July 1, 2010 (Vol. 30, No. 13)
John SantaLucia Jr. Ph.D. CEO and Co-Founder DNA Software
Gregory Boggy
Astrid Tuin
Sam Lee
Edward Sekinger, Ph.D.
Gordon Robertson, Ph.D.
Sensitive and Specific Primers and Probes Are Goals of New Modeling Platform
Common problems that arise during PCR assays include weak target-binding, competing target secondary structure, primer-dimer formation, and mishybridization of PCR probes and primers. These issues, rooted in DNA hybridization thermodynamics, often lead to artifacts that confound interpretation of assay results.
Most PCR assay design tools cannot design optimal PCR assays because they use overly simplified models of DNA hybridization and do not account for undesired effects. In contrast, the Oligonucleotide Modeling Platform™ (OMP™)—the engine that drives DNA Software’s Visual OMP™, ThermoBLAST™, and PCR Pathfinder™ molecular biology tools—employs a secondary structure folding algorithm, a multistate coupled equilibrium model, and a comprehensive database of nearest neighbor thermodynamic parameters to account for all thermodynamic effects under assay conditions. This article highlights the use of DNA Software’s molecular biology tools to design optimal PCR assays.
An original set of PCR primers was designed against a Ptprd cDNA target using the default settings of Primer3 in an attempt to validate the presence of a candidate novel intron in the 3´-UTR region of the Ptprd transcript. cDNA was obtained by reverse transcription (using random hexamers) of a crude sample of adult mouse liver. After 40 cycles of PCR, an electrophoresis gel displayed many bands (lane 2 of the gel in Figure 1A), whereas only two were expected, representing native Ptprd transcript and Ptprd transcript with the candidate intron.
To determine whether the undesired amplicons resulted from primer mishybridization to mouse cDNA or to targets resulting from external contamination, primers were scanned with ThermoBLAST, at 52ºC and PCR buffer conditions, against the mouse transcriptome. The ThermoBLAST scan found all thermodynamically stable mishybridization sites and potential false amplicons.
The resulting ThermoBLAST Virtual Gel in Figure 1B shows that multiple undesired amplicons could arise from PCR with the original primers and mouse transcriptome background, suggesting that these primers were improperly designed.
Because of the poor performance of the original primers, two new sets of primers were designed with PCR Pathfinder and Primer3 to compare the quality of designs from these two programs. Tools such as Primer3 use matched melting temperatures to rank candidate primer sets, calculated using a two-state model of nucleic acid hybridization.
In contrast, PCR Pathfinder and Visual OMP rank candidate primers according to specificity (low percentage of off-target binding) and sensitivity (high percentage bound to target) of primers under assay conditions, calculated using the multistate coupled equilibrium model.
PCR Pathfinder offers an intuitive, wizard interface that guides users through the process of designing high-quality PCR assays with up to six targets, while Visual OMP provides maximum flexibility for designing and simulating any nucleic acid hybridization assay with unlimited multiplexing capabilities.
To ensure amplification specificity of Ptprd primers, PCR Pathfinder was used in conjunction with ThermoBLAST, which automatically scanned primers against the mouse transcriptome. For comparison, primers from Primer3 were scanned using conventional BLAST against the background of the mouse transcriptome.
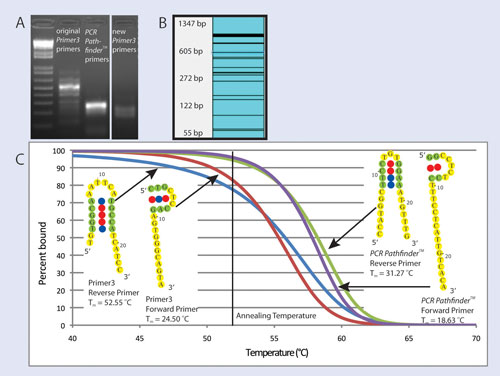
Visual OMP simulations of primer binding (Figure 1C) show that PCR Pathfinder primers are predicted to bind their target with 96% and 94% efficiency for forward and reverse primers, respectively, while forward and reverse Primer3 primers are predicted to bind their target with 82% and 77% efficiency, respectively.
The most stable secondary structures predicted by Visual OMP for each of the primers are also shown in Figure 1C. Secondary structures of the primers from PCR Pathfinder are of little concern because they are unstable as indicated by their low hairpin melting temperatures, whereas the reverse primer from Primer3 has a hairpin melting temperature above the annealing temperature, resulting in significant inhibition of primer binding observed in simulation results.
