February 1, 2007 (Vol. 27, No. 3)
Deepika deSilva
Jana Blackett
Developing a Simple and Inexpensive Genotyping Assay with LunaProbes
High-resolution melting has proven to be a highly sensitive method for mutation discovery. PCR is performed in the presence of the double-strand DNA binding dye LCGreen® Plus. Samples are then denatured on the Idaho Technology (www.idahotech.com) LightScanner® instrument. Nucleic acid melting is tracked by monitoring the fluorescence of the samples across a defined temperature range, generating melting profiles that can be used to identify the presence of sequence variation within the amplicon. Single point mutations, small insertions, and deletions can all be identified using high-resolution melting. The technique has been shown to have near 100% sensitivity and specificity when used on products up to 400 base pairs in length.
LunaProbes™, an extension of the high-resolution DNA melting technique, provides a simple and inexpensive alternative for genotyping. An unlabeled oligonucleotide probe, blocked at the 3´ end to prevent extension, is designed to sit over a SNP of interest and is included in the PCR reaction prior to amplification.
Genotyping is accomplished by monitoring the melting of the probe-target duplex post-PCR. Key to the success of this method is the use of asymmetric PCR, where one primer is used in excess, resulting in the overproduction of the target strand recognized by the probe and the use of LCGreen Plus dye, which is capable of producing a strong fluorescent signal from the probe-target interaction.
Genotyping by melting analysis is beneficial to investigators primarily due to the cost-effectiveness of the technique. In the past, using PCR methods for genotyping has required fluorescently labeled oligonucleotides. These methods can be costly and time consuming. With LunaProbes, all three possible genotypes that result from a given polymorphism can be detected with a single unlabeled probe.
Described is the assessment of a common polymorphism (c1155C>G) in the phenylalanine hydroxylase (PAH) gene using high-resolution melting, LCGreen Plus dye, and a LunaProbe.
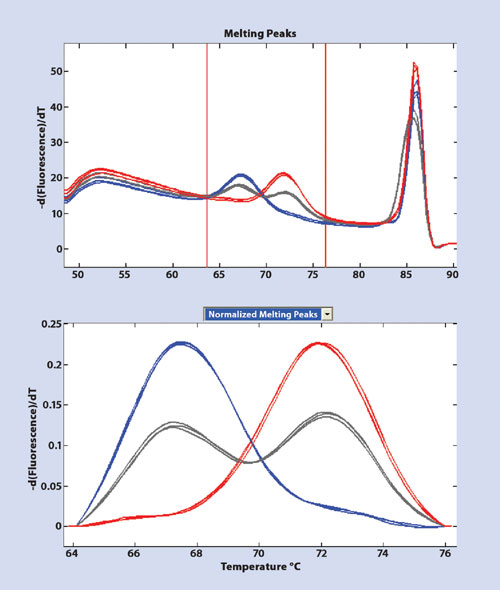
Figure 1
Methods
DNA samples from anonymous blood donors were extracted using a Qiagen(www.qiagen.com) kit. DNA samples were resuspended to a concentration of 10 ng/µL. A useful tool to verify that amplification of pseudogenes, or duplications will not occur is an In-Silico PCR tool (http://.genome.ucsc.edu).
While designing primers it is important to locate the mutation of interest and determine all the known variants present in the region of the amplicon to minimize the chance that the primers are placed over polymorphisms. When designing primers for genotyping, a Tm of about 60ºC is recommended. Probes should typically be designed to have a Tm of about 5º above the Tm of the primers.
Recommended probe length is between 20 and 35 bases, with the mutation site located in the middle of the probe to maximize the instability with the mismatched variant. To prevent extension of the probe during PCR, a 3´ block is required. Either C3 or phosphate may be used to block the probe at the 3´ end. However, over time the phosphate group can be lost, allowing extension beyond the probe site, therefore C3 is the preferred option.
In this study the SNP of interest is found in PAH exon 11, rs772897. The PCR product was 260 base pairs in length. The forward primer sequence was 5´ GATGCAGCAGGGAATACTGATC 3´, the reverse primer sequence was 5´ AGATGAGTGGCACCAGTCAGG 3´ and the probe 5´ GTTCCAGCCCCTGTATTACGTGGC-Blocked 3´.
Reaction Chemistry
Asymmetric PCR was performed in 10 µL reactions. LightScanner Master mix was used at a 1X concentration. The final concentration of template DNA was 10 ng. A 1:5 ratio of primers was used. The final concentrations of primers were: 0.1 µM limiting primer; 0.5 µM excess primer, and 0.5 µM probe. This asymmetry was designed so that the probe is bound to the strand produced in excess.
PCR was performed on a conventional block thermo cycler in a 96-well plate for 55 cycles with denaturation at 94ºC (30 seconds), annealing at 64ºC (30 seconds), and extension at 72ºC (30 seconds). A three-step PCR protocol was found to be essential for PCR success. Elongation at 72ºC allowed for robust amplification of the product without interference from probe-template interactions. Due to the reduced efficiency of asymmetric PCR, 55 cycles of amplification were required to generate sufficient template to produce a robust signal.
Melting Curve Acquistion and Analysis
Plates were taken from the thermo cylcer and subjected to a high-resolution denaturation step on the LightScanner. DNA melting was monitored from 55ºC to 85ºC, a process that took approximately five minutes. After data acquisition, the region of the probe melt was bracketed for further analysis, fluorescence was normalized to correct for nonspecific signal, and the melting curves were displayed as derivative melting peaks for simpler data read out. All of these analysis steps were performed automatically using the LightScanner genotyping software.
Results
DNA amplified in the presence of the DNA binding dye LC Green plus emits a high level of fluorescence when double stranded but loses fluorescence as the DNA duplex melts apart. In a LunaProbe genotyping experiment, fluorescent melting readouts are generated by dissociation of the short probe-target duplexes as well as the much larger target-target duplexes resulting in two distinct regions of DNA melting. Figure 1a illustrates the derivative curves obtained over the entire temperature range. Probe-target melting is observed between 65–75ºC, while the amplicon melting occurs between 82–87ºC.
In this example, a probe was designed to complement the G allele.
A perfectly matched probe-target hybrid has a characteristic melting temperature that is higher than a mismatched hybrid. Closer examination of the region of probe melting showed that samples with the G allele had a derivative melting peak at 72ºC, whereas samples harboring the C variant showed a melting peak at 67ºC. The heterozygous samples (grey) showed two peaks one at each temperature representing the combination of both alleles. Therefore, a single probe was able to recognize all three genotypes within the given sample set (Figure 1b).
Conclusion
High-resolution melting is not only a sensitive technique for mutation scanning, but also for genotyping. A LunaProbe used in conjunction with LCGreen Plus dye is a simple and effective way to genotype, eliminating the need for expensive fluorescent-labeled probes.
Deepika deSilva, Ph.D. is director of life science, and Jana Blackett is life science technical support specialist at Idaho Technology. Web: www.idahotech.com. Phone: (801) 736-6354. Email: [email protected].