigator, Center for Cancer Research Nanobiology Program, National Cancer Institute-Frederick. “Predicting RNA structure in a rapid and accurate way remains a challenge.
“Several years ago, Kevin Weeks and colleagues at the University of North Carolina, Chapel Hill developed a methodology called SHAPE (Selective 2´-Hydroxyl Acylation analyzed by Primer Extension). This method provides quantitative nucleotide resolution data that can be utilized to determine RNA secondary structure.”
The problem, according to Dr. Shapiro, is how to interpret the data. “A low signal indicates that there is base-pairing, but it doesn’t disclose the base-pairing partner or the type of base pair.
“In order to better understand the complexities of SHAPE, we utilized seven RNAs taken from the Protein Data Bank whose structure had already been solved. We prepared the RNAs by in vitro transcription, and folded the RNA into its native structure. This was followed by chemical modification of the RNAs with N-methyl-7-nitroisatoic anhydride (NMIA).
“Subsequently these sequences were subjected to primer extension. Due to stops in primer extension caused by NMIA-modified unpaired nucleotides, one is able to obtain bands on gels indicating the positions of these stops. The intensities of these bands are then converted to SHAPE values.”
Dr. Shapiro and his colleagues arrived at several conclusions. “First, the SHAPE signal is impacted to a large extent by the base-pairing state of a residue. We also found significant correlations with base-pair stacking. By comparing the known structures with the SHAPE data, we developed a method that converts raw SHAPE values into probabilities of base pairing.”
Overall the new methodology enhances understanding of the SHAPE methodology and could be particularly important for pharmaceutical applications.
“One could imagine applying this enhanced SHAPE analysis to help in the development of therapeutics that target specific RNAs that are involved in disease processes. Although we are still developing predictive methods, ultimately such probabilistic models will aid the interpretation of RNA structure for basic science as well as pharmaceutical development.”
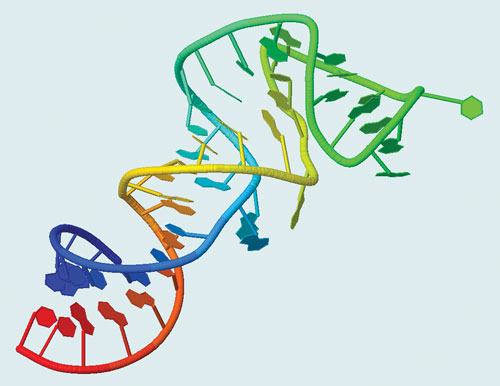
X-ray structure of the stem-loop II motif RNA element from the SARS virus genome (PDB ID: 1XJR) [Center for Cancer Research Nanobiology Program, NCI-Frederick]
Evolving Strategies
Examining gene expression via RNA analysis is a field on the move. Alice Lan, Ph.D., post-doctoral fellow in the laboratory of Donald R. Love, Ph.D., School of Biological Sciences, University of Auckland, provided a perspective on studies she performed previously on zebrafish.
“Over the years, whole mount in situ hybridization (WISH), another method of examining RNA transcripts, has been widely used among zebrafish researchers. Although WISH gives spatial patterns (in what tissues the transcripts are expressed), it is generally less sensitive and only semiquantitative (when coupled with imaging-analysis software). It is a usually a three-day protocol.
“However, qRT-PCR is a rapid, sensitive, and quantitative method for transcript analysis. It also allows multiplexing of differentially labeled probes so multiple transcripts can be examined at the same time.
“To examine expression during different stages of development simply requires extracting RNA from larvae at different time points. This is followed by cDNA synthesis and qPCR. The challenge is that one needs to validate the qRT-PCR results with another method. I think qRT-PCR and WISH experiments complement each other nicely.”
Dr. Lan is now applying her knowledge of RNA analysis to human clinical genetics. “We often need to correlate genetic variants (DNA) with gene expression.
“First, we have been emphasizing the use of multiple reference genes, rather than one reference gene, because how do you determine that a particular reference gene is stable without comparing it to other genes? Secondly, whenever possible, especially qPCR with SYBR Green, the amplicon should be sequenced to confirm their identity.
“Thirdly, disclosing detailed descriptions of RNA extraction protocol and data analysis is particularly important as the quality of RNA can greatly influence the efficiency of cDNA synthesis and qPCR. In terms of data analysis, readers should be able to easily figure out how the final ‘expression units’ are derived rather than second guessing what has been done to raw data.”
Tips for mRNA Capping
Capped RNAs can be made in vitro by incorporating cap analogs during transcription by enzymatic addition of cap structure after synthesis. Each method has advantages and drawbacks, according to G. Brett Robb, Ph.D., staff scientist at New England Biolabs:
- 5’-triphosphate transcripts can be capped using Vaccinia virus (VACV) mRNA capping enzyme with GTP and S-adenosyl methionine. This scalable approach allows for large amounts of RNA with m7GpppG cap structure to be generated.
- 5’-cap-specific labeling is possible using VACV. For high specific activity labeling, use the highest chemical concentration of radio-labeled GTP possible in the reaction.
- Co-transcriptional capping enables the incorporation of a variety of cap analogs including 3’-O-Me-m7GpppG (anti-reverse cap analog – ARCA), m7GpppG, GpppG, and ApppG.
- ARCA increases the yield of correctly capped mRNA since it incorporates in only one direction.
- Yields of co-transcriptionally capped mRNAs are influenced by many factors including concentration of NTPs and the ratio of cap analog to GTP. A ratio of 4:1 cap analog to GTP is suggested as a starting point.







