June 15, 2012 (Vol. 32, No. 12)
MaryAnn Labant
Aberrant expression of microRNA (miRNA) in human cancers is a common phenomenon. miRNAs regulate many tumor suppressor genes and oncogenes, therefore acting as oncogenes or tumor suppressor genes themselves to directly regulate cancer cell survival and proliferation.
In tumor cells, inhibiting miRNAs that are overexpressed, and restoring intracellular levels of those that are lost or otherwise underexpressed, are attractive approaches for cancer treatment. Additionally, the role miRNAs play in modulating cancer cell response to chemotherapeutic agents suggests that they may be a target for improving drug response in cancer therapy.
These naturally occurring 19–23 nucleotide long, single-stranded noncoding RNAs, which regulate gene expression largely by decreasing levels of target messenger RNAs, were highlighted at the recent Select Biosciences “Genomics Research” conference.
“As we start to look at the full genomic pattern of a tumor tissue as opposed to a normal tissue we see lots of changes over the course of tumor development. miRNA likely has hundreds of targets in the cell. The goal is to identify specific mutations and pair those with drugs that inhibit or activate pathways,” explained Anthony Saleh, Ph.D., IRTA scientist, National Institute on Deafness and Other Communication Disorders at NIH.
To determine the targets, researchers use genomic, transcriptomic, or proteomic approaches. The difficulty with omics-type experiments is the scope. Myriad miRNAs are overexpressed in cancer, although most are not very highly overexpressed. Targets are painstakingly narrowed down by expression level, consistent expression levels across samples, and biological function.
“We study miRNA expression in head and neck squamous cell carcinoma (HNSCC) lines grown from patients’ tumors and have identified a number of overexpressed miRNAs that are potentially oncogenic, in particular the miR-30 family,” continued Dr. Saleh.
“Other studies showed NF-kappaB was activating expression. NF-kappaB is, in general, an oncogene, promoting tumor formation. We also identified TP53 and Notch1, which are important tumor suppressors for HNSCC, as miR-30 targets. So in HNSCC miR-30 is being stimulated by an oncogene and suppressing two important tumor suppressor pathways.”
TP53, the most common tumor suppressor pathway, primarily functions to direct cell fate decisions. In 90% of HNSCC cases, TP53 is nonfunctional, deleted, or mutated.
Notch1 has different roles in different cancers and highlights the difficulty in studying cancer. In HNSCC it is a tumor cell suppressor. In other cancers it is an oncogene.
“Now we are determining if miR-30 might be a potential target for sensitizing tumors. If we decrease the levels of miR-30 through inhibitors, we may be able to slow down the cancer growth and, perhaps, kill it in combination with other chemotherapeutic drugs or radiation,” concluded Dr. Saleh.
miRNAs have a critical role in cancer, along with a unique mechanism of action, which make them an intriguing new class of diagnostic, prognostic, and therapeutic tools. Researchers have reversed the growth of lung tumors in mice using the naturally occurring tumor suppressor microRNA let-7. [Carol and Mike Werner/Photo Researchers]
Dx, Px, and Tx Tools
“If you consider the critical role of miRNAs in cancer coupled with their unique mechanism of action, it is clear that miRNAs represent a new class of diagnostic, prognostic, and therapeutic tools,” commented Alexander Pertsemlidis, Ph.D., assistant professor, Greehey Children’s Cancer Research Institute at the University of Texas Health Science Center at San Antonio.
“The idea of screening libraries of miRNA mimics, which increase intracellular levels of miRNAs, and inhibitors, which decrease those levels, has been around for a while, but is represented more in the patent literature than in the biomedical literature. We use high-throughput screening (HTS) to identify miRNA mimics and inhibitors that specifically sensitize cancer cells to widely used anticancer drugs.”
For example, miR-337-3p appears to be lost in lung cancer cells, with replacement selectively sensitizing cells to paclitaxel by enhancing taxane-induced G2/M arrest through downregulation of its targets. Increasing levels of miR-337-3p may provide a novel therapeutic tool for the treatment of non-small-cell lung cancer in conjunction with taxanes.
Conversely, miR-139-5p is present at aberrantly high levels in neuroendocrine lung cancer cells, and when removed with an inhibitor, selectively decreases tumor cell viability. Inhibiting miR-139-5p may therefore be a tool for treating neuroendocrine lung tumors.
Dr. Pertsemlidis’ investigations contribute to the understanding of miRNA roles in, and beyond, lung cancer pathogenesis. A number of miRNAs are aberrantly expressed in several different cancer types. Mimics or inhibitors that are selectively cytotoxic in lung cancer may also have therapeutic applications in breast cancer, and miRNA targets important to disease progression in neuroendocrine lung cancers may also play critical roles in other neuroendocrine cancers, such as neuroblastoma.
