June 15, 2010 (Vol. 30, No. 12)
Catherine Shaffer
Technology Is Proving Its Worth in Assessment of Drug Efficacy, Toxicity, and Dosing
With ongoing advances in genomic technologies, clinicians are eager to move personalized medicine from the realm of theory to practice. Investigating whether the technology is mature enough, and if so, how to implement it, has become a hot topic of discussion in the field of personalized medicine.
Thus far, some of the most clinic-ready technology exists in the field of pharmacogenomics, where genes have been linked to drug efficacy, toxicity, and dosing. In addition to practical considerations regarding science and patient care, scientists are now tackling the thorny issues of policy and ethics.
The “Personalized Medicine in the Clinic” meeting held recently at Arizona State University in Phoenix featured dual tracks for discussing both the science and policy implications of the latest pharmacogenomic advances.
It’s an open question whether personalized medicine is mature enough to be of use in a real-world clinical setting. Some researchers are eager to begin using genomic data to predict patient risk, others doubt that it has practical utility yet. The Coriell Institute for Medical Research is seeking to answer this question in conjunction with the Coriell Personalized Medicine Collaborative (CPMC), a large, longitudinal study on the use of personal genome data in clinical patient care.
Intended as a model for ethical implementation of personalized medicine, the CPMC will ultimately enroll 100,000 participants who will be able to receive information about their genetic risks and significant pharmacogenomic information. The project will also collect long-term data about patient health and behavior changes with respect to their specific genetic variant results.
Michael Christman, Ph.D., president and CEO of Coriell, joined the institute in 2007 and initiated the CPMC. Of all of the types of personalized medicine data that will be screened, he has the highest expectations for pharmacogenomics. “You hear it debated whether or not personalized medicine is really ready for prime time yet. I think that, in the area of pharmacogenomics, it is.”
As an example of the difference that good pharmacogenomic data can make, Dr. Christman cited Plavix, one of the best-selling prescription drugs for heart-related ailments. A simple genetic test shows that it doesn’t work in about one quarter of people taking it. Instead, those people have negative medical outcomes. With appropriate screening for this genetic variant, many people could be saved the time, expense, and negative effects of a therapy that is not appropriate for them.
When this type of genetic mismatch with a drug is multiplied across the thousands of prescription drugs used in medicine, the costs are staggering. “We spend $300 billion a year on prescription drugs,” noted Dr. Christman. “About half of that is wasted, as most drugs only work in about half of the people who take them.”
Dr. Christman emphasized the uniqueness of the CPMC research study at the meeting, including free participation and genetic counseling; the establishment of many hospital partners; and the inclusion of nongenetic risk factors in the risk results. Currently, 4,500 people are enrolled in the study.
A committee reviews genetic information to confirm that it is actionable before it is reported to the participants. Participants can then choose what results they want to view and, further, what information to share with their physicians and decide whether to make any changes in lifestyle, behavior, or medication. A separate review board oversees pharmacogenomic information, to decide which gene-drug pairs are valid and actionable. “That’s where I think the largest clinical impact will be,” said Dr. Christman.

In 2007, Coriell established the Genotyping and Microarray Center. It performs genome analyses for the Coriell Personalized Medicine Collaborative™, which seeks to better understand the impact of genome-informed medicine and to guide its ethical, legal, and responsible implementation.
Wild Type vs. Mutation
Many personalized medicine strategies are centered on identifying variations or mutations in normal genes that affect the efficacy, dosing, or toxicity of a drug. However, it is also sometimes more useful to determine whether a patient is wild type for a given gene. That appears to be the case with nod2, an immune system protein with a mutant form that is implicated in Crohn disease. nod2 interacts with rip2, and it’s the nod2/rip2 binding complex that sets up the signaling cascade.
Derek Abbott, M.D., Ph.D., assistant professor of pathology at Case Western Reserve University, has been working on nod2 and rip2 for a number of years, and his group found that two drugs used in non-small-cell lung cancer inhibit rip2, and, therefore, could be effective therapies in diseases characterized by too much activity of the nod2/rip2 complex such as Crohn’s or other inflammatory bowel disease.
However, this would only be true for patients who had a functioning, wild-type copy of the nod2 gene. Because the Crohn mutant is not the only possible mutation for nod2, a test for the Crohn mutation will not suffice, and in order to benefit from these drugs, a separate test is needed to confirm wild-type status. “In medicine, there’s a general emphasis on the mutation, rather than on the normal protein,” Dr. Abbott said. “Genetic studies tell you what goes wrong with the gene, sometimes it makes sense to know if you’re wild type for a gene.”
