February 15, 2014 (Vol. 34, No. 4)
Nicholas Miliaras, Ph.D. ATCC
The digital polymerase chain reaction (dPCR) consists of many discrete, small-volume reactions to represent genetic information from a single sample.
If a DNA template is sufficiently diluted so that some of these reactions contain no target molecules, while others contain discrete amounts, statistical analysis can then be used to quantify the exact number of target molecules in the sample. Therefore, dPCR can be used to accurately determine the number of nucleic acid molecules in a sample without comparison to a reference standard, provided that many reactions are performed in parallel.
Because the reaction volumes for dPCR are generally very small, and the template must be sufficiently dilute to achieve single-molecule resolution, nucleic acids can be isolated from minimal quantities of specimen. The dilution of template also removes many of the inhibitors of PCR that are present in the original sample, further improving accuracy and efficiency.
These properties make dPCR a superior diagnostic tool for the clinic, as well as any application where extreme sensitivity or precise quantification is essential, such as identifying mutations or copy number variations in tumor cells, or examining gene expression at the single-cell level.
While dPCR uses established thermal cycling technologies and enzymes, and fluorescent probes are used to read the amplification signal, as they are in qPCR, its essence lies in the partitioning of the sample into many small reactions. The number of reactions that can be run depends on the equipment and the process it uses for partitioning.
There are two types of dPCR machines currently available: chip-based and digital droplet PCR (ddPCR). For chip-based dPCR systems, such as the LifeTechnologies QuantStudio™, the sample is mixed with reagents and loaded into individual reaction chambers in plates that are about the size of a microscope slide. The reaction mixture is loaded into these small partitions by hydrophobic/hydrophilic interactions and capillary forces and the machine monitors the reactions as they occur.
In ddPCR, employed by Bio-Rad and RainDance machines, the samples are first mixed with the reagents and dispersed into nanoliter-sized droplets. The droplets for each sample are then placed in a tube and PCR is performed in a thermocycler. A droplet reader then detects a fluorescent signal to determine whether or not reactions have occurred. Unlike chip-based systems, ddPCR does not use physical partitions to separate the reactions, but the properties of the droplets themselves. For further discussion on dPCR technologies, please see Nature Methods 9, 541-544 (2012).
A Chip-Based System
The QuantStudio™ 3D dPCR system became available from Life Technologies in June 2013. According to Stephen Jackson, Ph.D., associate director of product applications at Life Technologies, approximately 20,000 reactions can be run on a single, dime-sized, silicon-wafer chip in this system. All of the partitions are of identical size. This system offers the additional advantages of a single-step workflow, where a 15–20 µL sample of the template DNA and reaction mix is loaded and processed by the system, which quantifies it as copies of nucleic acid per µL.
The QuantStudio™ 3D dPCR system is compact; the thermal cycler fits easily on the benchtop; and the reader instrument is “the size of a shoebox.” It is also priced to be within the budget of most research scientists, with a system that includes the QuantStudio™ 3D Digital PCR Instrument, Applied Biosystems® GeneAmp® 9700 dual flat-block thermal cycler, chip loader, and 384 digital PCR chips for under $50,000.
While there is a risk of cross contamination for any PCR system that processes a large number of reactions, Dr. Jackson believes that the risk will be reduced in this system, since each sample is loaded on a separate chip and the reactions are physically separated by partitions. With an eye toward the certification of dPCR as an in vitro clinical diagnostic, Dr. Jackson adds that “no mixing of patient samples is something the FDA will pay attention to.”

The Life Technologies™ QuantStudio™ 3D Digital PCR System uses a microfluidic chip that partitions samples based on hydrophobic/hydrophilic interactions and partitioning valves.
Detection of Rare Noncoding RNAs
The dilution of template into a very large number of reactions makes dPCR ideal for identifying targets that would otherwise be undetectable in a sample containing many thousands of copies of a genome, such as preoncogenic mutations. David Dodd, Ph.D., a postdoctoral fellow in Dr. David Corey’s laboratory at the University of Texas Southwestern Medical Center in Dallas, uses a Bio-Rad QX100 ddPCR System to detect long noncoding RNAs (lncRNAs) in the nucleus that regulate gene activity. Differences in the expression of lncRNAs have been associated with several diseases.
According to Dr. Dodd, dPCR is essential for this area of research, where knowledge of the exact number of a particular lncRNA can reveal whether it acts on a gene on the same chromosome (cis), or a different one (trans). “You get real numbers really quickly,” Dr. Dodd says, “and you don’t need to rely on a standard curve, as in real-time PCR, or quantitative PCR (qPCR), which can introduce problems due to structural variations in the RNA targets.”
Still, he does not think that the ddPCR system he uses will replace qPCR; although dPCR is important for his work, he mentions that few of his colleagues in academia use it because of the high cost of the equipment. Because of this, he sees it more as a service that would be offered by core facilities than something each lab would purchase. There are also “lots of hands-on steps” to prepare the droplets, he says, further adding to the inconvenience.
