July 1, 2011 (Vol. 31, No. 13)
K. John John Morrow Jr. Ph.D. President Newport Biotech
Technology Improving a Range of Studies from Ion Channels to Pathogen Detection
Multiplex assays are based on single-analyte tests or low-to-midplex procedures that typically predate the rise of their multiplex versions. They may include vast numbers of analytes in a single assay and require specialized technologies or miniaturization to achieve a higher degree of parallelization. Multiplexing and related issues were the focus of CHI’s “World Pharma Congress,” which was held last month, and will be on the agenda at its “Next-Generation Sequencing Summit,” which will be held in August.
“About 1 percent of the U.S. population suffers from chronic neuropathic pain, and available treatments have numerous undesirable features,” says Michael Finley, Ph.D., who is currently assay development team leader at Merck & Co. “This unmet medical need has driven an intensive effort to identify specific pain treatments.”
For these reasons, Dr. Finley and his former colleagues at Johnson & Johnson Pharmaceutical Research & Development focused on the neuronal voltage-gated (N-type) calcium channel, an appealing molecular target for the development of novel analgesics.
The N-type calcium channel is localized in the presynaptic terminals of pain fibers in the dorsal horn of the spinal cord, substantiating its pivotal role in pain response. Previously, Dr. Finley and his coworkers engineered an HEK cell line using transduction of the appropriate gene sequences that expressed the N-type calcium channel.
This cell line can be evaluated using both a fluorescence-based calcium detection system for higher throughput, as well as an automated patch-clamp electrophysiology for higher quality, albeit at lower throughput. Using the HEK cells with known inhibitors of the N-type calcium channel, Dr. Finley demonstrated that the results between the two assays were comparable.
In a large-scale evaluation, the Finley team tested about 3,000 compounds in the calcium-imaging assay in a series of eight-point dose-titration experiments. A subset of 134 of these compounds with IC50 values <50 nM were further tested in three-point dose titration in the automated patch-clamp electrophysiology assay.
The two assays showed a modest positive correlation with an approximately tenfold right shift in potency in the automated patch-clamp assay. In order to better understand the potential in vivo efficacy of the compounds, a third assay measured release of neuropeptides from primary neuronal cultures.
Zinc-Finger Protein Array
A novel approach to the multiplexed detection of pathogens in blood has been developed by David Segal, Ph.D., associate professor at UC Davis Genome Center, in collaboration with Hsueh-Chia Chang, Ph.D., Bayer professor in the department of chemical and biomolecular engineering at the University of Notre Dame.
“Our device is built in three stages,” Dr. Segal explains. “First, the pathogens are separated and concentrated from the blood; secondly the organisms are captured using antibody microarrays; and, finally, a visual detection system using engineered zinc fingered proteins produces the diagnostic signal.”
PCR-based methods are quick and sensitive, but Dr. Segal and his coworkers were aiming for multiplexible technology for which DNA hybridization is less than ideal. So they settled on a DNA binding protein, a zinc-finger scaffold that was engineered to bind to specific target sequences. However, each finger typically recognizes only three to four nucleotides of DNA.
For this reason, the investigators employed combinatorial mutagenesis to generate tandem zinc fingered domains that have the theoretical capacity to bind to 18 contiguous pairs of DNA.
A further development in the technology is the SEER-GFP feature (sequenced-enabled reassembly), which consists of split protein domains that can reassemble into an active complex only in the presence of specific, double-stranded DNA sequences. The investigators used a split beta lactamase protein that was brought together when the ZFPs found their target DNA. With a functional enzyme in place, a fluorescent substrate will generate a color chain from yellow to red.
Since the system detects double-stranded DNA, it is unnecessary to rely on production of single-stranded molecules for detection, thus the approach is more straightforward and faster than conventional PCR technology.
“We are currently working on optimization strategies in which our engineered ZFPs will detect specific pathogen DNAs in the presence of millions of nontarget sites,” Dr. Segal adds. These improvements, which are under way, will be required to realize an actual pathogen detection scenario. Moreover, in order to realize an operational system it will be necessary to integrate the components into a lab-on-a-chip device.
Dr. Segal stresses the significance of a straightforward, point-of-care device that generates an unambiguous and rapid readout, operable by an untrained individual. This would allow diagnosis of a wide range of infectious diseases by family physicians and healthcare workers far removed from high-tech diagnostic laboratories.
Ligand-Gated Ion Channels
ChanTest focuses on ion channels as a primary target for drug discovery and safety, according to Glenn E. Kirsch, Ph.D., senior director for pharmacology and program management. “There are currently, a wide range of possibilities, including well-worn paths such as hypertension, areas still in early development such as pain, and largely unexplored conditions, of which autoimmune disease is a tantalizing representative.”
Current investigations at ChanTest include 5-HT3a, the ionotropic serotonin receptor, a target for the treatment of nausea and vomiting. It figures largelyS in pain disorders and drug addiction. A second area of concern is the nAChR α3/β4 or nicotinic acetylcholine receptor, a long-known target for Alzheimer disease and alcohol abuse.
Two other appealing choices are the ASIC1a acid-sensitive ion channel and the Nav1.7 voltage-gated sodium channel, which both respond to pain in damaged tissues. Finally, the HERG cardiac potassium channel, which performs as a safety antitarget for drug-induced torsade arrhythmias, is also under investigation.
The ion-channel protocols take advantage of genetically modified, transfected HEK293 and CHO cells that were subjected to automated patch-clamp assays to measure their responses to a number of drug candidates. A planar patch-clamp electrode array can be used for recording whole-cell ion-channel activity simultaneously from multiple cells in a 384-well format.
