Biological assays in established and primary cells can greatly aid the drug discovery process. Stable cell (inheritable genotype) lines are chosen for their consistent profile and efficient measure of compound activity. Transient cell-based assays offer greater flexibility with regards to the choice of cell type but require introduction of exogenous DNA into the chosen cell line, prior to performing each experiment. Use of traditional lipid-based DNA delivery systems require DNA precomplex formation, and assays often suffer from low-end (and variable) target gene expression, leading to poor reproducibility of experimental data.
Improvements to transient cell-based assays can be made by use of more efficient delivery systems such as those based on adenovirus. These systems circumvent the necessity to prepare DNA complexes for each experiment, and allow high transduction efficiency and control of the level of target gene expression in the cell population.
GE Healthcare has launched a range of engineered vectors (Ad-A-Gene) that use adenovirus as the vehicle for efficient delivery of signal pathway sensors into mammalian cells. Ad-A-Gene vectors comprised of genes encoding key cellular proteins are fused either to a protein reporter (fluorescent) or a gene reporter (transcription factor regulating expression of nitroreductase).
The reporter is transiently maintained within the transduced cells, and cellular assays can be developed by monitoring the signal derived from the reporter group. We compare the efficiency and performance of adenoviral vectors to plasmid-based transfection methods to successfully deliver sensors into cells, and report how cells, transduced with either the EGFP-GCCR (enhanced green fluorescent protein-GCCR) fusion protein or GRE-NTR reporter, respond to dexamethasone.
Methods
Viral Transduction and Plasmid Transfection. Transduction was accomplished by adding the EGFP-GCCR or GRE-NTR adenovirus directly to cells at the appropriate multiplicity of infection (MOI).
Transfection of plasmid pDC515 containing the EGFP-GCCR open reading frame expressed from the human ubiquitin C promoter was performed according to the manufacturer’ recommendations.
Determining Transduction/Transfection Efficiency with IN Cell Analyzer 1000. EGFP-GCCR was used to determine the transduction/transfection efficiency of a number of cell lines. Image acquisition was performed using a 360/40-nm excitation filter (Hoechst), 475/20-nm excitation filter (EGFP), and 535/20-nm emission filter. An Object Intensity Analysis algorithm was used to measure the intensity of EGFP in the cytoplasmic region. EGFP intensities determined for nontransduced/nontransfected cells (plus standard deviation) were used to define a threshold. Cells in which EGFP cytoplasmic intensities exceeded the threshold were considered to be positive for the reporter.
Determining Transduction/ Transfection Efficiency by Flow Cytometry. A FACSCalibur flow cytometer was used to assess transduction/transfection efficiency. Non-transduced/non-transfected cells were used to define the threshold for non-expression (negative), and transduced/transfected cells were gated into a second group, corresponding to expression of EGFP (positive).
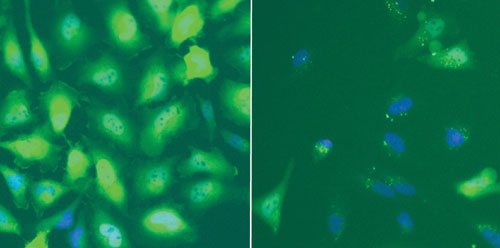
Functional EGFP-GCCR Assay. Cells were serum starved (1 h) prior to the addition of dexamethasone (30 min). Dexamethasone promotes the cytoplasm to nuclear translocation of GCCR. Cells were formalin-fixed and Hoechst nuclear stain was used to identify cells.
Functional GRE-NTR Assay. Growing cells were incubated (37C, 5% CO2) in either serum-free medium or serum-free medium containing dexamethasone for 24 h, followed by a 2 h further incubation in the presence of 1-µ CytoCy5S, and read on a Tecan Ultra Fluorimeter. Dexamethasone promotes the expression of NTR, which is detected through conversion of the pro-fluorescent substrate to a fluorescent product.