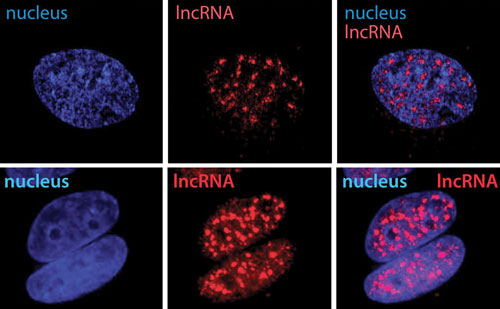
The distribution of lncRNAs within the cell can be determined by cellular fractionation followed by dPCR. These results can be verified by FISH, as in this image, where it is shown that many lncRNAs permanently reside in the nucleus. Moreover, the absolute number of transcripts per cell can be determined by dPCR, and this number gives clues as to their mechanism of action. Low-abundance ncRNAs are found to be cis acting while high-abundance ncRNAs are likely to act in trans. [K. Gagnon and L. Li, Corey Lab, UT Southwestern Medical Center]
Detection of MRSA Infection
While researchers welcome the increased precision and sensitivity of dPCR, perhaps its most significant impact will be in diagnostics, where a rapid detection of viral or bacterial infection, or mutations present in a type of cancer, are essential for determining the best course of treatment.
Donna Sullivan, Ph.D., professor of medicine at the University of Mississippi, has developed a technique for the rapid identification of methicillin-resistant Staphylococcus aureus (MRSA) infection, a major concern in hospitalized patients. Dr. Sullivan and colleagues have developed a ddPCR in vitro diagnostic (IVD) assay to detect the presence of the methicillin resistance (mecA) and Staphylococcus aureus-specific genes in extracts derived from patient nasal swabs using gene-specific probes and a Bio-Rad QX100 ddPCR system.
In the April 2013 issue of the Journal of Clinical Microbiology, they reported that in this assay ddPCR’s sensitivity, specificity, and kinetics were nearly identical to those in the qPCR. Also, ddPCR could detect MRSA with 95% confidence from a swab containing as few as 90–100 CFU of the strain. Significantly, the limit of detection for mecA was not affected by the presence of up to 105 CFU methicillin-sensitive Staphylococcus aureus (MSSA) or another potentially confounding strain in the reaction mix. Their results also agreed with the Cepheid MRSA GeneXpert assay, a qPCR assay and FDA-approved IVD for MRSA infection.
Dr. Sullivan is a strong proponent of ddPCR as a diagnostic tool. She is collaborating with others to develop ddPCR IVDs to identify KRAS mutations in FFPE samples for colorectal carcinoma. She finds that the strength of ddPCR is that it is “more sensitive [than qPCR] and could give results with much smaller sample volumes.” Also, the nasal swab samples for the MRSA study did not require an additional spin column purification step following extraction.
However, she identified potential hurdles to the wide adoption of ddPCR, including an insufficient number of users having access to the system. As with Dr. Dodd, she had issues with the high cost and time required to make droplets for ddPCR.
Single-Cell Genomic Analysis
Most diseases start at the level of the single cell—whether the initiating event is a preoncogenic mutation giving rise to a tumor, or a viral or bacterial infection—a single cell is the first to generate or receive a signal that initiates a series of events resulting in illness. How can experimental techniques that are designed for populations of thousands of cells be useful for understanding what is going on in one cell?
Since the principle of dPCR is to amplify DNA at the single-molecule level, it is ideally suited for genetic investigation of single cells. Richard Mathies, Ph.D., professor of chemistry at the University of California, Berkeley, has developed a method for extracting genomic DNA from single cells for further analysis using dPCR.
“The fundamental idea is that every cell is different. Cells are not a Gaussian distribution about the mean, so how do you identify their individual differences?” Dr. Mathies asks. To answer this question, his team harvests a population of cells and mixes them in an emulsion that contains low-melting agarose and reverse-primer-conjugated beads. This mixture is then dispersed into droplets that contain encapsulated single cells. Following cell lysis, forward primers and a reaction mixture are introduced in an emulsifying oil and thermally cycled, so that specific genetic mutations can be quantified by flow cytometry, or the target can be amplified again for sequencing.
Dr. Mathies’ approach for single-cell genomic analysis has many powerful applications ranging from forensics to identifying which splice variants of a gene are expressed in a single cell. He notes that probes for up to 20 disease-related genes could be included in a single droplet, for investigating pathology at the level of the individual cell.
Dr. Mathies also observed challenges to widespread and effective use of single-cell digital PCR. First among them is “courage of convictions” as he puts it. “We tell everyone when they are a grad student that when something doesn’t work, you need to add more of it…increasing the concentration is always the answer. This is not true if you want to see single molecules. You need to use less, or it will be lost in the noise.”
Another major factor related to this is lab cleanliness, since even minute amounts of DNA can contaminate the results. “The real test is seeing nothing,” states Dr. Mathies. “You must have zero background.”
Technical challenges for single-cell dPCR include the droplet generator technology (only one exists—in Dr. Mathies’ lab) and suitable emulsifying oil. “The suppliers keep changing or they get acquired. Then the formulation changes, and it doesn’t work anymore.”
An Exciting Time to Go Digital
Digital PCR allows for the detection of single genomes, a feat that was nearly impossible a few years ago. As Dr. Mathies observes, this is possibly the first instance in modern scientific history where it is possible to have too much of something; the minimal amount of template required works to the advantage of dPCR as a diagnostic tool in areas where large amounts of sample are not available.
dPCR is less than a decade old, and while the high cost of equipment and methods for droplet preparation may deter some from using it, dPCR is becoming more accessible. dPCR may never replace qPCR or even traditional PCR entirely, but it does offer distinct advantages. And, like all technologies, it will likely become more precise and affordable with time.