Standardizing RNAi Screens
Cellecta designs and constructs pooled libraries expressing short hairpin RNAs (shRNA) and performs genetic screens using these libraries to identify genes required for any cellular response with a strong selectable phenotype, such as genes required for growth and proliferation of cancer cells.
shRNA consists of a sense strand, short loop sequence, and antisense strand that, when expressed in cells, interferes with gene expression similar to synthetic siRNA. In addition to single shRNA screening, Cellecta has developed an approach to combinatorially screen shRNA pairs to discover synthetically lethal and synergistically lethal gene combinations.
The company’s shRNA libraries for pooled screening have optimized structures to increase stability, protocols for uniform en masse cloning of large heterogeneous pools of shRNA sequences into the lentiviral library vector, and specially designed shRNA-specific barcode sequences incorporated for HT sequencing to enable accurate assessment of the libraries’ shRNAs.
“It is not trivial to create a complex yet representative library that expresses multiple shRNAs to different targets, with each construct containing a unique sequenceable barcode that identifies the specific shRNA it contains,” commented Alex Chenchik, Ph.D., president and CSO.
With grant funding from the NIH National Center for Research Resources and National Human Genome Institute, Cellecta established The Decipher Project to help standardize the process of RNAi-based screens. “Cellecta’s goal with Decipher is to provide researchers investigating disease progression, cell-signaling pathways, or genes involved in a range of biological processes, the basic tools to conduct genome-scale genetic screens,” said Dr. Chenchik.
Through Decipher, Cellecta offers academic labs no-cost access to three plasmid human shRNA libraries that target 15,000 genes, and two for mouse that target 10,000 genes. In exchange, researchers agree to publicly share all genetic screen data generated after publication. In the last year, over 100 researchers accessed the Decipher libraries, according to Dr. Chenchik.
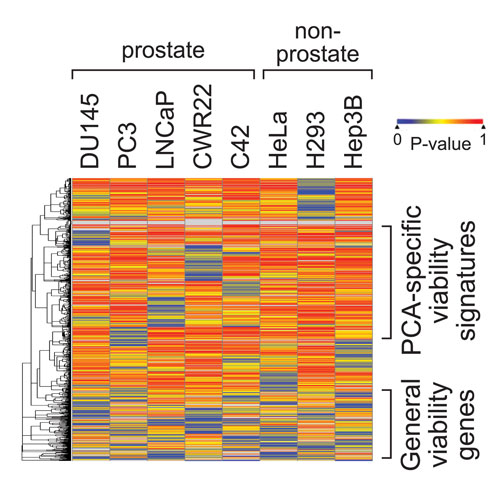
Cellecta RNAi screening heatmap shows genes that were identified as essential in eight different cell lines. The screen was conducted to find essential genes in blood cancers that could be potential drug targets. Each band represents a gene. The left hierarchy indicates the gene groupings. “More red” bands indicate that the shRNAs, targeting the gene, did not significantly affect the cell; “more blue” bands indicate that shRNAs were lethal to the cell line, so the gene is likely to be essential to survival of the cells.
Knockout Mice Models
Cell culture models provide valuable information concerning miRNA expression. The question is whether agents that inhibit the miRNA activity decrease activity so low that it is essentially the same as removing the gene. A low expression does not provide a true phenotype.
Gene targeting, or gene knockout, experiments are often used to inactivate single genes in mouse systems. Conditional knockout is based on a tissue-specific inactivation of the gene of interest, which can be achieved by means of a recombinase, such as CRE recombinase.
“Years ago, in collaboration with Brian Harfe, Ph.D., Cliff Tabin, Ph.D., and Phil Sharp, Ph.D., we created a conditional Dicer knockout mouse. These low-resolution studies informed us as to the overall importance for Dicer, a miRNA processing enzyme, in a given tissue. However, ablation of Dicer affects nearly all miRNAs and it is difficult, if not impossible, to identify which miRNA(s) are responsible for a given phenotype,” said Michael McManus, Ph.D., assistant professor, UCSF.
“So I started a challenging project to ablate 100 evolutionarily conserved noncoding RNA genes in the mouse, in the hopes that we might gain better insight into the roles for miRNA expression in a given tissue. We mutate the miRNA genes and then also do a gene replacement with a colorimetric reporter gene, the LacZ gene. Now we can see at a very granular level the specific cell types that are expressing that miRNA.”
The reagents have been released for public use and are available to investigators from the public repositories.