Simulated Clinical Decision Making
Peter Tonellato, Ph.D., visiting professor and senior research scientist at the Center for Biomedical Informatics and department of pathology at Harvard Medical School, discussed his work on the practical use of an individual’s whole genome data and information. He specifically described the tools, technologies, and thought processes behind the Laboratory for Personalized Medicine at Harvard, through a set of model experiments projecting clinical parameters and predictions based on genome data.
Dr. Tonellato used an algorithm that maps an individual genome onto a reference genome, and then identifies the individual’s clinically important sequence variants. These variants are then used to generate useful clinical information for sample patients. In one example, they predicted an optimal warfarin dose for James Watson—whose whole genome has been published. The predicted therapeutic dose for Watson (if he were to hypothetically need this medication) is 4.9 mg per day. This is very close to the recommended best-practice clinical dosing of 5 mg per day. Consequently, one might argue that the genetic variant does not contribute to the best-care practice.
However, when the same exercise was performed on an anonymous African American male genome, the resulting optimal dose was 6.9 mg per day. That difference then becomes clinically significant. “We didn’t contrive it, we just ran it. This is a case when genome data is materially important in actionable healthcare information,” said Dr. Tonellato.
In a second model experiment, Dr. Tonellato calculated the relative risk of breast cancer for an anonymous female based on height, weight, age at first birth, and simulated variants of BRCA1 and BRCA2 for that individual. For this representative female with no variants in BRCA1 and BRCA2, the relative lifetime risk of breast cancer was 24.8%, compared to 8% for the general population.
That risk rose to 91.9% with one variant of BRCA1 or BRCA2 and up again to 99.8% for other possible variants in BRCA1 and BRCA2. “Boiling the whole genome down to a few genotypes demonstrates that it is valuable and impacts healthcare decisions dramatically.
One anticipates that an individual’s genome will provide a lifetime of enriched data and information that will help guide preventive measures, diagnosis, and optimal treatment.”
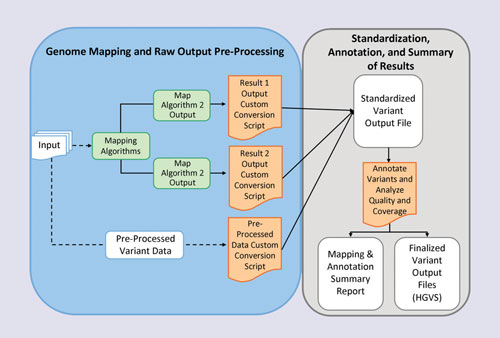
Scientists at Harvard Medical School are working on establishing practical uses of an individual’s whole genome data and information. They have employed an algorithm that maps an individual genome onto a reference genome and then identifies the individual’s clinically important sequence variants. These variants are then used to generate useful clinical information for sample patients.
New Liability Issues
Although the practical and scientific aspects of clinical pharmacogenomics are beginning to be understood, legal liability is an emerging issue with no easy answers. Gary Marchant, Ph.D., J.D., Lincoln professor of emerging technologies, law and ethics at Sandra Day O’Connor College of Law, spoke about the potential for lawsuits as personalized medicine approaches percolate into the clinic.
“Very few doctors have training in genetics to handle those issues; most of them don’t look at the primary literature, and it’s all subject to retrospective analysis in the courtroom,” Dr. Marchant said. For example, in Dr. Tonellato’s hypothetical case of warfarin dosing, if a physician chooses the FDA recommended standard dose of 5 mg per day and the patient has a stroke, will a jury then find the physician at fault for not ordering a genetic test and calculating his personalized dosage of warfarin?
“It seems to me this is really going to be a unique problem. With today’s rapid scientific and technical change, how do you have legal standards keep pace with that?” asked Dr. Marchant.
One possible outcome of the liability risk of pharmacogenomics in the clinic is overtesting, as doctors try to protect themselves from liability by ordering every possible test they can think of. The cumulative cost of overtesting is tremendous, and given the size of the genome and the expense of individual tests, the availability of pharmacogenomic information could lead to an explosion in healthcare expenses if the liability issues are not recognized and addressed.
Another cost of overtesting is burdening patients with information that may not be useful or actionable, leading only to anxiety.
