June 1, 2010 (Vol. 30, No. 11)
Vicki Glaser Writer GEN
Optimizing Separation Speed and Efficiency Gives Tremendous Boost to Critical Parameters
The plasma glycome represents a potential source of disease biomarkers. Changes in glycosylation of glycoproteins are often associated with inflammatory and other acute-phase disease responses and may provide a tool for early disease detection. In breast cancer, detectable glycan changes associated with inflammatory activity in tumors typically precede metastatic activity.
It is also possible to characterize changes in the glycosylation of proteins from cells shed by tumors, and such glycan-based biomarkers might prove useful for detecting circulating tumor cells. In prostate cancer, although the relative level of prostate specific antigen (PSA) is a commonly used biomarker for prostate cancer screening, it is neither sufficiently sensitive nor specific.
Pauline Rudd, Ph.D., NIBRT professor of glycobiology at University College, Dublin, and colleagues at the National Institute for Bioprocessing Research and Training (NIBRT), have identified a subset of PSA that has an altered glycosylation pattern in the prostate cancer setting.
“Altered glycosylation of acute-phase proteins and IgG suggests that cancer regulates certain pathways favoring cancer cell survival,” Dr. Rudd observes. In ovarian cancer, for example, N-glycosylation changes in serum glycoproteins have been associated with a decrease in galactosylation of IgG and an increase in sialyl Lewis X.
Dr. Rudd’s group has developed a robotic HPLC-based platform designed to perform quantitative glycan analysis and glycoprofiling to support both research on glycoproteins and bioprocessing applications from early-stage design through process analytical technology and biopharmaceutical production. Their work has led to the development of a glycome database called GlycoBase that contains structures of plasma glycans compiled from 1,008 individuals. The database includes more than 400 glycan structures, including 117 from the plasma glycome.
Dr. Rudd’s group used HPLC analysis of fluorophore-labeled glycans combined with sialidase digestion to separate and quantify individual glycans into 33 chromatographic peaks. Dr. Rudd reports “surprisingly large biological variability at the population level.”
GlycoBase provides the elution positions for labeled N-glycan structures and the predicted products of exoglycosidase digestion. The database is being used to study the variability, heritability, and lifestyle factors that affect the composition of the human plasma N-glycan.
Dr. Rudd cites several advantages of using HPLC for glycome analysis, including the sensitivity and reproducibility of the technology and its ability to yield quantitative results. Furthermore, HPLC resolves molecules on the basis of their hydrophobic/hydrophilic properties and is able to analyze both charged and neutral glycans simultaneously and to distinguish between isomers.
Dr. Rudd’s group automated the HPLC analytical workflow in a 96-well plate format from sample processing through data interpretation. N-linked sugars can be analyzed at concentrations as low as the femtomole range and isolated from microgram quantities of glycoprotein mixtures using traditional amide columns.
Technology Heats Up
In May, Waters announced a partnership with NIBRT focused on creating a database for glycan analysis based on the company’s UltraPerformance Liquid Chromatography® (UPLC®) platform. NIBRT will develop, maintain, and license the database, which should launch in 2011 and will be co-marketed by NIBRT and Waters.
The database will contain chromatographic retention times for sets of glycan structures associated with biotherapeutic compounds. It is intended for use by biopharmaceutical manufacturers as a tool for evaluating the structure of glycosylated proteins and as a means of monitoring product integrity and bioprocess control. Researchers will use the Acquity UPLC system with an ethylene bridged hybrid glycan separation column and fluorescence detection to separate the glycans released from glycoproteins in the form of 2-aminobenzamide derivatives.
Peter Carr, Ph.D., professor in the department of chemistry at the University of Minnesota, Minneapolis, describes three main technological advances in HPLC “that have come together to push it forward”: the instrumentation that enabled ultrahigh pressure LC (UHPLC), which was based on the work of Jim Jorgenson, professor in the department of chemistry at the University of North Carolina, Chapel Hill; the development of micropellicular particles, composed of a solid inner core that is chromatographically inactive and impermeable, surrounded by a thin crust of porous material that is chromatographically active; and the use of high temperatures, which has been the focus of Dr. Carr’s research.