PCR performed with the redesigned primers resulted in the bands present in lanes 3 and 4 of Figure 1A for PCR Pathfinder and Primer3 designed primers, respectively. Both primer sets appear to be specific, resulting in the expected two bands; however, comparing the bright bands of lane 3 with the weak bands of lane 4 indicates that the primers designed in PCR Pathfinder bind more efficiently than the primers designed in Primer3, as predicted by simulation in Visual OMP.
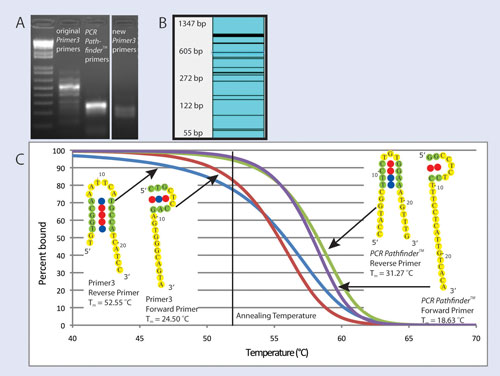
Figure 1. (A) Electrophoresis gel showing PCR products from primers targeting Ptprd; (B) Log-scale virtual gel that shows multiple undesired amplicons predicted by ThermoBLAST scanning of original Ptprd primers against the mouse transcriptome; (C) Visual OMP simulation of Ptprd primers binding to their desired target as a function of temperature. Also shown are the most stable unimolecular secondary structures and their melting temperatures for each primer as predicted by Visual OMP.
Luminex xTAG Technology
Multiplex real-time PCR is unable to detect all PCR targets in highly multiplexed assays, due to the limited number of channels that can be detected in a real-time assay. The Luminex xTAG® detection technology can circumvent this limitation and is appropriate for situations that require a high degree of multiplexing.
Multiplex PCR reactions are difficult to design, however, because amplicons are often synthesized with unequal efficiencies due to unequal hybridization thermodynamics, product inhibition of DNA polymerase, and primer-dimer interactions. Visual OMP is well suited for multiplex PCR because it designs primers so that all amplicons are synthesized with equal efficiency, while simultaneously avoiding undesired thermodynamic effects.
To validate the Visual OMP and Luminex xTAG technologies for designing and quantifying multiplex PCR experiments, an eight-plex PCR assay was designed in Visual OMP for detection by Luminex xTAG technology. cDNA targets were obtained following reverse transcription (using polyT primers) of eight polyadenylated RNAs spiked into human liver total RNA at different concentrations.
PCR primers were designed in Visual OMP and scanned with ThermoBLAST against the human transcriptome database. Simulating all primers together with their targets at actual reaction conditions in Visual OMP confirmed that the primers were specific (less than 0.01% primer binding to each possible off-target sequence) and sensitive for their targets, binding at greater than 99.9% efficiency, suggesting that all targets should amplify equally well.
Figure 2 shows data obtained by Luminex xTAG technology from Visual OMP-designed monoplex and multiplex PCR assays. The singleplex and multiplex reactions resulted in similar trends in intensity with respect to copy number, indicating that assay design in Visual OMP resulted in a reliable multiplex PCR assay.
PCR assay design and optimization is typically a complex and time-intensive process, particularly for multiplexed assays where difficulties scale with the number of targets in the reaction. Enabled by accurate simulation capabilities, PCR Pathfinder, Visual OMP, and ThermoBLAST allow users to design optimal PCR assays without the need for trial-and-error optimization.
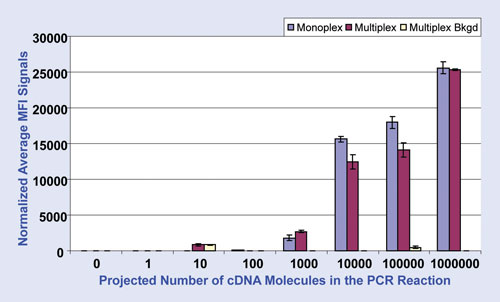
Figure 2. Luminex xTAG data from monoplex (purple) and multiplex (magenta) PCR assays using primers designed in Visual OMP. Each of the RNA sequences was spiked into the reverse transcription reaction at a different concentration, resulting in the projected cDNA copy numbers shown. xTAG data shown are normalized average values of the median fluorescence intensity (MFI) obtained from PCR assays repeated in triplicate, after 28 cycles of PCR.
John SantaLucia Jr., Ph.D. ([email protected]), is CSO, Gregory Boggy is business development manager, and Astrid Tuin is director of customer solutions at DNA Software. Web: www.dnasoftware.com. Sam Lee is a Ph.D. candidate at the British Columbia Cancer Research Center. Edward Sekinger, Ph.D., is senior research scientist in the assay advanced technology group at Luminex. Gordon Robertson, Ph.D., is staff scientist at the British Columbia Genome Sciences Center.