November 15, 2009 (Vol. 29, No. 20)
Richard Hill Ph.D.
Research Using Smaller Collections Reported to Shorten Discovery Efforts
Chemical libraries have long been a mainstay in the search for new pharmaceutical compounds, and they have been created using many different paradigms. Vast diverse collections of unique compounds have been screened at high throughput to find appropriate effects on target proteins. Such large libraries continue to be used for drug discovery, but screening smaller, more focused libraries, can provide more efficient solutions with better overall hit rates.
Ten years ago, BioFocus®, a Galapagos company, launched a nonexclusive compound library specifically designed to target serine-threonine and tyrosine kinases. This library, SoftFocus® Kinase library one (SFK01), was designed to mimic the binding of ATP to the catalytic (hinge) region and had a rather simplistic design based around an aminopyrimidine core (Figure 1).
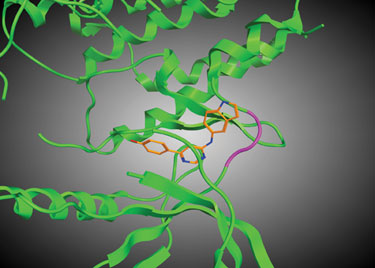
Figure 1. Ribbon diagram showing predicted binding of an example SFK01 compound to FGFR (hinge region shown in purple)
Targeting Kinases
Protein kinases are enzymes that phosphorylate substrate proteins at specified residues such as serine, threonine, and tyrosine. The phosphorylation of the substrate protein initiates a cascade, that in turn, modulates the transcription of a gene or set of genes. Kinases play pivotal roles in modulating diverse cellular activities, including growth, differentiation, metabolism, adhesion, motility, and death, and have been implicated as important mediators of certain forms of cancer.
Kinases, therefore, represent key druggable target proteins. The initial realization that most kinases possess highly conserved catalytic domains initially made kinase targets ideally suited for compound screening via focused chemical libraries in which the library compounds were specifically created to bind to the catalytic (hinge) regions.
Despite the slightly higher costs generally associated with the design and synthesis of such focused libraries compared to large and diverse compound collections, true savings can be gained as a result of shortened project cycles coupled with the reduced costs of screening, storage, and quality control of a smaller screening library.
The increasing wealth of structural data available along with a number of new techniques such as in silico design has enabled the continual development of kinase-focused collections, providing increasingly more sophisticated chemical structures. One of the key benefits of this is that the current range of SoftFocus kinase libraries has been designed to target additional binding modes to those involving the hinge region; most notably the DFG-out binding mode and the novel binding mode first observed in the kinase PIM-1.
In Silico Design
Current in silico design processes enable automated docking and scoring of scaffold ideas into a variety of known x-ray structures that have been selected, not only for broad coverage of the kinome, but also different conformational states of individual kinase enzymes. The design premise is that, if the core of the molecule or scaffold contains the key recognition groups for binding into one of the known conformational states, it has the potential to target any kinase.
However, as different side chains or monomers are added, the potential to gain specificity for one target over another becomes reality. The various docking methods used in the design of the BioFocus libraries include hinge binding, the DFG-out model, and novel binding modes.
Hinge Binding
Hinge-binding library designs are validated by docking a minimally substituted scaffold into various different high-resolution kinase x-ray structures. These structures have been selected from across the phylogenetic tree to ensure broad coverage of tyrosine and serine/threonine kinases.
The BioFocus Kinase Toolkit™ provides a two-dimensional map (2D Roadmap) of the key ligand-binding features within a customized ATP-site model, allowing predictions of affinity, selectivity, and likely off-target issues based on the compositions of the individual sub-sites. This knowledge can then be used to select the appropriate side chains or monomers with which to decorate the scaffolds.
DFG-Out Model
An approach to the design of kinase libraries with higher selectivity potential is to target the DFG-out allosteric pocket adjacent to the ATP site. BioFocus has developed a generalized binding model of the DFG-out pocket that enables the targeting of a range of inactive kinase conformations.
Binding Modes
A library design strategy focusing on alternative ligand-kinase binding modes is largely based on the novel binding modes observed in the cocrystal structure of a potent compound from a SoftFocus library (SFK33), bound to the kinase PIM-1 (Figure 2). In this case, the compound binds to a catalytic lysine residue while making no contact with the hinge region. This binding mode provides a unique paradigm for novel library design.
Only those scaffold ideas that pass this in silico docking are progressed to the next stage of evaluation, which includes a novelty check on structures and adherence to key hit/lead physicochemical property checks. Screening such libraries has generated information-rich data, generally consisting of higher hit rates (compared to screening diverse collections of unrelated compounds) together with key structure-activity relationship data that can speed up hit-to-lead programs.
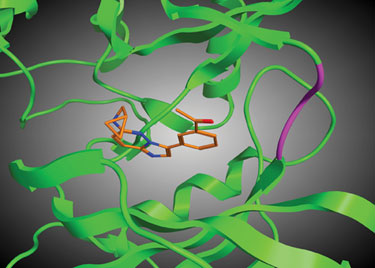
Figure 2. Ribbon diagram showing the binding of a potent SFK33 compound to the kinase PIM-1
Focusing on Success
By constantly refining the compounds using in silico methods, including those described above and with the development of innovative in silico models and applications such as Cresset BioMolecular’s molecular Field technology, the latest advances in kinase research can be incorporated into novel libraries. Consequently, this may also provide a strong intellectual property position to those who screen them. Indeed, based on the screening of SoftFocus kinase libraries alone, over 70 known patents have been applied for or granted.
A recent example from Galapagos highlights this. Following a screen of some 16,000 BioFocus focused kinase compounds, three hit series were identified that showed structure-activity relationships against a novel rheumatoid arthritis target. Two of these compound series were progressed to the hit-to-lead phase and subsequently one series was optimized, and a compound is currently undergoing clinical trials. The project time from screening to preclinical candidate nomination was three years.
There is no doubt that the drug discovery process should be shortened whenever possible. It is essential to improve the lead discovery process. Focused compound libraries such as the SoftFocus collection have the inherent capabilities to provide a robust hit discovery process, with good hit-to-lead conversion rates and shorter development times.
Furthermore, such libraries can maximize the benefit of new techniques such as in silico modeling, enabling them to be consistently developed over time. BioFocus has also maintained flexibility in its library-generation processes using these techniques, which has enabled the development of novel libraries with proven results.
Richard Hill, Ph.D. ([email protected]), is director of discovery
products, Asia and Pacific at BioFocus. Web: www.biofocus.com.