Dr. Carr describes temperature as “the third dimension in HPLC.” He is using higher temperatures to decrease fluid velocity, which yields an increase in the flow rate without the need for higher pressures and raises the diffusion coefficients to enhance mass transfer and limit peak broadening.
To achieve the benefits of increased temperature without compromising the quality of the chromatographic separation, Dr. Carr’s group has developed thermally stable stationary phases—zirconia-based and hyper-crosslinked silica-based phases. They are using these media to perform ultrafast high-temperature LC (UFHTLC) as the second dimension in comprehensive 2-D LC/LC for analytical applications, in which all of the material that exits the first column goes into the second column.
The increased speed enabled by the high temperatures is not only necessary to handle the large number of samples produced by the first-dimension LC separation, but it also provides another, unexpected benefit. “As you do the second dimension faster, your overall resolving power improves,” says Dr. Carr. If the speed of the second dimension is less than optimal and, consequently, “you don’t take enough fractions out of the first dimension,” the fractions will be too big, remixing can occur, and “you lose information that is already there.”
By running the second dimension at optimal speed, which Dr. Carr estimates from experimental data as a separation every 15–20 seconds, it is possible to achieve maximal peak capacity.
In 2-D LC/LC, the cycle time of the second dimension has a greater impact on the overall 2-D peak capacity than does the first dimension, explains Dr. Carr, and he recommends the use of strategies such as higher temperatures to optimize the speed of the second dimension rather than options such as microparticle-based media combined with high pressures, or long monolithic columns, to accelerate the first-dimension separation.
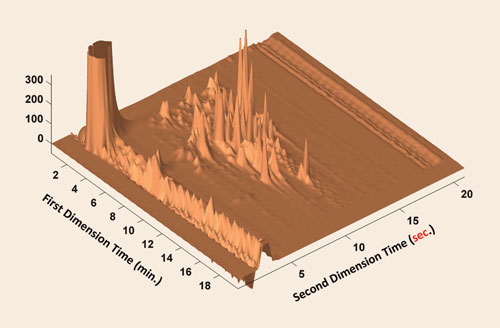
2-D LC of BSA tryptic peptides: Peak capacity of ~1,100 at a rate of 1 peak/second [Peter Carr, Ph.D., University of Minnesota]
Core-Based Particle Design
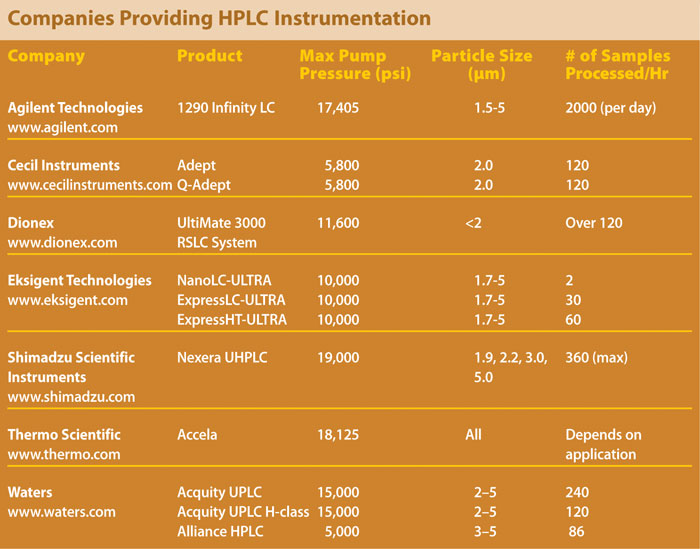
The first major advance in faster HPLC led to the development of UHPLC instrumentation, marked by the introduction of Waters’ Acquity UPLC technology for high-speed chromatography in 2004. UHPLC systems, which include Agilent Technologies’ Infinity LC and 1200 Series Rapid Resolution LC (RRLC), and Shimadzu Scientific Instruments’ Prominence UFLC, achieve faster analysis speeds without loss of chromatographic fidelity.