 Results
Results
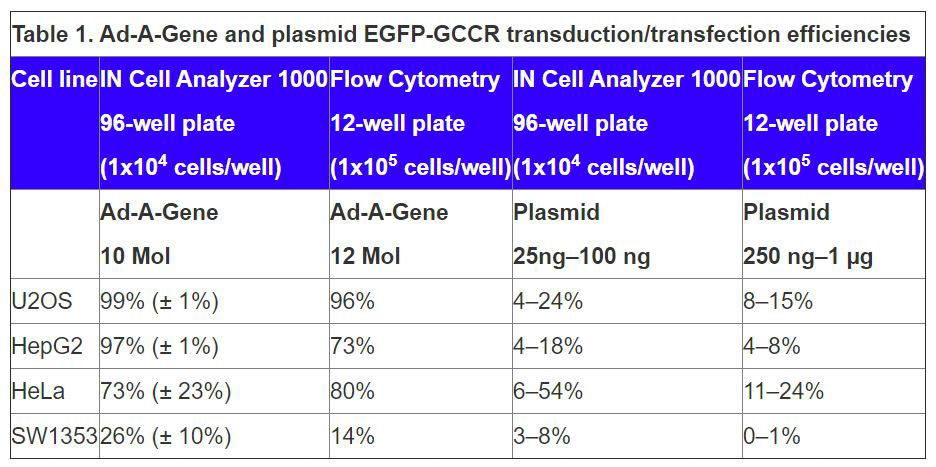
Adenoviral Transduction. Comparable values for transduction efficiency were determined by the IN Cell Analyzer 1000 and by flow cytometry . HepG2, HeLa, and U2OS cells transduce efficiently (73????%) with SW1353 cells being less susceptible. Data are comparable to those in a previous study, which reported adenoviral-mediated transduction efficiencies for U2OS and SW1353 cells.
Lipid-based Transfection. Comparable values for transfection efficiency were determined by the IN Cell Analyzer 1000 and by flow cytometry (Table 1). EGFP-GCCR plasmid efficiencies using a leading commercial lipid system were lower than equivalent transduction obtained with Ad-A-Gene vectors.
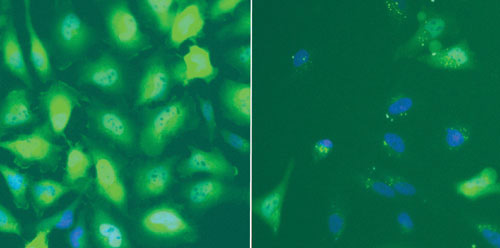
Ad-A-Gene Transduction and Plasmid-based TransfectionComparison in an Assay System. The EGFP-GCCR Ad-A-Gene transduced all cell types tested with consistent efficiencies. In contrast, plasmid- transfection efficiencies exhibited a large variation that was cell type-dependent. HeLa and U2OS cells were more amenable to plasmid-transfection compared to either HepG2 or SW1353 cells (Figure 1).