“Given the number of investigators exploring miRNAs in human diseases, it is only a matter of time before we see interesting studies that take advantage of this increased fantastic resource. The genome is a vast sea of uncharted territory, containing many noncoding RNAs with unknown function, and we would like to expand our project to the study of other noncoding RNAs,” concluded Dr. McManus.

Modification of this genetically modified miRNA gene allows for the programmed removal of the miRNA in specific tissues. A second modification with a colorimetric reporter gene allows for tracking the expression of the miRNA in the mouse. Specifically, everywhere the miRNA is expressed, the tissue will turn blue. [Michael McManus, University of California San Francisco]
Delivery and Targeting Hurdles
“Since the turn of the century when miRNAs were first implicated in human cancers, the field of miRNA research has grown exponentially,” said Christopher Cheng, Ph.D., postdoctoral researcher, Yale University. “Much of the research has involved elucidating the role of miRNAs in cancer, and we are now at a time when our understanding of miRNA biology will inform the development of new therapeutics. Due to their familiar chemical nature, there is a prevailing hope that miRNAs may be readily exploited as therapeutic tools by dovetailing off of established nucleic-acid delivery strategies.”
Two potential ways, with different delivery-system requirements, exist to use miRNAs as therapeutics. One is to deliver a precursor or synthetic tumor suppressor miRNA, and the other is to deliver an agent that inhibits endogenous oncogenic miRNA.
Delivery of tumor suppressor miRNAs can capitalize on established liposomal-based or nanoscale strategies for delivering siRNA, such as those pioneered by Tekmira or Alnylam.
Challenges abound. As with most nucleic-acid therapeutics in vivo, without an effective carrier, rapid degradation would occur. Delivery vehicles provide protection against degradation, but these carriers can be cleared from the bloodstream by the immune system and as a result, typically accumulate in the liver.
Cellular challenges emerge once the agents enter the cells of interest, usually through endocytosis. In the endocytotic pathway, which is one of the cell’s natural ways for taking up extracellular components, agents can eventually be degraded in lysosomes. Therefore, a delivery vehicle must be designed to escape the endocytosis pathway, which is no easy feat.
Unlike tumor suppressor miRNAs, miRNA inhibitors do not need to directly interact with the RNA interference machinery, which allows more freedom to employ chemical enhancements and modifications and opens up new delivery options for miRNA antagonists.
Targeting remains a main hurdle. Principles learned from studies over the last several decades to improve the delivery of therapeutics to tumors are now being applied to the miRNA field.
There is still a long way to go. Currently, drugs do not exist to target all of the identified mutations. For over 30 years, researchers have been trying to target TP53, the most mutated gene in cancer, and have yet to be successful.
Clinical Application of a Tumor Suppressor miRNA
The recent discovery of human miRNAs has been a revelation to biologists who have sought to understand the impact of the small, regulatory RNAs on cell function and human health.
The most active area of research has been in cancer where more than 6,000 peer-reviewed publications have identified and characterized a handful of mi-?RNAs that regulate key cancer-associated genes and pathways. More than 200 review articles have contemplated the potential use of tumor suppressor mi-RNAs to treat cancer patients.
Researchers at Mirna Therapeutics say they are preparing to test whether the optimism surrounding miRNA-based therapies in cancer has been warranted. The company’s lead therapeutic candidate is a mimic of miR-34, a tumor suppressor miRNA that functions within the p53 pathway.
“miR-34 induces cell cycle arrest, senescence, and apoptosis in cancer cells by regulating the expression of more than 20 oncogenes including MET, MYC, WNT, MYB, RRAS, NANOG, BCL2, and CDK4,” says David Brown, Ph.D., director of research at the company.
“We overcame a key hurdle for the miR-34-based therapy when we in-licensed a systemic delivery technology from Marina Biotechnology after an extensive evaluation of liposome- and polymer-based formulations that were developed for other oligonucleotide-based therapies.”
The formulated miR-34 mimic called MRX34 has produced complete tumor regression in two separate orthotopic models of liver cancer and has also displayed therapeutic activity in models of lung and metastatic colorectal cancer, according to Dr. Brown.
“Unlike many siRNA-based therapies, MRX34 has displayed no immunostimulatory activity in rodents and human whole blood, and preliminary toxicology studies have revealed no adverse effects in normal tissues at therapeutic doses,” he continues.
“Additional toxicology studies scheduled for the second half of 2012 will support an Investigational New Drug Application that will clear the way for a Phase I clinical trial in the beginning of 2013. As the first clinical test for a tumor suppressor miRNA, the trial will provide the first glimpse into the therapeutic potential of these small, regulatory RNAs.”