The combination of UHPLC technology and sub-2 micron particles maximized the efficiency of these systems. Agilent, for example developed the Zorbax Rapid Resolution HT columns, which are packed with porous 1.8-micron particles; the Zorbax line includes the company’s 300StableBond wide-bore 300Å columns.
The third main advance described by Dr. Carr was the development of smaller pellicular particles that offer a short diffusion distance and the combined benefits of the pressure drop created by a 2.8-micron particle and the narrow peaks achieved with sub-2 micron particles. Compared to sub-2 micron particles, the pellicular particles do not require high back pressures and can be used on conventional HPLC instruments.
Advanced Materials Technology’s Halo™ media is based on the company’s Fused-Core™ technology. Sigma-Aldrich recently added the Peptide ES-C18 column, based on a 160Å fused-core particle design, to its line of Ascentis® Express columns. Agilent offers its Poroshell columns with particles containing a solid silica core and porous outer shell.
Phenomenex introduced its 2.6 micron Kinetex core shell product in August 2009. The particle contains a 1.9-micron solid core and a 0.35-micron porous outer layer and offers pressure stability up to 600 Bar in 4.6 mm internal diameter columns and up to 1,000 Bar in 2.1 mm internal diameter columns. The company’s 1.7-micron core shell particle has a 1.25-micron core and a 0.23-micron shell and provides pressure stability up to 1,000 Bar. Both particles are available as C18, pentafluorophenyl (PFP, for separating isomeric and halogenated compounds), and hydrophilic interaction LC (HILIC) columns.
“If a customer is using a 5-micron particle, switching to Kinetex provides a threefold increase in efficiency; switching from a 3-micron particle provides twice the efficiency,” says Terrell Mathews, head of product management at Phenomenex. Many customers are using the core shell particles not only to increase throughput, “but to increase chromatographic resolution and sensitivity.”
The core shell technology “will be our platform for future development,” Mathews says. The company plans to develop new phases and new chemistries based on these particles. Customers are asking for a range of larger particle sizes, additional chemistries, and a selection of different column dimensions, he notes.
Scale-Down Strategies
The use of HPLC to separate molecules in a mixture for analytical purposes has benefited in recent years from advances in nanotechnology that have enabled greatly scaled down applications that reduce sample and reagent needs, speed processing, and lower costs. In application areas such as proteomics research and quantitative analysis of biomarkers or therapeutic drug levels in preclinical or clinical tissue samples, sample scarcity is driving efforts to downsize separation processes.
Frantisek Svec, Ph.D., organic and macromolecular synthesis facility director at the Molecular Foundry, Lawrence Berkeley National Laboratory, and colleagues have developed a strategy for producing second-generation porous polymer monolithic columns with modifiable properties in capillary columns.
Monoliths allow for faster separations than traditional packed columns due to higher flow rates and mass transfer that is driven by convection rather than diffusion, explains Dr. Svec. They offer an additional advantage: “you make them in situ and avoid the tedious, complicated packing” needed to make a capillary column or microfluidic separation chamber using traditional media.
Although polymer-based HPLC monoliths date back to the early 1990s, early forms had relatively small surface areas and were primarily suitable for separating large molecules. Dr. Svec’s group has developed a two-step approach to control the monolith’s porous properties: they first prepare a monolith with a typical porous structure and then perform in situ hyper-crosslinking reactions to create nanopores and mesopores.
The group has optimized several methods for modifying the surface chemistry of these monoliths, including co-polymerization with monomers that provide a desired functionality, chemical modification, photografting of polymer chains onto the pore surface and the addition of nanoparticles such as nanotubes or carbon 60 buckyballs.