Advanced Rare Cancer
In his talk on targeted therapy for rare cancer, Glen Weiss, M.D., associate investigator in the cancer and cell biology division at Tgen clinical research services, described work carried out at the University of Arizona College of Medicine on what makes a successful strategy for developing molecular targets and companion therapies in cancer.
The main problem in cancer is that not all patients (or the cancers they have) are the same. Yet currently, patients with the same cancer are primarily treated with the same type of chemotherapy. This therapy, not surprisingly, produces inconsistent results. Identifying markers that can be used to better target cancer with therapies most likely to work can improve results.
Some initial work that has come from Dr. Weiss’s group is in patients with advanced cancer. The team evaluated what potential targets a tumor might have for treatment—not necessarily those approved for a particular cancer. In conjunction with the Molecular Profiling Institute, which was a spinout from Tgen, they surveyed more than 110 patients, looking at gene expression in their tumors, as well as protein staining, picking out some common genes that predict response to certain drugs. Almost 100% of patients had a target for which there is a drug on the market.
Following up on that work, Dr. Weiss and his team conducted a prospective study on patients with advanced cancer. They analyzed tumor biopsy samples for genes and proteins in the cancer to try to identify a possible treatment, even if that treatment did not match their original cancer diagnosis. For example, if someone had breast cancer, and their tests indicated they could benefit from a colon cancer regimen, then they were given that treatment.
The results of this study were that 27% of the patients had an improvement of at least 30% greater duration of control of their cancer, compared to their last type of conventional cancer treatment.This confirms that genetic profiling can be useful for cancer treatment. The utility is especially attractive for rare cancers where there is less information to indicate which treatment is best.
Dr. Weiss also presented research into the use of locking signaling of Hedgehog pathway to improve outcomes for basal cell carcinoma (BCC), again showing some modest success in personalizing therapy for patients with advanced BCC. “So to have these real dramatic blockbuster successes with drugs, instead of just treating everyone with the more common cancers all the same, we’ll have to delve into the biology of what makes those individual cancers unique in each patient.
“We’re going to have to conduct much larger searches, paying attention to identified tumor targets, to find the small group of patients that will get great results with a treatment; rather than treat large groups of patients without evaluating their tumor targets, which results in minimal to no positive results.”
Psychiatric Therapy
Cancer is just one example of a category of disease whose treatment is affected by genetic variation. Psychiatric medications also have a great deal of variation in effect, dosing, and toxicity between individuals, and much of this difference has been linked to specific genes.
David Mrazek, M.D., chair of the department of psychiatry and psychology at Mayo Clinic, has focused his research on antidepressant and antipsychotic medications. His team tests for five gene variants. Three are drug-metabolizing enzymes, including the cytochrome P450 2D6 gene variants. Some 2D6 variants are completely ineffective, meaning the enzyme can’t metabolize medications like Paxil and pyroxitine. Of the two copies that each individual possesses, if both are ineffective, you have no 2D6 activity and can’t metabolize pyroxitine. Other classes of drugs affected by P450 enzymes include SSRI inhibitors and many antipsychotic medications.
A case study illustrated how genetic testing can help schizophrenics in the real world. Some individuals have a form of the gene that is highly inducible, and highly active when the person smokes, and smoking is a common habit among people with schizophrenia.
When a person with schizophrenia is admitted to a psychiatric hospital for treatment, they are typically not allowed to smoke, and thus their enzyme returns to a basal (noninduced) level. Once the patient is stabilized on an appropriate dose of olanzipine, an antipsychotic medication, they are released from the hospital and typically begin smoking again. This causes the enzyme to be induced once more, and destabilizes the illness, because the dose that was perfect in the hospital is now no longer enough to control symptoms.
Future
Increasingly, patients want to be tested and told which drugs will work for them. “We’re getting closer to that, but we’re not quite there,” Dr. Mrazek said. “We can say this is a medicine you shouldn’t take, because it’s going to be very hard to get the dose right, or this is a medicine you shouldn’t take because it might not help you, but we still don’t have enough of an indication to say we can find a medicine and it will have a 90 percent probability of working.”
Pharmacogenomics is a branch of personalized medicine that is rapidly entering clinical use. It has tremendous potential to help patients, but only if the information is organized and managed properly. Potential pitfalls include a too-narrow focus on specific mutations of SNPs, over- and undertesting due to lack of training or fear of liability, and information overload as the genome is explored. Simulation and prediction strategies can demonstrate how this information is useful in the clinic, but ultimately it will be necessary to test real genes and drugs in real patients in large-scale prospective studies such as CPMC.