When coupled with robotic liquid handling, the technique can screen thousands of compounds per day against both ligand- and voltage-gated ion channels.
Dr. Kirsch describes this technology as “a second-generation Ion Works® (Molecular Devices) instrument with the capability to improve high-throughput ion-channel assays. This is brought about by extending the range of targets to include fast desensitizing ligand-gated channels via continuous recording during rapid compound addition.”

ChanTest claims to offer the world’s most complete library of validated human ion channel-expressing cell lines.
Allosteric Modulators
“Allosteric modulators are regulatory molecules that do not bind to the same site as the receptor’s natural ligand, but rather to alternative sites on the protein, modulating the binding and signaling of the natural ligand,” explains Patricia McDonald, Ph.D., associate director at the Scripps Research Institute. This characteristic offers unique opportunities for the development of a new class of molecular pharmacological agents.
Dr. McDonald and her team focus on the GPCR family of transmembrane spanning receptors that transmit signals from outside the cell to inside the cell through interactions with their cognate G proteins. They are activated by a diverse array of ligands that include light, odorants, hormones, and neurotransmitters. They are noted for their participation in many disease processes, making them important candidates for drug development.
Dr. McDonald has studied arrestin, which binds to activated receptors in a phosphorylation-dependent manner. Arrestin binding to the receptor blocks further G protein mediated signaling, terminating the signal, and desensitizing the receptor, an important regulatory process that prevents the cell from overstimulation.
“The CellKey System is a cell-based label-free technology that measures changes in cellular impedance (Z) using cellular dielectric spectroscopy (CDS),” Dr. McDonald explains.
CellKey measures the integrated response of the cell to receptor activation, permitting interrogation of the mechanism of action of candidate compounds. Following receptor activation, changes in cell morphology, cell adherence, and cell-to-cell interactions contribute to changes in Z. Using this technology the team has developed a strategy to monitor the efficacies of drugs on both receptor activation and desensitization in real time.
“We can detect complex receptor behavior in response to multiple classes of pharmacological agents including allosteric modulators and evaluate their potential therapeutic effectiveness providing valuable insight into how these agents may behave in vivo,” Dr. McDonald says.
Thomas Briese, Ph.D., associate director of the Center for Infection and Immunity and a faculty member at Columbia University, says that the center builds on a bioinformatics approach for sequence analysis and data evaluation, and employs MassTag multiplex PCR, GreeneChip cDNA microarrays, and next-generation sequencing to build an overall picture of the universe of pathogens that threaten global health.
Investigators from the center have created sequence analysis algorithms that deliver automated retrieval, filtering, and alignment of sequences from databases. The goals of the program include characterization of microflora, investigation of outbreaks and pathogenesis of chronic diseases, design and construct field systems for on-site deployment in the developing world, and assessment of animals for potential human-risk agents.
The need for such efforts was amply demonstrated by the West Nile outbreak in New York in 1999, the 2003 SARS outbreak, and more recently, in a nosocomial hemorrhagic fever cluster caused by the Lujo virus.
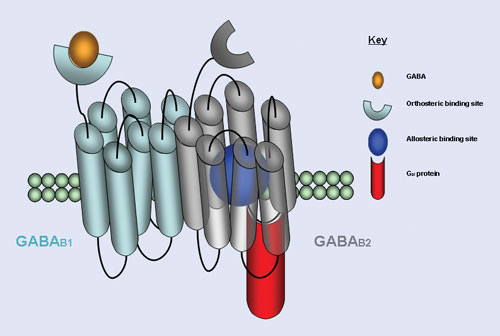
GPCR allosteric modulators bind to sites topographically distinct from the binding site of the receptor’s natural ligand, providing a novel approach to identify new GPCR-focused therapeutics devoid of side effects, according to researchers at Scripps Research Institute.
Pathogen Detection and Discovery
“The discovery process for new viruses is still in its infancy,” says Dr. Briese, and indeed, the causative agent is never identified in a vast number of enteric and respiratory infections.
“There are approximately 50,000 vertebrate species. Even if each has only 20 endemic viruses, we can expect the existence of one million vertebrate viruses. Given the current state of knowledge, this means that more than 99 percent of vertebrate viruses remain to be discovered.”
The center’s pathogen discovery platform expands on classical culture-based methods by bringing the speed and sensitivity of modern molecular diagnostics to bear. As Dr. Briese explains, to balance costs, sensitivity, and breadth, clinical specimens are first screened by multiplex MassTag PCR, which uses small photocleavable reporter tags detected by mass spectroscopy to test for up to 30 known pathogens in a single assay.
If no agent is identified or if no candidate agents are obvious, samples are analyzed by GreeneChip microarrays, which can identify viruses, bacteria, fungi, and parasites for which sequence information is available. If an agent is not indicated by either method, unbiased high-throughput sequencing analysis, the only current method capable of detecting uncharacterized agents, is used.
In all cases, analysis ideally culminates in sequencing and identification. However, proving causation requires extensive subsequent analyses and isolation of the candidate pathogen through classical methods.
According to Yu-Tsueng Liu, M.D., Ph.D., assistant adjunct professor, medicine at UC San Diego, the multiplexing technologies, which have co-evolved with the human genome project, will have a great impact in the next revolution of pathogen diagnosis. Because nearly all current tests rely on prior knowledge of the pathogen and the experience of the clinician, microarrays and next-generation sequencers will bring a vast assortment of new options and the possibility of precise identification, without which effective treatment cannot proceed.
K. John Morrow Jr., Ph.D. ([email protected]), is president of Newport Biotech and a contributing editor for GEN.