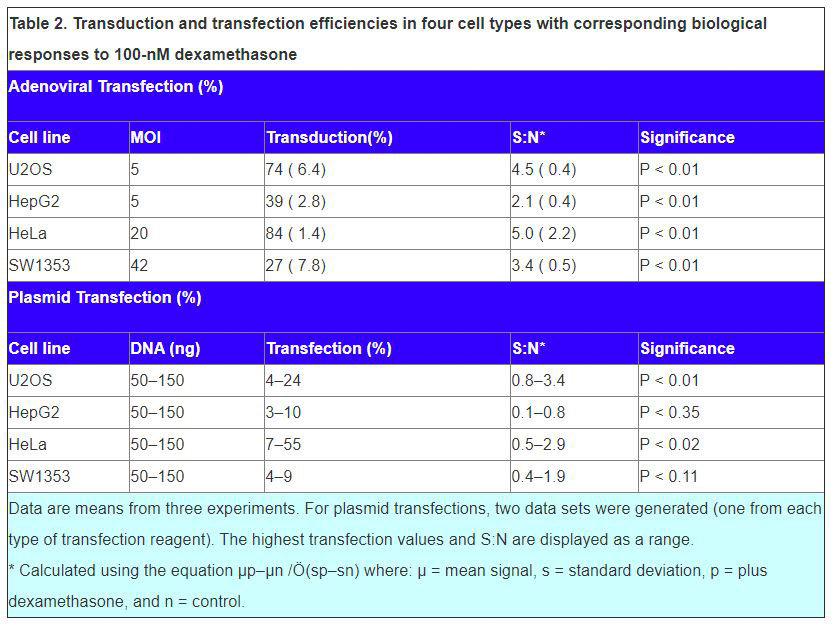
Data of dexamethasone response of cells transduced with the EGFP-GCCR adenovirus show acceptable S:N ratios and p-values in all cell types. In SW1353 and HepG2 cells (which exhibited lower transduction levels at the MOI used), the Nuclear Trafficking Analysis algorithm was able to resolve an acceptable dexamethasone response.
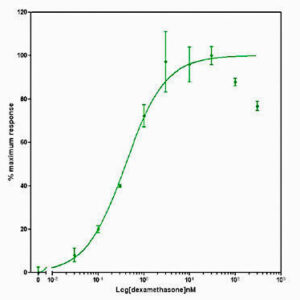
Plasmid-transfection efficiencies were comparably low in all cell types. Acceptable S:N ratios and p-values were obtained in HeLa and U2OS cells, although significant variation in S:N ratio was observed. In general, poor S:N ratios and p-values were determined for plasmid- transfected HepG2 and SW1353 cells.
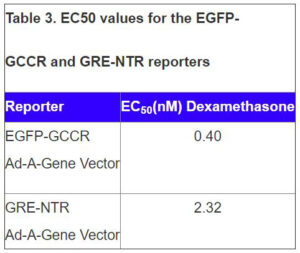
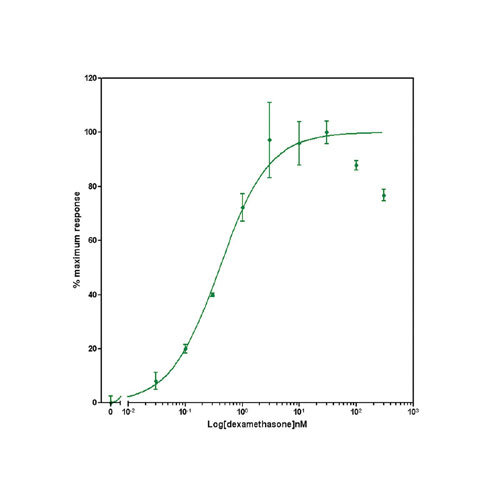
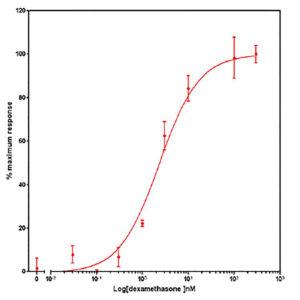
Data presented in Figure 2 shows the response of EGFP-GCCR and GRE-NTR to increasing concentrations of dexamethasone. This data demonstrates the successful transduction and development of two Ad-A-Gene vectors into assays for GCCR biology.
 Conclusion
Conclusion
Adenoviral transduction has several advantages over the current plasmid-transfection methods. Ad-A-Gene protocols are simple to perform, involving the direct addition of virus to cells. The use of Ad-A-Gene vectors does not require any additional reagents. Observable adenoviral transduction efficiency is high and reproducible in many of the cell types tested when used at an optimal MOI. The data also show that adenoviral transduction exhibits a robust and measurable assay response to dexamethasone in cells transduced with either the EGFP-GCCR or GRE-NTR adenovirus.
Tables