“We are also engaged in developing flat monolithic devices,” what Dr. Svec describes as “a new incarnation of thin layer chromatography.” These are 50-micrometer thin super-hydrophobic monolithic layers on glass substrates in which virtual channels are created using photolithography. This technique yields a microfluidic device without the need for microfabrication.
A recent paper described the use of porous polymer monolithic layers to separate mixtures of six peptides and three oligonucleotides in one minute using a thin layer electrochromatography technique (Woodward, S.D., et al., Analytical Chemistry, 2010). Separation was achieved using pressurized planar electrochromatography with a negatively charged layer prepared by co-grafting 2-acrylamido-2-methyl-1-propanesulfonic acid and 2-hydroxyethyl methacrylate.
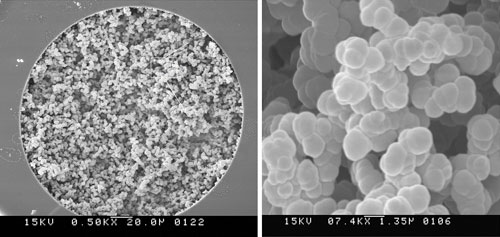

Researchers in the Molecular Foundry at Lawrence Berkeley National Laboratory have developed a strategy for producing second-generation porous polymer monolithic columns with modifiable properties in a capillary column. A scanning electron micrograph of a porous polymer monolith in a fused silica capillary column is shown.
Robert Kennedy, Ph.D., professor of chemistry at University of Michigan at Ann Arbor, and his team are optimizing droplet technology for collecting small protein fractions as they come off a capillary HPLC column and applying a segmented flow technique to capture nanoliter fractions for subsequent offline characterization using mass spectrometry. This scale-down technology is designed to overcome the effects of flow and dispersion that can cause protein fractions that have been separated by capillary HPLC and collected in capillary tubes to remix before delivery to the mass spectrometer.
By decoupling the capillary LC separation from MS analysis, Dr. Kennedy is able to collect nanoscale protein fractions that can either be fed directly into an MS system, divided, or modified prior to characterization.
Dr. Kennedy and colleagues derived the idea for applying segmented flow and droplet technology to the HPLC/MS interface from techniques designed to manipulate aqueous plugs segmented in oil. In another paper in Analytical Chemistry (2010, in press) Li, Pei, Song, and ‘Dr. Kennedy describe their method for segmented flow, which is achieved by slowing the HPLC flow rate when a sample of interest comes through the capillary LC, followed by collection of nanoliter fractions by forming plugs of effluent divided by an immiscible oil layer.
These plugs are stored in tubing for off-line delivery to an electrospray ionization mass spectrometer. The oil is siphoned away as each plug reaches the tip of the tubing, before injection into the ionization chamber. The authors have demonstrated no loss of chromatographic resolution using this off-line analytical technique.
The ability to decouple fraction collection and MS offers several advantages, according to Dr. Kennedy, the first being more time to do MS analysis of select fractions even if the peaks coming off the chromatogram are narrow and follow in rapid succession. Conversely, if the aim is to maximize the efficiency of MS analysis, decoupling allows for the collection of small batches of HPLC fractions that can then be run on the mass spectrometer while the next batch of fractions is coming off the HPLC, minimizing instrument downtime between analyses.
Another benefit of this decoupling technique and droplet technology is the potential to perform multiple parallel analyses on a single sample by dividing a fraction into daughter droplets and running individual daughter droplets on MS, NMR, or other types of analytical systems.
Decoupling also allows for manipulation of the protein captured in a particular fraction before it is analyzed. Dr. Kennedy’s group is experimenting with digestion of a protein into its component peptides followed by peptide analysis, and with derivitization techniques, both performed directly in the droplets.

A novel fraction collector for capillary LC has been developed at the University of Michigan at Ann Arbor. LC effluent is pumped into a tee with a flow of oil delivered to a second arm. The oil segments the LC effluent into plugs with nanoliter to picoliter volume for storage and manipulation. Photo on the lower left shows aqueous solution being segmented. A series of plugs stored in a capillary tube is shown on the right.







